An Analysis About Heterogeneity Among Cancers Based on the DNA Methylation Patterns Yang Liu1, Yue Gu2,Musu1, Hui Liu2, Shumei Zhang3* and Yan Zhang1*
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

Genome-Wide Association Study Identifies Loci for Arterial Stiffness
www.nature.com/scientificreports OPEN Genome-wide association study identifes loci for arterial stifness index in 127,121 UK Biobank Received: 1 February 2019 Accepted: 5 June 2019 participants Published: xx xx xxxx Kenneth Fung1, Julia Ramírez 2, Helen R. Warren2,3, Nay Aung 1, Aaron M. Lee1, Evan Tzanis2,3, Stefen E. Petersen 1,3 & Patricia B. Munroe2,3 Arterial stifness index (ASI) is a non-invasive measure of arterial stifness using infra-red fnger sensors (photoplethysmography). It is a well-suited measure for large populations as it is relatively inexpensive to perform, and data can be acquired within seconds. These features raise interest in using ASI as a tool to estimate cardiovascular disease risk as prior work demonstrates increased arterial stifness is associated with elevated systolic blood pressure, and ASI is predictive of cardiovascular disease and mortality. We conducted genome-wide association studies (GWASs) for ASI in 127,121 UK Biobank participants of European-ancestry. Our primary analyses identifed variants at four loci reaching genome-wide signifcance (P < 5 × 10−8): TEX41 (rs1006923; P = 5.3 × 10−12), FOXO1 (rs7331212; P = 2.2 × 10−11), C1orf21 (rs1930290, P = 1.1 × 10−8) and MRVI1 (rs10840457, P = 3.4 × 10−8). Gene- based testing revealed three signifcant genes, the most signifcant gene was COL4A2 (P = 1.41 × 10−8) encoding type IV collagen. Other candidate genes at associated loci were also involved in smooth muscle tone regulation. Our fndings provide new information for understanding the development of arterial stifness. Arterial stifness measures have been reported as independent markers of vascular ageing1,2, hypertension3,4, car- diovascular disease (CVD)5,6 and mortality6,7. -

Supplementary Table 1.Xlsx
SUPPLEMENTARY APPENDIX Chromothripsis is linked to TP53 alteration, cell cycle impairment, and dismal outcome in acute myeloid leukemia with complex karyotype Frank G. Rücker, 1 Anna Dolnik, 1 Tamara J. Blätte, 1 Veronica Teleanu, 1 Aurélie Ernst, 2 Felicitas Thol, 3 Michael Heuser, 3 Arnold Ganser, 3 Hartmut Döhner, 1 Konstanze Döhner *1 and Lars Bullinger *1,4 1Department of Internal Medicine III, University Hospital of Ulm, Albert-Einstein-Allee 23; 2Division of Molecular Genetics, German Cancer Consor - tium (DKTK), German Cancer Research Center (DKFZ), Heidelberg; 3Department of Hematology, Hemostasis, Oncology, and Stem Cell Transplanta - tion, Hannover Medical School and 4Department of Hematology, Oncology and Tumor Immunology, Charité-University Medicine Berlin, Campus Virchow Klinikum, Berlin, Germany *KD and LB contributed equally to this work. Correspondence: [email protected] doi:10.3324/haematol.2017.180497 Supplementary Table 1 Top 200 differentially expressed genes in chromothripsis‐positive (n=13) vs –negative (n=14) CK‐AML (p‐value <.05) Symbol ID LogFC p value PSMB10 5699 ‐1,652 4,81E‐05 ADAMTS6 11174 ‐1,736 1,50E‐04 SLC7A6 9057 ‐1,317 6,80E‐04 VANGL1 81839 2,403 6,94E‐04 ZNF197 10168 1,409 7,30E‐04 TAS2R62P 338399 ‐1,404 9,22E‐04 PLA2G15 23659 ‐1,455 0,001 CBFB 865 ‐1,453 0,001 CNOT8 9337 ‐1,449 0,001 ACD 65057 ‐1,329 0,001 CKLF 51192 ‐1,657 0,002 CMTM3 123920 ‐1,48 0,002 FAM192A 80011 ‐1,408 0,002 SSH3 54961 ‐1,279 0,002 ZNF415 55786 1,425 0,002 BEX2 84707 1,57 0,002 GCNT2 2651 1,732 0,002 PIM1 5292 1,935 -

Qt4vh1p2c4 Nosplash E372185
Copyright 2014 by Janine Micheli-Jazdzewski ii Dedication I would like to dedicate this thesis to Rock, who is not with us anymore, TR, General Jack D. Ripper, and Page. Thank you for sitting with me while I worked for countless hours over the years. iii Acknowledgements I would like to express my special appreciation and thanks to my advisor Dr. Deanna Kroetz, you have been a superb mentor for me. I would like to thank you for encouraging my research and for helping me to grow as a research scientist. Your advice on both research, as well as on my career have been priceless. I would also like to thank my committee members, Dr. Laura Bull, Dr. Steve Hamilton and Dr. John Witte for guiding my research and expanding my knowledge on statistics, genetics and clinical phenotypes. I also want to thank past and present members of my laboratory for their support and help over the years, especially Dr. Mike Baldwin, Dr. Sveta Markova, Dr. Ying Mei Liu and Dr. Leslie Chinn. Thanks are also due to my many collaborators that made this research possible including: Dr. Eric Jorgenson, Dr. David Bangsberg, Dr. Taisei Mushiroda, Dr. Michiaki Kubo, Dr. Yusuke Nakamura, Dr. Jeffrey Martin, Joel Mefford, Dr. Sarah Shutgarts, Dr. Sulggi Lee and Dr. Sook Wah Yee. A special thank you to the RIKEN Center for Genomic Medicine that generously performed the genome-wide genotyping for these projects. Thanks to Dr. Steve Chamow, Dr. Bill Werner, Dr. Montse Carrasco, and Dr. Teresa Chen who started me on the path to becoming a scientist. -
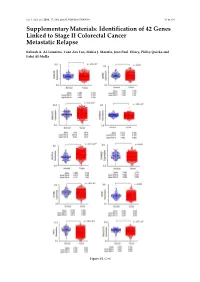
Identification of 42 Genes Linked to Stage II Colorectal Cancer Metastatic Relapse
Int. J. Mol. Sci. 2016, 17, 598; doi:10.3390/ijms17040598 S1 of S16 Supplementary Materials: Identification of 42 Genes Linked to Stage II Colorectal Cancer Metastatic Relapse Rabeah A. Al-Temaimi, Tuan Zea Tan, Makia J. Marafie, Jean Paul Thiery, Philip Quirke and Fahd Al-Mulla Figure S1. Cont. Int. J. Mol. Sci. 2016, 17, 598; doi:10.3390/ijms17040598 S2 of S16 Figure S1. Mean expression levels of fourteen genes of significant association with CRC DFS and OS that are differentially expressed in normal colon compared to CRC tissues. Each dot represents a sample. Table S1. Copy number aberrations associated with poor disease-free survival and metastasis in early stage II CRC as predicted by STAC and SPPS combined methodologies with resident gene symbols. CN stands for copy number, whereas CNV is copy number variation. Region Cytoband % of CNV Count of Region Event Gene Symbols Length Location Overlap Genes chr1:113,025,076–113,199,133 174,057 p13.2 CN Loss 0.0 2 AKR7A2P1, SLC16A1 chr1:141,465,960–141,822,265 356,305 q12–q21.1 CN Gain 95.9 1 SRGAP2B MIR5087, LOC10013000 0, FLJ39739, LOC10028679 3, PPIAL4G, PPIAL4A, NBPF14, chr1:144,911,564–146,242,907 1,331,343 q21.1 CN Gain 99.6 16 NBPF15, NBPF16, PPIAL4E, NBPF16, PPIAL4D, PPIAL4F, LOC645166, LOC388692, FCGR1C chr1:177,209,428–177,226,812 17,384 q25.3 CN Gain 0.0 0 chr1:197,652,888–197,676,831 23,943 q32.1 CN Gain 0.0 1 KIF21B chr1:201,015,278–201,033,308 18,030 q32.1 CN Gain 0.0 1 PLEKHA6 chr1:201,289,154–201,298,247 9093 q32.1 CN Gain 0.0 0 chr1:216,820,186–217,043,421 223,235 q41 CN -

Sex-Specific Transcriptome Differences in Human Adipose
G C A T T A C G G C A T genes Article Sex-Specific Transcriptome Differences in Human Adipose Mesenchymal Stem Cells 1, 2, 3 1,3 Eva Bianconi y, Raffaella Casadei y , Flavia Frabetti , Carlo Ventura , Federica Facchin 1,3,* and Silvia Canaider 1,3 1 National Laboratory of Molecular Biology and Stem Cell Bioengineering of the National Institute of Biostructures and Biosystems (NIBB)—Eldor Lab, at the Innovation Accelerator, CNR, Via Piero Gobetti 101, 40129 Bologna, Italy; [email protected] (E.B.); [email protected] (C.V.); [email protected] (S.C.) 2 Department for Life Quality Studies (QuVi), University of Bologna, Corso D’Augusto 237, 47921 Rimini, Italy; [email protected] 3 Department of Experimental, Diagnostic and Specialty Medicine (DIMES), University of Bologna, Via Massarenti 9, 40138 Bologna, Italy; fl[email protected] * Correspondence: [email protected]; Tel.: +39-051-2094114 These authors contributed equally to this work. y Received: 1 July 2020; Accepted: 6 August 2020; Published: 8 August 2020 Abstract: In humans, sexual dimorphism can manifest in many ways and it is widely studied in several knowledge fields. It is increasing the evidence that also cells differ according to sex, a correlation still little studied and poorly considered when cells are used in scientific research. Specifically, our interest is on the sex-related dimorphism on the human mesenchymal stem cells (hMSCs) transcriptome. A systematic meta-analysis of hMSC microarrays was performed by using the Transcriptome Mapper (TRAM) software. This bioinformatic tool was used to integrate and normalize datasets from multiple sources and allowed us to highlight chromosomal segments and genes differently expressed in hMSCs derived from adipose tissue (hADSCs) of male and female donors. -

WO 2019/068007 Al Figure 2
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization I International Bureau (10) International Publication Number (43) International Publication Date WO 2019/068007 Al 04 April 2019 (04.04.2019) W 1P O PCT (51) International Patent Classification: (72) Inventors; and C12N 15/10 (2006.01) C07K 16/28 (2006.01) (71) Applicants: GROSS, Gideon [EVIL]; IE-1-5 Address C12N 5/10 (2006.0 1) C12Q 1/6809 (20 18.0 1) M.P. Korazim, 1292200 Moshav Almagor (IL). GIBSON, C07K 14/705 (2006.01) A61P 35/00 (2006.01) Will [US/US]; c/o ImmPACT-Bio Ltd., 2 Ilian Ramon St., C07K 14/725 (2006.01) P.O. Box 4044, 7403635 Ness Ziona (TL). DAHARY, Dvir [EilL]; c/o ImmPACT-Bio Ltd., 2 Ilian Ramon St., P.O. (21) International Application Number: Box 4044, 7403635 Ness Ziona (IL). BEIMAN, Merav PCT/US2018/053583 [EilL]; c/o ImmPACT-Bio Ltd., 2 Ilian Ramon St., P.O. (22) International Filing Date: Box 4044, 7403635 Ness Ziona (E.). 28 September 2018 (28.09.2018) (74) Agent: MACDOUGALL, Christina, A. et al; Morgan, (25) Filing Language: English Lewis & Bockius LLP, One Market, Spear Tower, SanFran- cisco, CA 94105 (US). (26) Publication Language: English (81) Designated States (unless otherwise indicated, for every (30) Priority Data: kind of national protection available): AE, AG, AL, AM, 62/564,454 28 September 2017 (28.09.2017) US AO, AT, AU, AZ, BA, BB, BG, BH, BN, BR, BW, BY, BZ, 62/649,429 28 March 2018 (28.03.2018) US CA, CH, CL, CN, CO, CR, CU, CZ, DE, DJ, DK, DM, DO, (71) Applicant: IMMP ACT-BIO LTD. -

Sean Raspet – Molecules
1. Commercial name: Fructaplex© IUPAC Name: 2-(3,3-dimethylcyclohexyl)-2,5,5-trimethyl-1,3-dioxane SMILES: CC1(C)CCCC(C1)C2(C)OCC(C)(C)CO2 Molecular weight: 240.39 g/mol Volume (cubic Angstroems): 258.88 Atoms number (non-hydrogen): 17 miLogP: 4.43 Structure: Biological Properties: Predicted Druglikenessi: GPCR ligand -0.23 Ion channel modulator -0.03 Kinase inhibitor -0.6 Nuclear receptor ligand 0.15 Protease inhibitor -0.28 Enzyme inhibitor 0.15 Commercial name: Fructaplex© IUPAC Name: 2-(3,3-dimethylcyclohexyl)-2,5,5-trimethyl-1,3-dioxane SMILES: CC1(C)CCCC(C1)C2(C)OCC(C)(C)CO2 Predicted Olfactory Receptor Activityii: OR2L13 83.715% OR1G1 82.761% OR10J5 80.569% OR2W1 78.180% OR7A2 77.696% 2. Commercial name: Sylvoxime© IUPAC Name: N-[4-(1-ethoxyethenyl)-3,3,5,5tetramethylcyclohexylidene]hydroxylamine SMILES: CCOC(=C)C1C(C)(C)CC(CC1(C)C)=NO Molecular weight: 239.36 Volume (cubic Angstroems): 252.83 Atoms number (non-hydrogen): 17 miLogP: 4.33 Structure: Biological Properties: Predicted Druglikeness: GPCR ligand -0.6 Ion channel modulator -0.41 Kinase inhibitor -0.93 Nuclear receptor ligand -0.17 Protease inhibitor -0.39 Enzyme inhibitor 0.01 Commercial name: Sylvoxime© IUPAC Name: N-[4-(1-ethoxyethenyl)-3,3,5,5tetramethylcyclohexylidene]hydroxylamine SMILES: CCOC(=C)C1C(C)(C)CC(CC1(C)C)=NO Predicted Olfactory Receptor Activity: OR52D1 71.900% OR1G1 70.394% 0R52I2 70.392% OR52I1 70.390% OR2Y1 70.378% 3. Commercial name: Hyperflor© IUPAC Name: 2-benzyl-1,3-dioxan-5-one SMILES: O=C1COC(CC2=CC=CC=C2)OC1 Molecular weight: 192.21 g/mol Volume -

Prognostic Classifier Based on Genome-Wide DNA Methylation Profiling in Well-Differentiated Thyroid Tumors
CLINICAL RESEARCH ARTICLE Downloaded from https://academic.oup.com/jcem/article-abstract/102/11/4089/4082866 by Universidade Estadual Paulista J�lio de Mesquita Filho user on 06 May 2019 Prognostic Classifier Based on Genome-Wide DNA Methylation Profiling in Well-Differentiated Thyroid Tumors Mariana Bisarro dos Reis,1,2* Mateus Camargo Barros-Filho,1* F´abio Albuquerque Marchi,1 Caroline Moraes Beltrami,1 Hellen Kuasne,1 Clovis ´ Antonioˆ Lopes Pinto,3 Srikant Ambatipudi,4,5 Zdenko Herceg,4 Luiz Paulo Kowalski,6 and Silvia Regina Rogatto2,7 1International Research Center, CIPE, A.C. Camargo Cancer Center and National Institute of Science and Technology in Oncogenomics, S~ao Paulo 01509-010, SP, Brazil; 2Department of Urology, Faculty of Medicine, UNESP, S~ao Paulo State University, Botucatu 18618-970, SP, Brazil; 3Department of Pathology, A.C. Camargo Cancer Center, S~ao Paulo 01509-010, SP, Brazil; 4Epigenetics Group; International Agency for Research on Cancer (IARC), Lyon 69372, France; 5MRC Integrative Epidemiology Unit, University of Bristol, Bristol BS8 1TH, United Kingdom; 6Department of Head and Neck Surgery and Otorhinolaryngology, A.C. Camargo Cancer Center, S~ao Paulo 01509-010, SP, Brazil; and 7Department of Clinical Genetics, Vejle Hospital and Institute of Regional Health Research, University of Southern Denmark, Vejle, 7100, Denmark Context: Even though the majority of well-differentiated thyroid carcinoma (WDTC) is indolent, a number of cases display an aggressive behavior. Cumulative evidence suggests that the deregulation of DNA methylation has the potential to point out molecular markers associated with worse prognosis. Objective: To identify a prognostic epigenetic signature in thyroid cancer. -

Genetic Characterization of Greek Population Isolates Reveals Strong Genetic Drift at Missense and Trait-Associated Variants
ARTICLE Received 22 Apr 2014 | Accepted 22 Sep 2014 | Published 6 Nov 2014 DOI: 10.1038/ncomms6345 OPEN Genetic characterization of Greek population isolates reveals strong genetic drift at missense and trait-associated variants Kalliope Panoutsopoulou1,*, Konstantinos Hatzikotoulas1,*, Dionysia Kiara Xifara2,3, Vincenza Colonna4, Aliki-Eleni Farmaki5, Graham R.S. Ritchie1,6, Lorraine Southam1,2, Arthur Gilly1, Ioanna Tachmazidou1, Segun Fatumo1,7,8, Angela Matchan1, Nigel W. Rayner1,2,9, Ioanna Ntalla5,10, Massimo Mezzavilla1,11, Yuan Chen1, Chrysoula Kiagiadaki12, Eleni Zengini13,14, Vasiliki Mamakou13,15, Antonis Athanasiadis16, Margarita Giannakopoulou17, Vassiliki-Eirini Kariakli5, Rebecca N. Nsubuga18, Alex Karabarinde18, Manjinder Sandhu1,8, Gil McVean2, Chris Tyler-Smith1, Emmanouil Tsafantakis12, Maria Karaleftheri16, Yali Xue1, George Dedoussis5 & Eleftheria Zeggini1 Isolated populations are emerging as a powerful study design in the search for low-frequency and rare variant associations with complex phenotypes. Here we genotype 2,296 samples from two isolated Greek populations, the Pomak villages (HELIC-Pomak) in the North of Greece and the Mylopotamos villages (HELIC-MANOLIS) in Crete. We compare their genomic characteristics to the general Greek population and establish them as genetic isolates. In the MANOLIS cohort, we observe an enrichment of missense variants among the variants that have drifted up in frequency by more than fivefold. In the Pomak cohort, we find novel associations at variants on chr11p15.4 showing large allele frequency increases (from 0.2% in the general Greek population to 4.6% in the isolate) with haematological traits, for example, with mean corpuscular volume (rs7116019, P ¼ 2.3 Â 10 À 26). We replicate this association in a second set of Pomak samples (combined P ¼ 2.0 Â 10 À 36). -

Mrna Expression Profiles Obtained from Microdissected Pancreatic Cancer Cells Can Predict Patient Survival
www.impactjournals.com/oncotarget/ Oncotarget, 2017, Vol. 8, (No. 62), pp: 104796-104805 Research Paper mRNA expression profiles obtained from microdissected pancreatic cancer cells can predict patient survival Ana-Barbara García-García1,2,*, M. Carmen Gómez-Mateo3,7,*, Rebeca Hilario2, Pilar Rentero-Garrido2, Alvaro Martínez-Domenech4, Veronica Gonzalez-Albert2, Andres Cervantes5, Pablo Marín-Garcia6, Felipe Javier Chaves1,2, Antonio Ferrández- Izquierdo3 and Luis Sabater4 1CIBER of Diabetes and Associated Metabolic Diseases (CIBERDEM), Madrid, Spain 2Unidad de Genómica y Diagnóstico Genético. Fundación Investigación Clínico de Valencia, Instituto de Investigación Sanitaria Clínico de Valencia (INCLIVA), Valencia, Spain 3Department of Pathology, Faculty of Medicine and Odontology, University of Valencia and Clinical Hospital of Valencia, and Instituto de Investigación Sanitaria Clínico de Valencia (INCLIVA), Valencia, Spain 4Department of Surgery, University of Valencia, Hospital Clínico Universitario, Instituto de Investigación Sanitaria Clínico de Valencia (INCLIVA), Valencia, Spain 5Hematology and Medical Oncology Unit, Biomedical Research Institute INCLIVA and Department of Medicine, University of Valencia, Valencia, Spain 6Bioinformatics Unit. Fundación Investigación Clínico de Valencia, Instituto de Investigación Sanitaria Clínico de Valencia (INCLIVA), Valencia, Spain 7Current/Present address: Pathology Department, Hospital Universitario Donostia, San Sebastian, Spain *These authors have contributed equally to this work Correspondence -

Clinical, Molecular, and Immune Analysis of Dabrafenib-Trametinib
Supplementary Online Content Chen G, McQuade JL, Panka DJ, et al. Clinical, molecular and immune analysis of dabrafenib-trametinib combination treatment for metastatic melanoma that progressed during BRAF inhibitor monotherapy: a phase 2 clinical trial. JAMA Oncology. Published online April 28, 2016. doi:10.1001/jamaoncol.2016.0509. eMethods. eReferences. eTable 1. Clinical efficacy eTable 2. Adverse events eTable 3. Correlation of baseline patient characteristics with treatment outcomes eTable 4. Patient responses and baseline IHC results eFigure 1. Kaplan-Meier analysis of overall survival eFigure 2. Correlation between IHC and RNAseq results eFigure 3. pPRAS40 expression and PFS eFigure 4. Baseline and treatment-induced changes in immune infiltrates eFigure 5. PD-L1 expression eTable 5. Nonsynonymous mutations detected by WES in baseline tumors This supplementary material has been provided by the authors to give readers additional information about their work. © 2016 American Medical Association. All rights reserved. Downloaded From: https://jamanetwork.com/ on 09/30/2021 eMethods Whole exome sequencing Whole exome capture libraries for both tumor and normal samples were constructed using 100ng genomic DNA input and following the protocol as described by Fisher et al.,3 with the following adapter modification: Illumina paired end adapters were replaced with palindromic forked adapters with unique 8 base index sequences embedded within the adapter. In-solution hybrid selection was performed using the Illumina Rapid Capture Exome enrichment kit with 38Mb target territory (29Mb baited). The targeted region includes 98.3% of the intervals in the Refseq exome database. Dual-indexed libraries were pooled into groups of up to 96 samples prior to hybridization. -

OR56A3 Rabbit Polyclonal Antibody – TA332338 | Origene
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for TA332338 OR56A3 Rabbit Polyclonal Antibody Product data: Product Type: Primary Antibodies Applications: WB Recommended Dilution: WB Reactivity: Human Host: Rabbit Isotype: IgG Clonality: Polyclonal Immunogen: The immunogen for Anti-OR56A3 Antibody is: synthetic peptide directed towards the C- terminal region of Human OR56A3. Synthetic peptide located within the following region: RAVLRLKAEGAVAKALSTCGSHFMLILFFSTILLVFVLTHVAKKKVSPDV Formulation: Liquid. Purified antibody supplied in 1x PBS buffer with 0.09% (w/v) sodium azide and 2% sucrose. Note that this product is shipped as lyophilized powder to China customers. Conjugation: Unconjugated Storage: Store at -20°C as received. Stability: Stable for 12 months from date of receipt. Predicted Protein Size: 35 kDa Gene Name: olfactory receptor family 56 subfamily A member 3 Database Link: NP_001003443 Entrez Gene 390083 Human Q8NH54 Background: Olfactory receptors interact with odorant molecules in the nose, to initiate a neuronal response that triggers the perception of a smell. The olfactory receptor proteins are members of a large family of G-protein-coupled receptors (GPCR) arising from single coding- exon genes. Olfactory receptors share a 7-transmembrane domain structure with many neurotransmitter and hormone receptors and are responsible for the recognition and G protein-mediated transduction of odorant signals. The olfactory receptor gene family is the largest in the genome. The nomenclature assigned to the olfactory receptor genes and proteins for this organism is independent of other organisms. This product is to be used for laboratory only.