Number 7 July 2012
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Integrating Single-Step GWAS and Bipartite Networks Reconstruction Provides Novel Insights Into Yearling Weight and Carcass Traits in Hanwoo Beef Cattle
animals Article Integrating Single-Step GWAS and Bipartite Networks Reconstruction Provides Novel Insights into Yearling Weight and Carcass Traits in Hanwoo Beef Cattle Masoumeh Naserkheil 1 , Abolfazl Bahrami 1 , Deukhwan Lee 2,* and Hossein Mehrban 3 1 Department of Animal Science, University College of Agriculture and Natural Resources, University of Tehran, Karaj 77871-31587, Iran; [email protected] (M.N.); [email protected] (A.B.) 2 Department of Animal Life and Environment Sciences, Hankyong National University, Jungang-ro 327, Anseong-si, Gyeonggi-do 17579, Korea 3 Department of Animal Science, Shahrekord University, Shahrekord 88186-34141, Iran; [email protected] * Correspondence: [email protected]; Tel.: +82-31-670-5091 Received: 25 August 2020; Accepted: 6 October 2020; Published: 9 October 2020 Simple Summary: Hanwoo is an indigenous cattle breed in Korea and popular for meat production owing to its rapid growth and high-quality meat. Its yearling weight and carcass traits (backfat thickness, carcass weight, eye muscle area, and marbling score) are economically important for the selection of young and proven bulls. In recent decades, the advent of high throughput genotyping technologies has made it possible to perform genome-wide association studies (GWAS) for the detection of genomic regions associated with traits of economic interest in different species. In this study, we conducted a weighted single-step genome-wide association study which combines all genotypes, phenotypes and pedigree data in one step (ssGBLUP). It allows for the use of all SNPs simultaneously along with all phenotypes from genotyped and ungenotyped animals. Our results revealed 33 relevant genomic regions related to the traits of interest. -

Genetic and Genomic Analysis of Hyperlipidemia, Obesity and Diabetes Using (C57BL/6J × TALLYHO/Jngj) F2 Mice
University of Tennessee, Knoxville TRACE: Tennessee Research and Creative Exchange Nutrition Publications and Other Works Nutrition 12-19-2010 Genetic and genomic analysis of hyperlipidemia, obesity and diabetes using (C57BL/6J × TALLYHO/JngJ) F2 mice Taryn P. Stewart Marshall University Hyoung Y. Kim University of Tennessee - Knoxville, [email protected] Arnold M. Saxton University of Tennessee - Knoxville, [email protected] Jung H. Kim Marshall University Follow this and additional works at: https://trace.tennessee.edu/utk_nutrpubs Part of the Animal Sciences Commons, and the Nutrition Commons Recommended Citation BMC Genomics 2010, 11:713 doi:10.1186/1471-2164-11-713 This Article is brought to you for free and open access by the Nutrition at TRACE: Tennessee Research and Creative Exchange. It has been accepted for inclusion in Nutrition Publications and Other Works by an authorized administrator of TRACE: Tennessee Research and Creative Exchange. For more information, please contact [email protected]. Stewart et al. BMC Genomics 2010, 11:713 http://www.biomedcentral.com/1471-2164/11/713 RESEARCH ARTICLE Open Access Genetic and genomic analysis of hyperlipidemia, obesity and diabetes using (C57BL/6J × TALLYHO/JngJ) F2 mice Taryn P Stewart1, Hyoung Yon Kim2, Arnold M Saxton3, Jung Han Kim1* Abstract Background: Type 2 diabetes (T2D) is the most common form of diabetes in humans and is closely associated with dyslipidemia and obesity that magnifies the mortality and morbidity related to T2D. The genetic contribution to human T2D and related metabolic disorders is evident, and mostly follows polygenic inheritance. The TALLYHO/ JngJ (TH) mice are a polygenic model for T2D characterized by obesity, hyperinsulinemia, impaired glucose uptake and tolerance, hyperlipidemia, and hyperglycemia. -

Targeting PH Domain Proteins for Cancer Therapy
The Texas Medical Center Library DigitalCommons@TMC The University of Texas MD Anderson Cancer Center UTHealth Graduate School of The University of Texas MD Anderson Cancer Biomedical Sciences Dissertations and Theses Center UTHealth Graduate School of (Open Access) Biomedical Sciences 12-2018 Targeting PH domain proteins for cancer therapy Zhi Tan Follow this and additional works at: https://digitalcommons.library.tmc.edu/utgsbs_dissertations Part of the Bioinformatics Commons, Medicinal Chemistry and Pharmaceutics Commons, Neoplasms Commons, and the Pharmacology Commons Recommended Citation Tan, Zhi, "Targeting PH domain proteins for cancer therapy" (2018). The University of Texas MD Anderson Cancer Center UTHealth Graduate School of Biomedical Sciences Dissertations and Theses (Open Access). 910. https://digitalcommons.library.tmc.edu/utgsbs_dissertations/910 This Dissertation (PhD) is brought to you for free and open access by the The University of Texas MD Anderson Cancer Center UTHealth Graduate School of Biomedical Sciences at DigitalCommons@TMC. It has been accepted for inclusion in The University of Texas MD Anderson Cancer Center UTHealth Graduate School of Biomedical Sciences Dissertations and Theses (Open Access) by an authorized administrator of DigitalCommons@TMC. For more information, please contact [email protected]. TARGETING PH DOMAIN PROTEINS FOR CANCER THERAPY by Zhi Tan Approval page APPROVED: _____________________________________________ Advisory Professor, Shuxing Zhang, Ph.D. _____________________________________________ -

Systemic Surfaceome Profiling Identifies Target Antigens for Immune-Based Therapy in Subtypes of Advanced Prostate Cancer
Systemic surfaceome profiling identifies target antigens for immune-based therapy in subtypes of advanced prostate cancer John K. Leea,b,c, Nathanael J. Bangayand, Timothy Chaie, Bryan A. Smithf, Tiffany E. Parivaf, Sangwon Yung, Ajay Vashishth, Qingfu Zhangi,j, Jung Wook Parkf, Eva Coreyk, Jiaoti Huangi, Thomas G. Graeberc,d,l,m, James Wohlschlegelh, and Owen N. Wittec,d,f,n,o,1 aDivision of Hematology and Oncology, Department of Medicine, University of California, Los Angeles, CA 90095; bInstitute of Urologic Oncology, Department of Urology, University of California, Los Angeles, CA 90095; cJonsson Comprehensive Cancer Center, University of California, Los Angeles, CA 90095; dDepartment of Molecular and Medical Pharmacology, University of California, Los Angeles, CA 90095; eStanford University School of Medicine, Palo Alto, CA 94305; fDepartment of Microbiology, Immunology, and Medical Genetics, University of California, Los Angeles, CA 90095; gYale School of Medicine, New Haven, CT 06510; hDepartment of Biological Chemistry, University of California, Los Angeles, CA 90095; iDepartment of Pathology, Duke University School of Medicine, Durham, NC 27708; jDepartment of Pathology, China Medical University, 110001 Shenyang, People’s Republic of China; kDepartment of Urology, University of Washington School of Medicine, Seattle, WA 98195; lCrump Institute for Molecular Imaging, University of California, Los Angeles, CA 90095; mUCLA Metabolomics Center, University of California, Los Angeles, CA 900095; nParker Institute for Cancer Immunotherapy, -

Delineation of Key Regulatory Elements Identifies Points Of
DELINEATION OF KEY REGULATORY ELEMENTS IDENTIFIES POINTS OF VULNERABILITY IN THE MITOGEN-ACTIVATED SIGNALING NETWORK SUPPLEMENTARY MATERIALS List of contents Supplementary Figures with legends 1. Figure S1: Distribution of primary siRNA screen data, and standardization of assay procedure. 2. Figure S2: Scatter plot of screen data. 3. Figure S3: Functional relevance of the identified targets and Calculation of residence time from PDT and cell cycle distribution. 4. Figure S4: FACS profiles for ABL1 and AKT1. Table for data in Figure 5B. 5. Figure S5: Venn diagram showing the results of the comparative analysis of other screen results 6. Figure S6: Dose response profiles for the AKT1 + ABL1 inhibitor combination for CH1, list of the 14 cell lines and their description, effect of ABL1+AKT1 inhibitor combination on increase in apoptotic cells and G1 arrest in 14 cell lines, effects of CHEK1 inhibitor on combination C1,C2 on 4 cell lines. Supplementary Tables 1. Table S1: siRNA screen results for targeted kinases and phosphatases. 2. Table S2: Gene expression status of the validated hits. 3. Table S3: Role played by identified RNAi hits in regulation of cell cycle, the effect on PDTs along with phase-specific RTs. 4. Table S4: List of molecules classified as cell cycle targets. 5. Table S5: High confidence network used for graph theory analysis. 6. Table S6: Occurrences of nodes in shortest path networks. 7. Table S7: Network file used as SNAVI background. 8. Table S8: Classification of nodes present in modules according to specificity. Legends for tables Supplementary Experimental Procedures References Figure S1 A 450 400 G1 S 350 G2 300 250 200 150 100 50 Distribution of molecules Distribution 0 -6-4-20246 Z-score 350 200 400 G1 S 300 G2 150 300 250 200 100 200 150 100 50 100 Distribution of molecules 50 0 0 0 -4 -2 0 2 4 -4-20246 -4-20246 Z-score B PLK1 GAPDH PLCg BTK PLCg CDC2A PLCg CHEK1 PLCg MET Distribution profiles of complete primary screen and western blots showing knockdown efficiency. -

The Proteomic Landscape of Resting and Activated CD4+ T Cells Reveal Insights Into Cell Differentiation and Function
International Journal of Molecular Sciences Article The Proteomic Landscape of Resting and Activated CD4+ T Cells Reveal Insights into Cell Differentiation and Function Yashwanth Subbannayya 1 , Markus Haug 1, Sneha M. Pinto 1, Varshasnata Mohanty 2, Hany Zakaria Meås 1, Trude Helen Flo 1, T.S. Keshava Prasad 2 and Richard K. Kandasamy 1,* 1 Centre of Molecular Inflammation Research (CEMIR), Department of Clinical and Molecular Medicine (IKOM), Norwegian University of Science and Technology, 7491 Trondheim, Norway; [email protected] (Y.S.); [email protected] (M.H.); [email protected] (S.M.P.); [email protected] (H.Z.M.); trude.fl[email protected] (T.H.F.) 2 Center for Systems Biology and Molecular Medicine, Yenepoya (Deemed to be University), Mangalore 575018, India; [email protected] (V.M.); [email protected] (T.S.K.P.) * Correspondence: [email protected] Abstract: CD4+ T cells (T helper cells) are cytokine-producing adaptive immune cells that activate or regulate the responses of various immune cells. The activation and functional status of CD4+ T cells is important for adequate responses to pathogen infections but has also been associated with auto-immune disorders and survival in several cancers. In the current study, we carried out a label-free high-resolution FTMS-based proteomic profiling of resting and T cell receptor-activated (72 h) primary human CD4+ T cells from peripheral blood of healthy donors as well as SUP-T1 cells. We identified 5237 proteins, of which significant alterations in the levels of 1119 proteins were observed between resting and activated CD4+ T cells. -
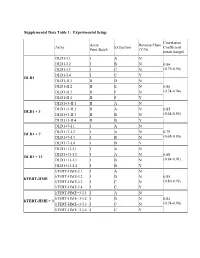
Table 3: Average Gene Expression Profiles by Chromosome
Supplemental Data Table 1: Experimental Setup Correlation Array Reverse Fluor Array Extraction Coefficient Print Batch (Y/N) mean (range) DLD1-I.1 I A N DLD1-I.2 I B N 0.86 DLD1-I.3 I C N (0.79-0.90) DLD1-I.4 I C Y DLD1 DLD1-II.1 II D N DLD1-II.2 II E N 0.86 DLD1-II.3 II F N (0.74-0.94) DLD1-II.4 II F Y DLD1+3-II.1 II A N DLD1+3-II.2 II A N 0.85 DLD1 + 3 DLD1+3-II.3 II B N (0.64-0.95) DLD1+3-II.4 II B Y DLD1+7-I.1 I A N DLD1+7-I.2 I A N 0.79 DLD1 + 7 DLD1+7-I.3 I B N (0.68-0.90) DLD1+7-I.4 I B Y DLD1+13-I.1 I A N DLD1+13-I.2 I A N 0.88 DLD1 + 13 DLD1+13-I.3 I B N (0.84-0.91) DLD1+13-I.4 I B Y hTERT-HME-I.1 I A N hTERT-HME-I.2 I B N 0.85 hTERT-HME hTERT-HME-I.3 I C N (0.80-0.92) hTERT-HME-I.4 I C Y hTERT-HME+3-I.1 I A N hTERT-HME+3-I.2 I B N 0.84 hTERT-HME + 3 hTERT-HME+3-I.3 I C N (0.74-0.90) hTERT-HME+3-I.4 I C Y Supplemental Data Table 2: Average gene expression profiles by chromosome arm DLD1 hTERT-HME Ratio.7 Ratio.1 Ratio.3 Ratio.3 Chrom. -

Differential Expression Profile Prioritization of Positional Candidate Glaucoma Genes the GLC1C Locus
LABORATORY SCIENCES Differential Expression Profile Prioritization of Positional Candidate Glaucoma Genes The GLC1C Locus Frank W. Rozsa, PhD; Kathleen M. Scott, BS; Hemant Pawar, PhD; John R. Samples, MD; Mary K. Wirtz, PhD; Julia E. Richards, PhD Objectives: To develop and apply a model for priori- est because of moderate expression and changes in tization of candidate glaucoma genes. expression. Transcription factor ZBTB38 emerges as an interesting candidate gene because of the overall expres- Methods: This Affymetrix GeneChip (Affymetrix, Santa sion level, differential expression, and function. Clara, Calif) study of gene expression in primary cul- ture human trabecular meshwork cells uses a positional Conclusions: Only1geneintheGLC1C interval fits our differential expression profile model for prioritization of model for differential expression under multiple glau- candidate genes within the GLC1C genetic inclusion in- coma risk conditions. The use of multiple prioritization terval. models resulted in filtering 7 candidate genes of higher interest out of the 41 known genes in the region. Results: Sixteen genes were expressed under all condi- tions within the GLC1C interval. TMEM22 was the only Clinical Relevance: This study identified a small sub- gene within the interval with differential expression in set of genes that are most likely to harbor mutations that the same direction under both conditions tested. Two cause glaucoma linked to GLC1C. genes, ATP1B3 and COPB2, are of interest in the con- text of a protein-misfolding model for candidate selec- tion. SLC25A36, PCCB, and FNDC6 are of lesser inter- Arch Ophthalmol. 2007;125:117-127 IGH PREVALENCE AND PO- identification of additional GLC1C fami- tential for severe out- lies7,18-20 who provide optimal samples for come combine to make screening candidate genes for muta- adult-onset primary tions.7,18,20 The existence of 2 distinct open-angle glaucoma GLC1C haplotypes suggests that muta- (POAG) a significant public health prob- tions will not be limited to rare descen- H1 lem. -

FXYD2 Monoclonal Antibody (M01), Human Gene Nomenclature for the Family Is Clone 1C3-B3 FXYD-Domain Containing Ion Transport Regulator
FXYD2 monoclonal antibody (M01), human gene nomenclature for the family is clone 1C3-B3 FXYD-domain containing ion transport regulator. Mouse FXYD5 has been termed RIC (Related to Ion Channel). Catalog Number: H00000486-M01 FXYD2, also known as the gamma subunit of the Na,K-ATPase, regulates the properties of that enzyme. Regulatory Status: For research use only (RUO) FXYD1 (phospholemman), FXYD2 (gamma), FXYD3 (MAT-8), FXYD4 (CHIF), and FXYD5 (RIC) have been Product Description: Mouse monoclonal antibody shown to induce channel activity in experimental raised against a full length recombinant FXYD2. expression systems. Transmembrane topology has been established for two family members (FXYD1 and Clone Name: 1C3-B3 FXYD2), with the N-terminus extracellular and the C-terminus on the cytoplasmic side of the membrane. Immunogen: FXYD2 (AAH05302.1, 1 a.a. ~ 64 a.a) The Type III integral membrane protein encoded by this full-length recombinant protein with GST tag. MW of the gene is the gamma subunit of the Na,K-ATPase present GST tag alone is 26 KDa. on the plasma membrane. Although the Na,K-ATPase does not depend on the gamma subunit to be functional, Sequence: it is thought that the gamma subunit modulates the MDRWYLGGSPKGDVDPFYYDYETVRNGGLIFAGLAFI enzyme's activity by inducing ion channel activity. VGLLILLSRRFRCGGNKKRRQINEDEP Mutations in this gene have been associated with renal Host: Mouse hypomagnesaemia. Alternatively spliced transcript variants encoding different isoforms have been found for Reactivity: Human this gene. [provided by RefSeq] Applications: ELISA, IHC-P, S-ELISA, WB-Ce, WB-Re References: (See our web site product page for detailed applications 1. -

Single-Cell Transcriptomes Reveal a Complex Cellular Landscape in the Middle Ear and Differential Capacities for Acute Response to Infection
fgene-11-00358 April 9, 2020 Time: 15:55 # 1 ORIGINAL RESEARCH published: 15 April 2020 doi: 10.3389/fgene.2020.00358 Single-Cell Transcriptomes Reveal a Complex Cellular Landscape in the Middle Ear and Differential Capacities for Acute Response to Infection Allen F. Ryan1*, Chanond A. Nasamran2, Kwang Pak1, Clara Draf1, Kathleen M. Fisch2, Nicholas Webster3 and Arwa Kurabi1 1 Departments of Surgery/Otolaryngology, UC San Diego School of Medicine, VA Medical Center, La Jolla, CA, United States, 2 Medicine/Center for Computational Biology & Bioinformatics, UC San Diego School of Medicine, VA Medical Center, La Jolla, CA, United States, 3 Medicine/Endocrinology, UC San Diego School of Medicine, VA Medical Center, La Jolla, CA, United States Single-cell transcriptomics was used to profile cells of the normal murine middle ear. Clustering analysis of 6770 transcriptomes identified 17 cell clusters corresponding to distinct cell types: five epithelial, three stromal, three lymphocyte, two monocyte, Edited by: two endothelial, one pericyte and one melanocyte cluster. Within some clusters, Amélie Bonnefond, Institut National de la Santé et de la cell subtypes were identified. While many corresponded to those cell types known Recherche Médicale (INSERM), from prior studies, several novel types or subtypes were noted. The results indicate France unexpected cellular diversity within the resting middle ear mucosa. The resolution of Reviewed by: Fabien Delahaye, uncomplicated, acute, otitis media is too rapid for cognate immunity to play a major Institut Pasteur de Lille, France role. Thus innate immunity is likely responsible for normal recovery from middle ear Nelson L. S. Tang, infection. The need for rapid response to pathogens suggests that innate immune The Chinese University of Hong Kong, China genes may be constitutively expressed by middle ear cells. -

Anti-ASAP2 (Aa 5-194) Polyclonal Antibody (DPABH-00543) This Product Is for Research Use Only and Is Not Intended for Diagnostic Use
Anti-ASAP2 (aa 5-194) polyclonal antibody (DPABH-00543) This product is for research use only and is not intended for diagnostic use. PRODUCT INFORMATION Antigen Description Activates the small GTPases ARF1, ARF5 and ARF6. Regulates the formation of post-Golgi vesicles and modulates constitutive secretion. Modulates phagocytosis mediated by Fc gamma receptor and ARF6. Modulates PXN recruitment to focal contacts and cell migration. Immunogen Synthetic peptide, corresponding to a region within N terminal amino acids 5-194 of Human PAG3 (UniProt ID: O43150-2). Isotype IgG Source/Host Rabbit Species Reactivity Mouse, Human Purification Immunogen affinity purified Conjugate Unconjugated Applications WB Format Liquid Size 100 μl Buffer pH: 7.00; Constituents: 0.75% Glycine, 1.21% Tris, 20% Glycerol Preservative None Storage Shipped at 4°C. Upon delivery aliquot and store at -20°C or -80°C. Avoid repeated freeze / thaw cycles. GENE INFORMATION Gene Name ASAP2 ArfGAP with SH3 domain, ankyrin repeat and PH domain 3 [ Homo sapiens ] Official Symbol ASAP2 Synonyms ASAP2; ArfGAP with SH3 domain, ankyrin repeat and PH domain 2; PAP; PAG3; AMAP2; DDEF2; SHAG1; CENTB3; Pap-alpha; arf-GAP with SH3 domain, ANK repeat and PH domain- 45-1 Ramsey Road, Shirley, NY 11967, USA Email: [email protected] Tel: 1-631-624-4882 Fax: 1-631-938-8221 1 © Creative Diagnostics All Rights Reserved containing protein 2; centaurin, beta 3; PYK2 C terminus-associated protein; pyk2 C-terminus- associated protein; development and differentiation enhancing factor 2; development and differentiation-enhancing factor 2; paxillin-associated protein with ARF GAP activity 3; Entrez Gene ID 8853 Protein Refseq NP_001128663.1 UniProt ID O43150 Pathway Arf6 trafficking events; Endocytosis; Fc gamma R-mediated phagocytosis; Function ARF GTPase activator activity; enzyme activator activity; protein binding; zinc ion binding 45-1 Ramsey Road, Shirley, NY 11967, USA Email: [email protected] Tel: 1-631-624-4882 Fax: 1-631-938-8221 2 © Creative Diagnostics All Rights Reserved.