Microarray-Based Prediction of Cytotoxicity of Tumor Cells to Arsenic Trioxide
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

Genetic and Genomic Analysis of Hyperlipidemia, Obesity and Diabetes Using (C57BL/6J × TALLYHO/Jngj) F2 Mice
University of Tennessee, Knoxville TRACE: Tennessee Research and Creative Exchange Nutrition Publications and Other Works Nutrition 12-19-2010 Genetic and genomic analysis of hyperlipidemia, obesity and diabetes using (C57BL/6J × TALLYHO/JngJ) F2 mice Taryn P. Stewart Marshall University Hyoung Y. Kim University of Tennessee - Knoxville, [email protected] Arnold M. Saxton University of Tennessee - Knoxville, [email protected] Jung H. Kim Marshall University Follow this and additional works at: https://trace.tennessee.edu/utk_nutrpubs Part of the Animal Sciences Commons, and the Nutrition Commons Recommended Citation BMC Genomics 2010, 11:713 doi:10.1186/1471-2164-11-713 This Article is brought to you for free and open access by the Nutrition at TRACE: Tennessee Research and Creative Exchange. It has been accepted for inclusion in Nutrition Publications and Other Works by an authorized administrator of TRACE: Tennessee Research and Creative Exchange. For more information, please contact [email protected]. Stewart et al. BMC Genomics 2010, 11:713 http://www.biomedcentral.com/1471-2164/11/713 RESEARCH ARTICLE Open Access Genetic and genomic analysis of hyperlipidemia, obesity and diabetes using (C57BL/6J × TALLYHO/JngJ) F2 mice Taryn P Stewart1, Hyoung Yon Kim2, Arnold M Saxton3, Jung Han Kim1* Abstract Background: Type 2 diabetes (T2D) is the most common form of diabetes in humans and is closely associated with dyslipidemia and obesity that magnifies the mortality and morbidity related to T2D. The genetic contribution to human T2D and related metabolic disorders is evident, and mostly follows polygenic inheritance. The TALLYHO/ JngJ (TH) mice are a polygenic model for T2D characterized by obesity, hyperinsulinemia, impaired glucose uptake and tolerance, hyperlipidemia, and hyperglycemia. -

Bioinformatics Analyses of Genomic Imprinting
Bioinformatics Analyses of Genomic Imprinting Dissertation zur Erlangung des Grades des Doktors der Naturwissenschaften der Naturwissenschaftlich-Technischen Fakultät III Chemie, Pharmazie, Bio- und Werkstoffwissenschaften der Universität des Saarlandes von Barbara Hutter Saarbrücken 2009 Tag des Kolloquiums: 08.12.2009 Dekan: Prof. Dr.-Ing. Stefan Diebels Berichterstatter: Prof. Dr. Volkhard Helms Priv.-Doz. Dr. Martina Paulsen Vorsitz: Prof. Dr. Jörn Walter Akad. Mitarbeiter: Dr. Tihamér Geyer Table of contents Summary________________________________________________________________ I Zusammenfassung ________________________________________________________ I Acknowledgements _______________________________________________________II Abbreviations ___________________________________________________________ III Chapter 1 – Introduction __________________________________________________ 1 1.1 Important terms and concepts related to genomic imprinting __________________________ 2 1.2 CpG islands as regulatory elements ______________________________________________ 3 1.3 Differentially methylated regions and imprinting clusters_____________________________ 6 1.4 Reading the imprint __________________________________________________________ 8 1.5 Chromatin marks at imprinted regions___________________________________________ 10 1.6 Roles of repetitive elements ___________________________________________________ 12 1.7 Functional implications of imprinted genes _______________________________________ 14 1.8 Evolution and parental conflict ________________________________________________ -

A Flexible Microfluidic System for Single-Cell Transcriptome Profiling
www.nature.com/scientificreports OPEN A fexible microfuidic system for single‑cell transcriptome profling elucidates phased transcriptional regulators of cell cycle Karen Davey1,7, Daniel Wong2,7, Filip Konopacki2, Eugene Kwa1, Tony Ly3, Heike Fiegler2 & Christopher R. Sibley 1,4,5,6* Single cell transcriptome profling has emerged as a breakthrough technology for the high‑resolution understanding of complex cellular systems. Here we report a fexible, cost‑efective and user‑ friendly droplet‑based microfuidics system, called the Nadia Instrument, that can allow 3′ mRNA capture of ~ 50,000 single cells or individual nuclei in a single run. The precise pressure‑based system demonstrates highly reproducible droplet size, low doublet rates and high mRNA capture efciencies that compare favorably in the feld. Moreover, when combined with the Nadia Innovate, the system can be transformed into an adaptable setup that enables use of diferent bufers and barcoded bead confgurations to facilitate diverse applications. Finally, by 3′ mRNA profling asynchronous human and mouse cells at diferent phases of the cell cycle, we demonstrate the system’s ability to readily distinguish distinct cell populations and infer underlying transcriptional regulatory networks. Notably this provided supportive evidence for multiple transcription factors that had little or no known link to the cell cycle (e.g. DRAP1, ZKSCAN1 and CEBPZ). In summary, the Nadia platform represents a promising and fexible technology for future transcriptomic studies, and other related applications, at cell resolution. Single cell transcriptome profling has recently emerged as a breakthrough technology for understanding how cellular heterogeneity contributes to complex biological systems. Indeed, cultured cells, microorganisms, biopsies, blood and other tissues can be rapidly profled for quantifcation of gene expression at cell resolution. -

Targeting PH Domain Proteins for Cancer Therapy
The Texas Medical Center Library DigitalCommons@TMC The University of Texas MD Anderson Cancer Center UTHealth Graduate School of The University of Texas MD Anderson Cancer Biomedical Sciences Dissertations and Theses Center UTHealth Graduate School of (Open Access) Biomedical Sciences 12-2018 Targeting PH domain proteins for cancer therapy Zhi Tan Follow this and additional works at: https://digitalcommons.library.tmc.edu/utgsbs_dissertations Part of the Bioinformatics Commons, Medicinal Chemistry and Pharmaceutics Commons, Neoplasms Commons, and the Pharmacology Commons Recommended Citation Tan, Zhi, "Targeting PH domain proteins for cancer therapy" (2018). The University of Texas MD Anderson Cancer Center UTHealth Graduate School of Biomedical Sciences Dissertations and Theses (Open Access). 910. https://digitalcommons.library.tmc.edu/utgsbs_dissertations/910 This Dissertation (PhD) is brought to you for free and open access by the The University of Texas MD Anderson Cancer Center UTHealth Graduate School of Biomedical Sciences at DigitalCommons@TMC. It has been accepted for inclusion in The University of Texas MD Anderson Cancer Center UTHealth Graduate School of Biomedical Sciences Dissertations and Theses (Open Access) by an authorized administrator of DigitalCommons@TMC. For more information, please contact [email protected]. TARGETING PH DOMAIN PROTEINS FOR CANCER THERAPY by Zhi Tan Approval page APPROVED: _____________________________________________ Advisory Professor, Shuxing Zhang, Ph.D. _____________________________________________ -

Delineation of Key Regulatory Elements Identifies Points Of
DELINEATION OF KEY REGULATORY ELEMENTS IDENTIFIES POINTS OF VULNERABILITY IN THE MITOGEN-ACTIVATED SIGNALING NETWORK SUPPLEMENTARY MATERIALS List of contents Supplementary Figures with legends 1. Figure S1: Distribution of primary siRNA screen data, and standardization of assay procedure. 2. Figure S2: Scatter plot of screen data. 3. Figure S3: Functional relevance of the identified targets and Calculation of residence time from PDT and cell cycle distribution. 4. Figure S4: FACS profiles for ABL1 and AKT1. Table for data in Figure 5B. 5. Figure S5: Venn diagram showing the results of the comparative analysis of other screen results 6. Figure S6: Dose response profiles for the AKT1 + ABL1 inhibitor combination for CH1, list of the 14 cell lines and their description, effect of ABL1+AKT1 inhibitor combination on increase in apoptotic cells and G1 arrest in 14 cell lines, effects of CHEK1 inhibitor on combination C1,C2 on 4 cell lines. Supplementary Tables 1. Table S1: siRNA screen results for targeted kinases and phosphatases. 2. Table S2: Gene expression status of the validated hits. 3. Table S3: Role played by identified RNAi hits in regulation of cell cycle, the effect on PDTs along with phase-specific RTs. 4. Table S4: List of molecules classified as cell cycle targets. 5. Table S5: High confidence network used for graph theory analysis. 6. Table S6: Occurrences of nodes in shortest path networks. 7. Table S7: Network file used as SNAVI background. 8. Table S8: Classification of nodes present in modules according to specificity. Legends for tables Supplementary Experimental Procedures References Figure S1 A 450 400 G1 S 350 G2 300 250 200 150 100 50 Distribution of molecules Distribution 0 -6-4-20246 Z-score 350 200 400 G1 S 300 G2 150 300 250 200 100 200 150 100 50 100 Distribution of molecules 50 0 0 0 -4 -2 0 2 4 -4-20246 -4-20246 Z-score B PLK1 GAPDH PLCg BTK PLCg CDC2A PLCg CHEK1 PLCg MET Distribution profiles of complete primary screen and western blots showing knockdown efficiency. -
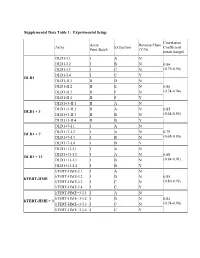
Table 3: Average Gene Expression Profiles by Chromosome
Supplemental Data Table 1: Experimental Setup Correlation Array Reverse Fluor Array Extraction Coefficient Print Batch (Y/N) mean (range) DLD1-I.1 I A N DLD1-I.2 I B N 0.86 DLD1-I.3 I C N (0.79-0.90) DLD1-I.4 I C Y DLD1 DLD1-II.1 II D N DLD1-II.2 II E N 0.86 DLD1-II.3 II F N (0.74-0.94) DLD1-II.4 II F Y DLD1+3-II.1 II A N DLD1+3-II.2 II A N 0.85 DLD1 + 3 DLD1+3-II.3 II B N (0.64-0.95) DLD1+3-II.4 II B Y DLD1+7-I.1 I A N DLD1+7-I.2 I A N 0.79 DLD1 + 7 DLD1+7-I.3 I B N (0.68-0.90) DLD1+7-I.4 I B Y DLD1+13-I.1 I A N DLD1+13-I.2 I A N 0.88 DLD1 + 13 DLD1+13-I.3 I B N (0.84-0.91) DLD1+13-I.4 I B Y hTERT-HME-I.1 I A N hTERT-HME-I.2 I B N 0.85 hTERT-HME hTERT-HME-I.3 I C N (0.80-0.92) hTERT-HME-I.4 I C Y hTERT-HME+3-I.1 I A N hTERT-HME+3-I.2 I B N 0.84 hTERT-HME + 3 hTERT-HME+3-I.3 I C N (0.74-0.90) hTERT-HME+3-I.4 I C Y Supplemental Data Table 2: Average gene expression profiles by chromosome arm DLD1 hTERT-HME Ratio.7 Ratio.1 Ratio.3 Ratio.3 Chrom. -

Anti-ASAP2 (Aa 5-194) Polyclonal Antibody (DPABH-00543) This Product Is for Research Use Only and Is Not Intended for Diagnostic Use
Anti-ASAP2 (aa 5-194) polyclonal antibody (DPABH-00543) This product is for research use only and is not intended for diagnostic use. PRODUCT INFORMATION Antigen Description Activates the small GTPases ARF1, ARF5 and ARF6. Regulates the formation of post-Golgi vesicles and modulates constitutive secretion. Modulates phagocytosis mediated by Fc gamma receptor and ARF6. Modulates PXN recruitment to focal contacts and cell migration. Immunogen Synthetic peptide, corresponding to a region within N terminal amino acids 5-194 of Human PAG3 (UniProt ID: O43150-2). Isotype IgG Source/Host Rabbit Species Reactivity Mouse, Human Purification Immunogen affinity purified Conjugate Unconjugated Applications WB Format Liquid Size 100 μl Buffer pH: 7.00; Constituents: 0.75% Glycine, 1.21% Tris, 20% Glycerol Preservative None Storage Shipped at 4°C. Upon delivery aliquot and store at -20°C or -80°C. Avoid repeated freeze / thaw cycles. GENE INFORMATION Gene Name ASAP2 ArfGAP with SH3 domain, ankyrin repeat and PH domain 3 [ Homo sapiens ] Official Symbol ASAP2 Synonyms ASAP2; ArfGAP with SH3 domain, ankyrin repeat and PH domain 2; PAP; PAG3; AMAP2; DDEF2; SHAG1; CENTB3; Pap-alpha; arf-GAP with SH3 domain, ANK repeat and PH domain- 45-1 Ramsey Road, Shirley, NY 11967, USA Email: [email protected] Tel: 1-631-624-4882 Fax: 1-631-938-8221 1 © Creative Diagnostics All Rights Reserved containing protein 2; centaurin, beta 3; PYK2 C terminus-associated protein; pyk2 C-terminus- associated protein; development and differentiation enhancing factor 2; development and differentiation-enhancing factor 2; paxillin-associated protein with ARF GAP activity 3; Entrez Gene ID 8853 Protein Refseq NP_001128663.1 UniProt ID O43150 Pathway Arf6 trafficking events; Endocytosis; Fc gamma R-mediated phagocytosis; Function ARF GTPase activator activity; enzyme activator activity; protein binding; zinc ion binding 45-1 Ramsey Road, Shirley, NY 11967, USA Email: [email protected] Tel: 1-631-624-4882 Fax: 1-631-938-8221 2 © Creative Diagnostics All Rights Reserved. -

Frequent Loss-Of-Heterozygosity in CRISPR-Cas9–Edited Early Human Embryos
Frequent loss-of-heterozygosity in CRISPR-Cas9–edited COLLOQUIUM PAPER early human embryos Gregorio Alanis-Lobatoa, Jasmin Zohrenb, Afshan McCarthya, Norah M. E. Fogartya,c, Nada Kubikovad,e, Emily Hardmana, Maria Grecof, Dagan Wellsd,g, James M. A. Turnerb, and Kathy K. Niakana,h,1 aHuman Embryo and Stem Cell Laboratory, The Francis Crick Institute, NW1 1AT London, United Kingdom; bSex Chromosome Biology Laboratory, The Francis Crick Institute, NW1 1AT London, United Kingdom; cCentre for Stem Cells and Regenerative Medicine, Guy’s Campus, King’s College London, SE1 9RT London, United Kingdom; dNuffield Department of Women’s and Reproductive Health, John Radcliffe Hospital, University of Oxford, OX3 9DU Oxford, United Kingdom; eJesus College, University of Oxford, OX1 3DW Oxford, United Kingdom; fAncient Genomics Laboratory, The Francis Crick Institute, NW1 1AT London, United Kingdom; gJuno Genetics, OX4 4GE Oxford, United Kingdom; and hThe Centre for Trophoblast Research, Department of Physiology, Development and Neuroscience, University of Cambridge, CB2 3EG Cambridge, United Kingdom Edited by Barbara J. Meyer, University of California, Berkeley, CA, and approved October 31, 2020 (received for review June 5, 2020) CRISPR-Cas9 genome editing is a promising technique for clinical homozygous WT embryos in both cases was not associated with applications, such as the correction of disease-associated alleles in use of the provided repair template for gene correction. Instead, somatic cells. The use of this approach has also been discussed in the authors suggest that in edited embryos the WT maternal the context of heritable editing of the human germ line. However, allele served as a template for the high-fidelity homology di- studies assessing gene correction in early human embryos report rected repair (HDR) pathway to repair the double-strand lesion low efficiency of mutation repair, high rates of mosaicism, and the caused by the Cas9 protein in the paternal allele (8). -

Detection of Copy Number Alterations in Metastatic Melanoma by a DNA
Human Cancer Biology Detection of Copy Number Alterations in Metastatic Melanoma by aDNAFluorescenceIn situ Hybridization Probe Panel and Array Comparative Genomic Hybridization: A Southwest Oncology Group Study (S9431) Stephen R. Moore,1Diane L. Persons,2 Jeffrey A. Sosman,3 Dolores Bobadilla,1Victoria Bedell,1 David D. Smith,1Sandra R.Wolman,4 Ralph J. Tuthill,5 Jim Moon,6 Vernon K. Sondak,7 and Marilyn L. Slovak1 Abstract Purpose: Gene copy number alteration (CNA) is common in malignant melanoma and is associated with tumor development and progression. The concordance between molecular cytogenetic techniques used to determine CNA has not been evaluated on a large set of loci in malignant melanoma. Experimental Design: A panel of 16 locus-specific fluorescence in situ hybridization (FISH) probes located on eight chromosomes was used to identify CNA in touch preparations of frozen tissue samples from19 patients with metastaticmelanoma (SWOG-9431). A subset ( n =11)was analyzed using bacterial artificial chromosome (BAC) array comparative genomic hybridization (aCGH) of DNA isolated directly from touch-preparation slides. Results: By FISH, most samples showed loss near or at WISP3 /6p21, CCND3/6q22, and CDKN2A /9p21 (>75% of samples tested). More than one third of CDKN2A /9p21losses were biallelic. Gains of NEDD9/6p24, MET/7q31, and MYC/8q24 were common (57%, 47%, and 41%, respectively) and CNA events involving 9p21/7p12.3 and MET were frequently coincident, suggesting gain of the whole chromosome 7. Changes were confirmed by aCGH, which also uncovered many discreet regions of change, larger than a single BAC. Overlapping segments observed in >45% of samples included many of the loci analyzed in the FISH study, in addition to other WNT pathway members, and genes associated withTP53 pathways and DNA damage response, repair, and stability. -

DDEF2 CRISPR/Cas9 KO Plasmid (H): Sc-403676
SANTA CRUZ BIOTECHNOLOGY, INC. DDEF2 CRISPR/Cas9 KO Plasmid (h): sc-403676 BACKGROUND APPLICATIONS The Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR) and DDEF2 CRISPR/Cas9 KO Plasmid (h) is recommended for the disruption of CRISPR-associated protein (Cas9) system is an adaptive immune response gene expression in human cells. defense mechanism used by archea and bacteria for the degradation of foreign genetic material (4,6). This mechanism can be repurposed for other 20 nt non-coding RNA sequence: guides Cas9 functions, including genomic engineering for mammalian systems, such as to a specific target location in the genomic DNA gene knockout (KO) (1,2,3,5). CRISPR/Cas9 KO Plasmid products enable the U6 promoter: drives gRNA scaffold: helps Cas9 identification and cleavage of specific genes by utilizing guide RNA (gRNA) expression of gRNA bind to target DNA sequences derived from the Genome-scale CRISPR Knock-Out (GeCKO) v2 library developed in the Zhang Laboratory at the Broad Institute (3,5). Termination signal Green Fluorescent Protein: to visually REFERENCES verify transfection CRISPR/Cas9 Knockout Plasmid CBh (chicken β-Actin 1. Cong, L., et al. 2013. Multiplex genome engineering using CRISPR/Cas hybrid) promoter: drives systems. Science 339: 819-823. 2A peptide: expression of Cas9 allows production of both Cas9 and GFP from the 2. Mali, P., et al. 2013. RNA-guided human genome engineering via Cas9. same CBh promoter Science 339: 823-826. Nuclear localization signal 3. Ran, F.A., et al. 2013. Genome engineering using the CRISPR-Cas9 system. Nuclear localization signal SpCas9 ribonuclease Nat. Protoc. 8: 2281-2308. -

MYBL1 Knockdown in a Triple Negative Breast Cancer Line: Evidence of Down-Regulation of MYBL2, TCF19 and Kif18b Expression
Open Access Austin Journal of Cancer and Clinical Research Research Article MYBL1 Knockdown in a Triple Negative Breast Cancer Line: Evidence of Down-Regulation of MYBL2, TCF19 and KIF18b Expression Player A*, Abraham N, Abdulrahman N, Nsende E, Cunningham S and Rogers S Abstract Department of Biology, Texas Southern University Purpose: The MYBL1 gene is a strong transcriptional activator, associated Houston TX, USA with cell cycle signaling and differentiation. Data show the gene is over- *Corresponding author: Audrey Player, Department expressed in triple negative breast cancers. Considering the possibility that of Biology, Texas Southern University, Houston Texas, MYBL1 might be involved in events associated with the pathogenesis of these USA cancers, we sought to identify genes associated with MYBL1 expression in triple negative breast cancer. Received: May 21, 2021; Accepted: June 12, 2021; Published: June 19, 2021 Methods: shRNA lentiviral knockdown was used to down-regulate the MYBL1 gene. Microarray analyses were used to identify genes either directly or indirectly affected by targeting MYBL1 knockdown. Data analyses was performed utilizing Affymetrix TAC 4.0, Chip X transcription factor analyses, Target Scan miRNA analyses, and STRING analyses was used to determine protein: protein interaction and pathway analyses. Web Gestalt and Gene Ontology were used to determine pathway and gene-set enrichments. Publicly available patient and cell line datasets were retrieved and processed using resources available in Gene Expression Omnibus and Oncomine. The polymerase chain reaction and western analyses were used to determine transcript and protein levels, respectively. Results: Knockdown of MYBL1 in a triple negative breast cell line led to down-regulation of MYBL2, TCF19, KIF18b along with an enrichment of cell cycle signaling genes. -

Interaction Networks of Lithium and Valproate Molecular Targets Reveal a Striking Enrichment of Apoptosis Functional Clusters and Neurotrophin Signaling
The Pharmacogenomics Journal (2012) 12, 328–341 & 2012 Macmillan Publishers Limited. All rights reserved 1470-269X/12 www.nature.com/tpj ORIGINAL ARTICLE Interaction networks of lithium and valproate molecular targets reveal a striking enrichment of apoptosis functional clusters and neurotrophin signaling A Gupta1, TG Schulze2, The overall neurobiological mechanisms by which lithium and valproate 3 1 stabilize mood in bipolar disorder patients have yet to be fully defined. The V Nagarajan , N Akula , therapeutic efficacy and dissimilar chemical structures of these medications 1 1 W Corona , X-y Jiang , suggest that they perturb both shared and disparate cellular processes. To N Hunter1, FJ McMahon1 and investigate key pathways and functional clusters involved in the global action SD Detera-Wadleigh1 of lithium and valproate, we generated interaction networks formed by well- supported drug targets. Striking functional similarities emerged. Intersecting 1Human Genetics Branch, National Institute of nodes in lithium and valproate networks highlighted a strong enrichment Mental Health, National Institutes of Health, of apoptosis clusters and neurotrophin signaling. Other enriched pathways 2 Bethesda, MD, USA; Department of Psychiatry included MAPK, ErbB, insulin, VEGF, Wnt and long-term potentiation and Psychotherapy, University Medical Center, Georg-August-Universita¨t, Go¨ttingen, Germany indicating a widespread effect of both drugs on diverse signaling systems. and 3Bioinformatics and Computational MAPK1/3 and AKT1/2 were the most preponderant nodes across pathways Biosciences Branch, Office of Cyber Infrastructure suggesting a central role in mediating pathway interactions. The conver- and Computational Biology, National Institute of gence of biological responses unveils a functional signature for lithium and Allergy and Infectious Diseases, National Institutes of Health, Bethesda, MD, USA valproate that could be key modulators of their therapeutic efficacy.