Qt4vh1p2c4 Nosplash E372185
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Rare Modifier Variants Alter the Severity of Cardiovascular Disease in Pseudoxanthoma Elasticum
fcell-09-612581 June 8, 2021 Time: 11:34 # 1 BRIEF RESEARCH REPORT published: 08 June 2021 doi: 10.3389/fcell.2021.612581 Rare Modifier Variants Alter the Severity of Cardiovascular Disease in Pseudoxanthoma Elasticum: Identification of Novel Candidate Modifier Genes and Disease Pathways Through Mixture of Effects Analysis Eva Y. G. De Vilder1,2,3, Ludovic Martin4, Georges Lefthériotis5, Paul Coucke1,6, Filip Van Nieuwerburgh7 and Olivier M. Vanakker1,6* Edited by: 1 2 Svetlana Komarova, Center for Medical Genetics, Ghent University Hospital, Ghent, Belgium, The Research Foundation – Flanders, Ghent, 3 4 McGill University, Canada Belgium, Department of Ophthalmology, Ghent University Hospital, Ghent, Belgium, Department of Dermatology, Angers University Hospital, Angers, France, 5 Department of Vascular Physiology and Sports Medicine, Angers University, Angers, Reviewed by: France, 6 Department of Biomolecular Medicine, Ghent University, Ghent, Belgium, 7 Department of Pharmaceutics, Carolina Beraldo Meloto, Laboratory of Pharmaceutical Biotechnology, Ghent University, Ghent, Belgium McGill University, Canada Koen van de Wetering, Thomas Jefferson University, Introduction: Pseudoxanthoma elasticum (PXE), an ectopic mineralization disorder United States caused by pathogenic ABCC6 variants, is characterized by skin, ocular and Jessica Bertrand, University Hospital Magdeburg, cardiovascular (CV) symptoms. Due to striking phenotypic variability without genotype- Germany phenotype correlations, modifier genes are thought to play a role in disease variability. *Correspondence: In this study, we evaluated the collective modifying effect of rare variants on the Olivier M. Vanakker [email protected] cardiovascular phenotype of PXE. Materials and Methods: Mixed effects of rare variants were assessed by Whole Exome Specialty section: This article was submitted to Sequencing in 11 PXE patients with an extreme CV phenotype (mild/severe). -

Genetic Variation Across the Human Olfactory Receptor Repertoire Alters Odor Perception
bioRxiv preprint doi: https://doi.org/10.1101/212431; this version posted November 1, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. Genetic variation across the human olfactory receptor repertoire alters odor perception Casey Trimmer1,*, Andreas Keller2, Nicolle R. Murphy1, Lindsey L. Snyder1, Jason R. Willer3, Maira Nagai4,5, Nicholas Katsanis3, Leslie B. Vosshall2,6,7, Hiroaki Matsunami4,8, and Joel D. Mainland1,9 1Monell Chemical Senses Center, Philadelphia, Pennsylvania, USA 2Laboratory of Neurogenetics and Behavior, The Rockefeller University, New York, New York, USA 3Center for Human Disease Modeling, Duke University Medical Center, Durham, North Carolina, USA 4Department of Molecular Genetics and Microbiology, Duke University Medical Center, Durham, North Carolina, USA 5Department of Biochemistry, University of Sao Paulo, Sao Paulo, Brazil 6Howard Hughes Medical Institute, New York, New York, USA 7Kavli Neural Systems Institute, New York, New York, USA 8Department of Neurobiology and Duke Institute for Brain Sciences, Duke University Medical Center, Durham, North Carolina, USA 9Department of Neuroscience, University of Pennsylvania School of Medicine, Philadelphia, Pennsylvania, USA *[email protected] ABSTRACT The human olfactory receptor repertoire is characterized by an abundance of genetic variation that affects receptor response, but the perceptual effects of this variation are unclear. To address this issue, we sequenced the OR repertoire in 332 individuals and examined the relationship between genetic variation and 276 olfactory phenotypes, including the perceived intensity and pleasantness of 68 odorants at two concentrations, detection thresholds of three odorants, and general olfactory acuity. -

Genome-Wide Association Study Identifies Loci for Arterial Stiffness
www.nature.com/scientificreports OPEN Genome-wide association study identifes loci for arterial stifness index in 127,121 UK Biobank Received: 1 February 2019 Accepted: 5 June 2019 participants Published: xx xx xxxx Kenneth Fung1, Julia Ramírez 2, Helen R. Warren2,3, Nay Aung 1, Aaron M. Lee1, Evan Tzanis2,3, Stefen E. Petersen 1,3 & Patricia B. Munroe2,3 Arterial stifness index (ASI) is a non-invasive measure of arterial stifness using infra-red fnger sensors (photoplethysmography). It is a well-suited measure for large populations as it is relatively inexpensive to perform, and data can be acquired within seconds. These features raise interest in using ASI as a tool to estimate cardiovascular disease risk as prior work demonstrates increased arterial stifness is associated with elevated systolic blood pressure, and ASI is predictive of cardiovascular disease and mortality. We conducted genome-wide association studies (GWASs) for ASI in 127,121 UK Biobank participants of European-ancestry. Our primary analyses identifed variants at four loci reaching genome-wide signifcance (P < 5 × 10−8): TEX41 (rs1006923; P = 5.3 × 10−12), FOXO1 (rs7331212; P = 2.2 × 10−11), C1orf21 (rs1930290, P = 1.1 × 10−8) and MRVI1 (rs10840457, P = 3.4 × 10−8). Gene- based testing revealed three signifcant genes, the most signifcant gene was COL4A2 (P = 1.41 × 10−8) encoding type IV collagen. Other candidate genes at associated loci were also involved in smooth muscle tone regulation. Our fndings provide new information for understanding the development of arterial stifness. Arterial stifness measures have been reported as independent markers of vascular ageing1,2, hypertension3,4, car- diovascular disease (CVD)5,6 and mortality6,7. -

Supplementary Table 9. Functional Annotation Clustering Results for the Union (GS3) of the Top Genes from the SNP-Level and Gene-Based Analyses (See ST4)
Supplementary Table 9. Functional Annotation Clustering Results for the union (GS3) of the top genes from the SNP-level and Gene-based analyses (see ST4) Column Header Key Annotation Cluster Name of cluster, sorted by descending Enrichment score Enrichment Score EASE enrichment score for functional annotation cluster Category Pathway Database Term Pathway name/Identifier Count Number of genes in the submitted list in the specified term % Percentage of identified genes in the submitted list associated with the specified term PValue Significance level associated with the EASE enrichment score for the term Genes List of genes present in the term List Total Number of genes from the submitted list present in the category Pop Hits Number of genes involved in the specified term (category-specific) Pop Total Number of genes in the human genome background (category-specific) Fold Enrichment Ratio of the proportion of count to list total and population hits to population total Bonferroni Bonferroni adjustment of p-value Benjamini Benjamini adjustment of p-value FDR False Discovery Rate of p-value (percent form) Annotation Cluster 1 Enrichment Score: 3.8978262119731335 Category Term Count % PValue Genes List Total Pop Hits Pop Total Fold Enrichment Bonferroni Benjamini FDR GOTERM_CC_DIRECT GO:0005886~plasma membrane 383 24.33290978 5.74E-05 SLC9A9, XRCC5, HRAS, CHMP3, ATP1B2, EFNA1, OSMR, SLC9A3, EFNA3, UTRN, SYT6, ZNRF2, APP, AT1425 4121 18224 1.18857065 0.038655922 0.038655922 0.086284383 UP_KEYWORDS Membrane 626 39.77128335 1.53E-04 SLC9A9, HRAS, -

Time-Series Plasma Cell-Free DNA Analysis Reveals Disease Severity of COVID-19 Patients
medRxiv preprint doi: https://doi.org/10.1101/2020.06.08.20124305; this version posted June 9, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. It is made available under a CC-BY-NC-ND 4.0 International license . Time-series plasma cell-free DNA analysis reveals disease severity of COVID- 19 patients Authors: Xinping Chen1†, Yu Lin2†, Tao Wu1†, Jinjin Xu2†, Zhichao Ma1†, Kun Sun2,5†, Hui Li1†, Yuxue Luo2,3†, Chen Zhang1, Fang Chen2, Jiao Wang1, Tingyu Kuo2,4, Xiaojuan Li1, Chunyu Geng2, Feng Lin1, Chaojie Huang2, Junjie Hu1, Jianhua Yin2, Ming Liu1, Ye Tao2, Jiye Zhang1, Rijing Ou2, Furong Xiao1, Huanming Yang2,6, Jian Wang2,6, Xun Xu2,7, Shengmiao Fu1*, Xin Jin2,3*, Hongyan Jiang1*, Ruoyan Chen2* Affiliations: 1Hainan General Hospital, Hainan Affiliated Hospital of Hainan Medical University, Hainan Provincial Key Laboratory of Cell and Molecular Genetic Translational Medicine, Haikou 570311, Hainan, China. 2BGI-Shenzhen, Shenzhen, 518083, Guangdong, China 3School of Medicine, South China University of Technology, Guangzhou 510006, Guangdong, China 4BGI Education Center, University of Chinese Academy of Sciences, Shenzhen 518083, Guangdong, China 5Shenzhen Bay Laboratory, Shenzhen 518132, Guangdong, China 6James D. Watson Institute of Genome Sciences, Hangzhou 310058, China 7Guangdong Provincial Key Laboratory of Genome Read and Write, BGI-Shenzhen, Shenzhen, 518120, China *Correspondence to: [email protected]; [email protected]; [email protected]; [email protected]. †These authors contributed equally to this work. Abstract: Clinical symptoms of coronavirus disease 2019 (COVID-19) range from asymptomatic to severe pneumonia and death. -
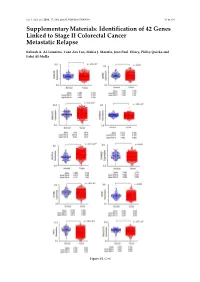
Identification of 42 Genes Linked to Stage II Colorectal Cancer Metastatic Relapse
Int. J. Mol. Sci. 2016, 17, 598; doi:10.3390/ijms17040598 S1 of S16 Supplementary Materials: Identification of 42 Genes Linked to Stage II Colorectal Cancer Metastatic Relapse Rabeah A. Al-Temaimi, Tuan Zea Tan, Makia J. Marafie, Jean Paul Thiery, Philip Quirke and Fahd Al-Mulla Figure S1. Cont. Int. J. Mol. Sci. 2016, 17, 598; doi:10.3390/ijms17040598 S2 of S16 Figure S1. Mean expression levels of fourteen genes of significant association with CRC DFS and OS that are differentially expressed in normal colon compared to CRC tissues. Each dot represents a sample. Table S1. Copy number aberrations associated with poor disease-free survival and metastasis in early stage II CRC as predicted by STAC and SPPS combined methodologies with resident gene symbols. CN stands for copy number, whereas CNV is copy number variation. Region Cytoband % of CNV Count of Region Event Gene Symbols Length Location Overlap Genes chr1:113,025,076–113,199,133 174,057 p13.2 CN Loss 0.0 2 AKR7A2P1, SLC16A1 chr1:141,465,960–141,822,265 356,305 q12–q21.1 CN Gain 95.9 1 SRGAP2B MIR5087, LOC10013000 0, FLJ39739, LOC10028679 3, PPIAL4G, PPIAL4A, NBPF14, chr1:144,911,564–146,242,907 1,331,343 q21.1 CN Gain 99.6 16 NBPF15, NBPF16, PPIAL4E, NBPF16, PPIAL4D, PPIAL4F, LOC645166, LOC388692, FCGR1C chr1:177,209,428–177,226,812 17,384 q25.3 CN Gain 0.0 0 chr1:197,652,888–197,676,831 23,943 q32.1 CN Gain 0.0 1 KIF21B chr1:201,015,278–201,033,308 18,030 q32.1 CN Gain 0.0 1 PLEKHA6 chr1:201,289,154–201,298,247 9093 q32.1 CN Gain 0.0 0 chr1:216,820,186–217,043,421 223,235 q41 CN -

Identification of Candidate Biomarkers and Pathways Associated with Type 1 Diabetes Mellitus Using Bioinformatics Analysis
bioRxiv preprint doi: https://doi.org/10.1101/2021.06.08.447531; this version posted June 9, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Identification of candidate biomarkers and pathways associated with type 1 diabetes mellitus using bioinformatics analysis Basavaraj Vastrad1, Chanabasayya Vastrad*2 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karnataka, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India bioRxiv preprint doi: https://doi.org/10.1101/2021.06.08.447531; this version posted June 9, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract Type 1 diabetes mellitus (T1DM) is a metabolic disorder for which the underlying molecular mechanisms remain largely unclear. This investigation aimed to elucidate essential candidate genes and pathways in T1DM by integrated bioinformatics analysis. In this study, differentially expressed genes (DEGs) were analyzed using DESeq2 of R package from GSE162689 of the Gene Expression Omnibus (GEO). Gene ontology (GO) enrichment analysis, REACTOME pathway enrichment analysis, and construction and analysis of protein-protein interaction (PPI) network, modules, miRNA-hub gene regulatory network and TF-hub gene regulatory network, and validation of hub genes were then performed. A total of 952 DEGs (477 up regulated and 475 down regulated genes) were identified in T1DM. GO and REACTOME enrichment result results showed that DEGs mainly enriched in multicellular organism development, detection of stimulus, diseases of signal transduction by growth factor receptors and second messengers, and olfactory signaling pathway. -

DNMT Inhibitors Increase Methylation at Subset of Cpgs in Colon
bioRxiv preprint doi: https://doi.org/10.1101/395467; this version posted August 25, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Title: DNMT inhibitors increase methylation at subset of CpGs in colon, bladder, lymphoma, 2 breast, and ovarian, cancer genome 3 Running title: Decitabine/azacytidine increases DNA methylation 4 Anil K Giri1, Tero Aittokallio1,2 5 1Institute for Molecular Medicine Finland, FIMM, University of Helsinki, Helsinki, Finland. 6 2Department of Mathematics and Statistics, University of Turku, Turku, Finland. 7 Correspondence to 8 Dr. Anil K Giri 9 Institute for Molecular Medicine Finland FIMM, University of Helsinki, Helsinki, Finland. 10 Email: [email protected] 11 Financial disclosure: This work was funded by the Academy of Finland (grants 269862, 292611, 12 310507 and 313267), Cancer Society of Finland, and the Sigrid Juselius Foundation. 13 Ethical disclosure: This study is an independent analysis of existing data available in the public 14 domain and does not involve any animal or human samples that have been collected by the authors 15 themselves. 16 Author contribution: AKG conceptualized, analyzed the data and wrote the manuscript. TA 17 critically revised and edited the manuscript. The authors report no conflict of interest. 18 19 Word count: 20 Figure number: 5 21 Table number: 1 22 23 Abstract 24 Background: DNA methyltransferase inhibitors (DNMTi) decitabine and azacytidine are approved 25 therapies for acute myeloid leukemia and myelodysplastic syndrome. -

Qt38n028mr Nosplash A3e1d84
! ""! ACKNOWLEDGEMENTS I dedicate this thesis to my parents who inspired me to become a scientist through invigorating scientific discussions at the dinner table even when I was too young to understand what the hippocampus was. They also prepared me for the ups and downs of science and supported me through all of these experiences. I would like to thank my advisor Dr. Elizabeth Blackburn and my thesis committee members Dr. Eric Verdin, and Dr. Emmanuelle Passegue. Liz created a nurturing and supportive environment for me to explore my own ideas, while at the same time teaching me how to love science, test my questions, and of course provide endless ways to think about telomeres and telomerase. Eric and Emmanuelle both gave specific critical advice about the proper experiments for T cells and both volunteered their lab members for further critical advice. I always felt inspired with a sense of direction after thesis committee meetings. The Blackburn lab is full of smart and dedicated scientists whom I am thankful for their support. Specifically Dr. Shang Li and Dr. Brad Stohr for their stimulating scientific debates and “arguments.” Dr. Jue Lin, Dana Smith, Kyle Lapham, Dr. Tet Matsuguchi, and Kyle Jay for their friendships and discussions about what my data could possibly mean. Dr. Eva Samal for teaching me molecular biology techniques and putting up with my late night lab exercises. Beth Cimini for her expertise with microscopy, FACs, singing, and most of all for being a caring and supportive friend. Finally, I would like to thank Dr. Imke Listerman, my scientific partner for most of the breast cancer experiments. -

The Mutational Landscape of Human Olfactory G Protein-Coupled Receptors
Jimenez et al. BMC Biology (2021) 19:21 https://doi.org/10.1186/s12915-021-00962-0 RESEARCH ARTICLE Open Access The mutational landscape of human olfactory G protein-coupled receptors Ramón Cierco Jimenez1,2, Nil Casajuana-Martin1, Adrián García-Recio1, Lidia Alcántara1, Leonardo Pardo1, Mercedes Campillo1 and Angel Gonzalez1* Abstract Background: Olfactory receptors (ORs) constitute a large family of sensory proteins that enable us to recognize a wide range of chemical volatiles in the environment. By contrast to the extensive information about human olfactory thresholds for thousands of odorants, studies of the genetic influence on olfaction are limited to a few examples. To annotate on a broad scale the impact of mutations at the structural level, here we analyzed a compendium of 119,069 natural variants in human ORs collected from the public domain. Results: OR mutations were categorized depending on their genomic and protein contexts, as well as their frequency of occurrence in several human populations. Functional interpretation of the natural changes was estimated from the increasing knowledge of the structure and function of the G protein-coupled receptor (GPCR) family, to which ORs belong. Our analysis reveals an extraordinary diversity of natural variations in the olfactory gene repertoire between individuals and populations, with a significant number of changes occurring at the structurally conserved regions. A particular attention is paid to mutations in positions linked to the conserved GPCR activation mechanism that could imply phenotypic variation in the olfactory perception. An interactive web application (hORMdb, Human Olfactory Receptor Mutation Database) was developed for the management and visualization of this mutational dataset. -

WO 2019/068007 Al Figure 2
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization I International Bureau (10) International Publication Number (43) International Publication Date WO 2019/068007 Al 04 April 2019 (04.04.2019) W 1P O PCT (51) International Patent Classification: (72) Inventors; and C12N 15/10 (2006.01) C07K 16/28 (2006.01) (71) Applicants: GROSS, Gideon [EVIL]; IE-1-5 Address C12N 5/10 (2006.0 1) C12Q 1/6809 (20 18.0 1) M.P. Korazim, 1292200 Moshav Almagor (IL). GIBSON, C07K 14/705 (2006.01) A61P 35/00 (2006.01) Will [US/US]; c/o ImmPACT-Bio Ltd., 2 Ilian Ramon St., C07K 14/725 (2006.01) P.O. Box 4044, 7403635 Ness Ziona (TL). DAHARY, Dvir [EilL]; c/o ImmPACT-Bio Ltd., 2 Ilian Ramon St., P.O. (21) International Application Number: Box 4044, 7403635 Ness Ziona (IL). BEIMAN, Merav PCT/US2018/053583 [EilL]; c/o ImmPACT-Bio Ltd., 2 Ilian Ramon St., P.O. (22) International Filing Date: Box 4044, 7403635 Ness Ziona (E.). 28 September 2018 (28.09.2018) (74) Agent: MACDOUGALL, Christina, A. et al; Morgan, (25) Filing Language: English Lewis & Bockius LLP, One Market, Spear Tower, SanFran- cisco, CA 94105 (US). (26) Publication Language: English (81) Designated States (unless otherwise indicated, for every (30) Priority Data: kind of national protection available): AE, AG, AL, AM, 62/564,454 28 September 2017 (28.09.2017) US AO, AT, AU, AZ, BA, BB, BG, BH, BN, BR, BW, BY, BZ, 62/649,429 28 March 2018 (28.03.2018) US CA, CH, CL, CN, CO, CR, CU, CZ, DE, DJ, DK, DM, DO, (71) Applicant: IMMP ACT-BIO LTD. -

Supplementary Table 2
Supplementary Table 2. Non-synonymous somatic mutations identified in the Discovery set of four NKTCL cases. Gene Sample Transcript Amino Mutation No. Nucleotide (Genomic) Symbol ID Accession ID Acid Type 1 ABCA4 7 CCDS747.1 g.chr1: 94564391 T>C p.T243A Missense 2 ACOX2 7 ENST00000492530 g.chr3: 58517538 C>T p.G7R Missense 3 ACSS3 10 CCDS9022.1 g.chr12: 81503369 C>A p.Y114X Nonsense 4 ADAMTS2 7 CCDS4444.1 g.chr5: 178540908 A>C p.I1199S Missense 5 AKAP8 7 CCDS12329.1 g.chr19: 15483121 C>T p.G300D Missense 6 ANGEL2 7 CCDS1512.1 g.chr1: 213168445 G>A p.P525S Missense 7 ANKZF1 59 CCDS42821.1 g.chr2: 220100475 C>T p.R617W Missense 8 APBA3 59 ENST00000439726 g.chr19: 3751342 insG fs Insertion 9 APLNR 7 CCDS7950.1 g.chr11: 57004458 A>T p.F7L Missense 10 ARMC7 7 CCDS11714.1 g.chr17: 73124795 G>A p.D87N Missense 11 ATM 7 CCDS31669.1 g.chr11: 108192038 G>A p.V2155M Missense 12 ATP6V0A1 7 CCDS45683.1 g.chr17: 40666340 A>T p.H762L Missense 13 BNIP3 7 CCDS7663.1 g.chr10: 133787372 C>T p.G41E Missense 14 BRCA1 10 CCDS11453.1 g.chr17: 41246517 G>A p.A344V Missense 15 BRI3 7 CCDS5656.1 g.chr7: 97911710 G>A p.V64I Missense 16 BSG 7 CCDS12032.1 g.chr19: 571548 A>G p.M1V Missense 17 BTAF1 7 CCDS7419.1 g.chr10: 93713569 A>G p.R214G Missense 18 BTBD11 59 CCDS31893.1 g.chr12: 107914407 G>A p.E427K Missense 19 BTN2A3 10 ENST00000465856 g.chr6: 26423337 G>A p.E86K Missense 20 C13orf23 31 CCDS45041.1 g.chr13: 39587665 delA fs Deletion 21 C2CD3 7 CCDS31636.1 g.chr11: 73753262 T>G p.S1833R Missense 22 C3orf63 31 CCDS46853.1 g.chr3: 56681175 A>C p.F530L Missense