Predator Complementarity Dampens Variability of Phytoplankton Biomass in a Diversity
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

RECORDS of the HAWAII BIOLOGICAL SURVEY for 1995 Part 2: Notes1
RECORDS OF THE HAWAII BIOLOGICAL SURVEY FOR 1995 Part 2: Notes1 This is the second of two parts to the Records of the Hawaii Biological Survey for 1995 and contains the notes on Hawaiian species of plants and animals including new state and island records, range extensions, and other information. Larger, more compre- hensive treatments and papers describing new taxa are treated in the first part of this Records [Bishop Museum Occasional Papers 45]. New Hawaiian Pest Plant Records for 1995 PATRICK CONANT (Hawaii Dept. of Agriculture, Plant Pest Control Branch, 1428 S King St, Honolulu, HI 96814) Fabaceae Ulex europaeus L. New island record On 6 October 1995, Hawaii Department of Land and Natural Resources, Division of Forestry and Wildlife employee C. Joao submitted an unusual plant he found while work- ing in the Molokai Forest Reserve. The plant was identified as U. europaeus and con- firmed by a Hawaii Department of Agriculture (HDOA) nox-A survey of the site on 9 October revealed an infestation of ca. 19 m2 at about 457 m elevation in the Kamiloa Distr., ca. 6.2 km above Kamehameha Highway. Distribution in Wagner et al. (1990, Manual of the flowering plants of Hawai‘i, p. 716) listed as Maui and Hawaii. Material examined: MOLOKAI: Molokai Forest Reserve, 4 Dec 1995, Guy Nagai s.n. (BISH). Melastomataceae Miconia calvescens DC. New island record, range extensions On 11 October, a student submitted a leaf specimen from the Wailua Houselots area on Kauai to PPC technician A. Bell, who had the specimen confirmed by David Lorence of the National Tropical Botanical Garden as being M. -

Hundreds of Species of Aquatic Macroinvertebrates Live in Illinois In
Illinois A B aquatic sowbug Asellus sp. Photograph © Paul P.Tinerella AAqquuaattiicc mayfly A. adult Hexagenia sp.; B. nymph Isonychia sp. MMaaccrrooiinnvveerrtteebbrraatteess Photographs © Michael R. Jeffords northern clearwater crayfish Orconectes propinquus Photograph © Michael R. Jeffords ruby spot damselfly Hetaerina americana Photograph © Michael R. Jeffords aquatic snail Pleurocera acutum Photograph © Jochen Gerber,The Field Museum of Natural History predaceous diving beetle Dytiscus circumcinctus Photograph © Paul P.Tinerella monkeyface mussel Quadrula metanevra common skimmer dragonfly - nymph Libellula sp. Photograph © Kevin S. Cummings Photograph © Paul P.Tinerella water scavenger beetle Hydrochara sp. Photograph © Steve J.Taylor devil crayfish Cambarus diogenes A B Photograph © ChristopherTaylor dobsonfly Corydalus sp. A. larva; B. adult Photographs © Michael R. Jeffords common darner dragonfly - nymph Aeshna sp. Photograph © Paul P.Tinerella giant water bug Belostoma lutarium Photograph © Paul P.Tinerella aquatic worm Slavina appendiculata Photograph © Mark J. Wetzel water boatman Trichocorixa calva Photograph © Paul P.Tinerella aquatic mite Order Prostigmata Photograph © Michael R. Jeffords backswimmer Notonecta irrorata Photograph © Paul P.Tinerella leech - adult and young Class Hirudinea pygmy backswimmer Neoplea striola mosquito - larva Toxorhynchites sp. fishing spider Dolomedes sp. Photograph © William N. Roston Photograph © Paul P.Tinerella Photograph © Michael R. Jeffords Photograph © Paul P.Tinerella Species List Species are not shown in proportion to actual size. undreds of species of aquatic macroinvertebrates live in Illinois in a Kingdom Animalia Hvariety of habitats. Some of the habitats have flowing water while Phylum Annelida Class Clitellata Family Naididae aquatic worm Slavina appendiculata This poster was made possible by: others contain still water. In order to survive in water, these organisms Class Hirudinea leech must be able to breathe, find food, protect themselves, move and reproduce. -

Underwater Breathing: the Mechanics of Plastron Respiration
J. Fluid Mech. (2008), vol. 608, pp. 275–296. c 2008 Cambridge University Press 275 doi:10.1017/S0022112008002048 Printed in the United Kingdom Underwater breathing: the mechanics of plastron respiration M. R. FLYNN† AND J O H N W. M. B U S H Department of Mathematics, Massachusetts Institute of Technology, 77 Massachusetts Avenue, Cambridge, MA 02139-4307, USA (Received 11 July 2007 and in revised form 10 April 2008) The rough, hairy surfaces of many insects and spiders serve to render them water-repellent; consequently, when submerged, many are able to survive by virtue of a thin air layer trapped along their exteriors. The diffusion of dissolved oxygen from the ambient water may allow this layer to function as a respiratory bubble or ‘plastron’, and so enable certain species to remain underwater indefinitely. Main- tenance of the plastron requires that the curvature pressure balance the pressure difference between the plastron and ambient. Moreover, viable plastrons must be of sufficient area to accommodate the interfacial exchange of O2 and CO2 necessary to meet metabolic demands. By coupling the bubble mechanics, surface and gas-phase chemistry, we enumerate criteria for plastron viability and thereby deduce the range of environmental conditions and dive depths over which plastron breathers can survive. The influence of an external flow on plastron breathing is also examined. Dynamic pressure may become significant for respiration in fast-flowing, shallow and well-aerated streams. Moreover, flow effects are generally significant because they sharpen chemical gradients and so enhance mass transfer across the plastron interface. Modelling this process provides a rationale for the ventilation movements documented in the biology literature, whereby arthropods enhance plastron respiration by flapping their limbs or antennae. -

Biological Diversity, Ecological Health and Condition of Aquatic Assemblages at National Wildlife Refuges in Southern Indiana, USA
Biodiversity Data Journal 3: e4300 doi: 10.3897/BDJ.3.e4300 Taxonomic Paper Biological Diversity, Ecological Health and Condition of Aquatic Assemblages at National Wildlife Refuges in Southern Indiana, USA Thomas P. Simon†, Charles C. Morris‡, Joseph R. Robb§, William McCoy | † Indiana University, Bloomington, IN 46403, United States of America ‡ US National Park Service, Indiana Dunes National Lakeshore, Porter, IN 47468, United States of America § US Fish and Wildlife Service, Big Oaks National Wildlife Refuge, Madison, IN 47250, United States of America | US Fish and Wildlife Service, Patoka River National Wildlife Refuge, Oakland City, IN 47660, United States of America Corresponding author: Thomas P. Simon ([email protected]) Academic editor: Benjamin Price Received: 08 Dec 2014 | Accepted: 09 Jan 2015 | Published: 12 Jan 2015 Citation: Simon T, Morris C, Robb J, McCoy W (2015) Biological Diversity, Ecological Health and Condition of Aquatic Assemblages at National Wildlife Refuges in Southern Indiana, USA. Biodiversity Data Journal 3: e4300. doi: 10.3897/BDJ.3.e4300 Abstract The National Wildlife Refuge system is a vital resource for the protection and conservation of biodiversity and biological integrity in the United States. Surveys were conducted to determine the spatial and temporal patterns of fish, macroinvertebrate, and crayfish populations in two watersheds that encompass three refuges in southern Indiana. The Patoka River National Wildlife Refuge had the highest number of aquatic species with 355 macroinvertebrate taxa, six crayfish species, and 82 fish species, while the Big Oaks National Wildlife Refuge had 163 macroinvertebrate taxa, seven crayfish species, and 37 fish species. The Muscatatuck National Wildlife Refuge had the lowest diversity of macroinvertebrates with 96 taxa and six crayfish species, while possessing the second highest fish species richness with 51 species. -

Aquatic Macroinvertebrates Section a Aquatic Macroinvertebrates (Exclusive of Mosquitoes)
I LLINOI S UNIVERSITY OF ILLINOIS AT URBANA-CHAMPAIGN PRODUCTION NOTE University of Illinois at Urbana-Champaign Library Large-scale Digitization Project, 2007. \oc iatural History Survey. Library iiAOs (ClSCi;; ILLINOIS - NATURAL HISTORY Ai . .ý . - I-w. Iv mk U16 OL SURVEY CHAPTER 9 AQUATIC MACROINVERTEBRATES SECTION A AQUATIC MACROINVERTEBRATES (EXCLUSIVE OF MOSQUITOES) Final Report October, 1985 Section of Faunistic Surveys and Insect Identification Technical Report by Allison R. Brigham, Lawrence M. Page, John D. Unzicker Mark J. Wetzel, Warren U. Brigham, Donald W. Webb, and Liane Suloway Prepared for Wetlands Research, Inc. 53 West Jackson Boulevard Chicago, IL 60604 Arjpp, Section of Faunistic Surveys and Insect Identification Technical Report 1985 (6) 6'Wa- CHAPTER 9 AQUATIC MACROINVERTEBRATES SECTION A AQUATIC MACROINVERTEBRATES (EXCLUSIVE OF MOSQUITOES) Allison R. Brigham, Lawrence M. Page, John D. Unzicker Mark J. Wetzel, Warren U. Brigham, Donald W. Webb, and Liane Suloway INTRODUCTION Aquatic macroinvertebrates are primary and secondary level consumers that play an important role in transferring energy through the different trophic levels of the food chains of aquatic ecosystems. These animals feed upon submerged and emergent macrophytes, plankton, and organic material suspended in the water column. Burrowing and feeding activities aid in the decomposition of plant and animal matter and the eventual recycling of nutrients. In addition, these organisms prey upon each other and serve as food for fishes, certain birds, and other animals. In general, aquatic macroinvertebrates have not been systematically surveyed in Illinois, and rarely have individual species been studied ecologically. This is due, in part, to the inconspicuous nature of most freshwater inverte- brates and the many taxonomic problems which preclude distributional, ecologi- cal, and other studies. -
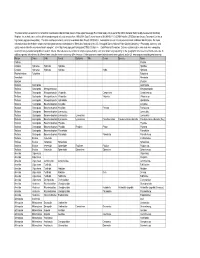
This Table Contains a Taxonomic List of Benthic Invertebrates Collected from Streams in the Upper Mississippi River Basin Study
This table contains a taxonomic list of benthic invertebrates collected from streams in the Upper Mississippi River Basin study unit as part of the USGS National Water Quality Assessemnt (NAWQA) Program. Invertebrates were collected from woody snags in selected streams from 1996-2004. Data Retreival occurred 26-JAN-06 11.10.25 AM from the USGS data warehouse (Taxonomic List Invert http://water.usgs.gov/nawqa/data). The data warehouse currently contains invertebrate data through 09/30/2002. Invertebrate taxa can include provisional and conditional identifications. For more information about invertebrate sample processing and taxonomic standards see, "Methods of analysis by the U.S. Geological Survey National Water Quality Laboratory -- Processing, taxonomy, and quality control of benthic macroinvertebrate samples", at << http://nwql.usgs.gov/Public/pubs/OFR00-212.html >>. Data Retrieval Precaution: Extreme caution must be exercised when comparing taxonomic lists generated using different search criteria. This is because the number of samples represented by each taxa list will vary depending on the geographic criteria selected for the retrievals. In addition, species lists retrieved at different times using the same criteria may differ because: (1) the taxonomic nomenclature (names) were updated, and/or (2) new samples containing new taxa may Phylum Class Order Family Subfamily Tribe Genus Species Taxon Porifera Porifera Cnidaria Hydrozoa Hydroida Hydridae Hydridae Cnidaria Hydrozoa Hydroida Hydridae Hydra Hydra sp. Platyhelminthes Turbellaria Turbellaria Nematoda Nematoda Bryozoa Bryozoa Mollusca Gastropoda Gastropoda Mollusca Gastropoda Mesogastropoda Mesogastropoda Mollusca Gastropoda Mesogastropoda Viviparidae Campeloma Campeloma sp. Mollusca Gastropoda Mesogastropoda Viviparidae Viviparus Viviparus sp. Mollusca Gastropoda Mesogastropoda Hydrobiidae Hydrobiidae Mollusca Gastropoda Basommatophora Ancylidae Ancylidae Mollusca Gastropoda Basommatophora Ancylidae Ferrissia Ferrissia sp. -

Great Lakes Entomologist the Grea T Lakes E N Omo L O G Is T Published by the Michigan Entomological Society Vol
The Great Lakes Entomologist THE GREA Published by the Michigan Entomological Society Vol. 45, Nos. 3 & 4 Fall/Winter 2012 Volume 45 Nos. 3 & 4 ISSN 0090-0222 T LAKES Table of Contents THE Scholar, Teacher, and Mentor: A Tribute to Dr. J. E. McPherson ..............................................i E N GREAT LAKES Dr. J. E. McPherson, Educator and Researcher Extraordinaire: Biographical Sketch and T List of Publications OMO Thomas J. Henry ..................................................................................................111 J.E. McPherson – A Career of Exemplary Service and Contributions to the Entomological ENTOMOLOGIST Society of America L O George G. Kennedy .............................................................................................124 G Mcphersonarcys, a New Genus for Pentatoma aequalis Say (Heteroptera: Pentatomidae) IS Donald B. Thomas ................................................................................................127 T The Stink Bugs (Hemiptera: Heteroptera: Pentatomidae) of Missouri Robert W. Sites, Kristin B. Simpson, and Diane L. Wood ............................................134 Tymbal Morphology and Co-occurrence of Spartina Sap-feeding Insects (Hemiptera: Auchenorrhyncha) Stephen W. Wilson ...............................................................................................164 Pentatomoidea (Hemiptera: Pentatomidae, Scutelleridae) Associated with the Dioecious Shrub Florida Rosemary, Ceratiola ericoides (Ericaceae) A. G. Wheeler, Jr. .................................................................................................183 -

Microsoft Outlook
Joey Steil From: Leslie Jordan <[email protected]> Sent: Tuesday, September 25, 2018 1:13 PM To: Angela Ruberto Subject: Potential Environmental Beneficial Users of Surface Water in Your GSA Attachments: Paso Basin - County of San Luis Obispo Groundwater Sustainabilit_detail.xls; Field_Descriptions.xlsx; Freshwater_Species_Data_Sources.xls; FW_Paper_PLOSONE.pdf; FW_Paper_PLOSONE_S1.pdf; FW_Paper_PLOSONE_S2.pdf; FW_Paper_PLOSONE_S3.pdf; FW_Paper_PLOSONE_S4.pdf CALIFORNIA WATER | GROUNDWATER To: GSAs We write to provide a starting point for addressing environmental beneficial users of surface water, as required under the Sustainable Groundwater Management Act (SGMA). SGMA seeks to achieve sustainability, which is defined as the absence of several undesirable results, including “depletions of interconnected surface water that have significant and unreasonable adverse impacts on beneficial users of surface water” (Water Code §10721). The Nature Conservancy (TNC) is a science-based, nonprofit organization with a mission to conserve the lands and waters on which all life depends. Like humans, plants and animals often rely on groundwater for survival, which is why TNC helped develop, and is now helping to implement, SGMA. Earlier this year, we launched the Groundwater Resource Hub, which is an online resource intended to help make it easier and cheaper to address environmental requirements under SGMA. As a first step in addressing when depletions might have an adverse impact, The Nature Conservancy recommends identifying the beneficial users of surface water, which include environmental users. This is a critical step, as it is impossible to define “significant and unreasonable adverse impacts” without knowing what is being impacted. To make this easy, we are providing this letter and the accompanying documents as the best available science on the freshwater species within the boundary of your groundwater sustainability agency (GSA). -

Butterflies of North America
Insects of Western North America 7. Survey of Selected Arthropod Taxa of Fort Sill, Comanche County, Oklahoma. 4. Hexapoda: Selected Coleoptera and Diptera with cumulative list of Arthropoda and additional taxa Contributions of the C.P. Gillette Museum of Arthropod Diversity Colorado State University, Fort Collins, CO 80523-1177 2 Insects of Western North America. 7. Survey of Selected Arthropod Taxa of Fort Sill, Comanche County, Oklahoma. 4. Hexapoda: Selected Coleoptera and Diptera with cumulative list of Arthropoda and additional taxa by Boris C. Kondratieff, Luke Myers, and Whitney S. Cranshaw C.P. Gillette Museum of Arthropod Diversity Department of Bioagricultural Sciences and Pest Management Colorado State University, Fort Collins, Colorado 80523 August 22, 2011 Contributions of the C.P. Gillette Museum of Arthropod Diversity. Department of Bioagricultural Sciences and Pest Management Colorado State University, Fort Collins, CO 80523-1177 3 Cover Photo Credits: Whitney S. Cranshaw. Females of the blow fly Cochliomyia macellaria (Fab.) laying eggs on an animal carcass on Fort Sill, Oklahoma. ISBN 1084-8819 This publication and others in the series may be ordered from the C.P. Gillette Museum of Arthropod Diversity, Department of Bioagricultural Sciences and Pest Management, Colorado State University, Fort Collins, Colorado, 80523-1177. Copyrighted 2011 4 Contents EXECUTIVE SUMMARY .............................................................................................................7 SUMMARY AND MANAGEMENT CONSIDERATIONS -

Life History of Neoplea Striola (Hemiptera: Pleidae)
The Great Lakes Entomologist Volume 19 Number 4 - Winter 1986 Number 4 - Winter Article 5 1986 December 1986 Life History of Neoplea Striola (Hemiptera: Pleidae) J. E. McPherson Southern Illinois University Follow this and additional works at: https://scholar.valpo.edu/tgle Part of the Entomology Commons Recommended Citation McPherson, J. E. 1986. "Life History of Neoplea Striola (Hemiptera: Pleidae)," The Great Lakes Entomologist, vol 19 (4) Available at: https://scholar.valpo.edu/tgle/vol19/iss4/5 This Peer-Review Article is brought to you for free and open access by the Department of Biology at ValpoScholar. It has been accepted for inclusion in The Great Lakes Entomologist by an authorized administrator of ValpoScholar. For more information, please contact a ValpoScholar staff member at [email protected]. McPherson: Life History of <i>Neoplea Striola</i> (Hemiptera: Pleidae) 1986 THE GREAT LAKES ENTOMOLOGIST 217 LIFE HISTORY OF NEOPLEA STRIOLA (HEMIPTERA: PLEIDAE) J. E. McPherson I ABSTRACT The life history of Neop/ea striola was studied in southern Illinois during 1983-1985. This apparently univoltine species overwintered as adults and became active in early March. First instars appeared in mid-May followed by marked overlapping of the subsequent instars. No active adults were found after November. The pygmy backswimmer Neop/ea striola (Fieber) occurs in America north of Mexico from Florida north to Ontario, west to Manitoba, south to Texas, and east to Florida (Sanderson 1982); it has also been reported from Utah (Drake and Chapman 1953). In Illinois. it ha'i been recorded only from the northern 112 of the state (Lauck 1959). -

An Overview of the Heteroptera of Illinois
The Great Lakes Entomologist Volume 22 Number 4 - Winter 1989 Number 4 - Winter Article 1 1989 December 1989 An Overview of the Heteroptera of Illinois J. E. McPherson Southern Illinois University Follow this and additional works at: https://scholar.valpo.edu/tgle Part of the Entomology Commons Recommended Citation McPherson, J. E. 1989. "An Overview of the Heteroptera of Illinois," The Great Lakes Entomologist, vol 22 (4) Available at: https://scholar.valpo.edu/tgle/vol22/iss4/1 This Peer-Review Article is brought to you for free and open access by the Department of Biology at ValpoScholar. It has been accepted for inclusion in The Great Lakes Entomologist by an authorized administrator of ValpoScholar. For more information, please contact a ValpoScholar staff member at [email protected]. McPherson: An Overview of the Heteroptera of Illinois 1989 THE GREAT LAKES ENTOMOLOGIST 177 AN OVERVIEW OF THE HETEROPTERA OF ILLINOIS l J. E. McPherson ,2 ABSTRACT A key to adults of all heteropteran families known to occur in Illinois is presented together with general information on the biologies of these families. Also included are general references on Heteroptera and on individual families, particularly if those references involve studies of fauna that were conducted in Illinois, adjacent states, or nearby parts of Canada. The Heteroptera (true bugs) is a large insect order that occurs worldwide and is represented in America north of Mexico by about 45 families. Of these, 36 are known to occur in Illinois. The order is a well defined group characterized by (1) a segmented beak that arises from the front of the head and (2) wings that, when present and well developed, lie flat on the abdomen with the first pair usually leathery basally and membranous distally. -
The Pleidae (Hemiptera, Heteroptera) of Thailand, with the Descriptions of Two New Species and a Discussion of Species from Southeast Asia
ZooKeys 973: 35–68 (2020) A peer-reviewed open-access journal doi: 10.3897/zookeys.973.54026 RESEARCH ARTICLE https://zookeys.pensoft.net Launched to accelerate biodiversity research The Pleidae (Hemiptera, Heteroptera) of Thailand, with the descriptions of two new species and a discussion of species from Southeast Asia Jerry L. Cook1, Robert W. Sites2, Akekawat Vitheepradit3 1 Department of Biological Sciences, Sam Houston State University, Huntsville, TX 77341, USA 2 Enns Ento- mology Museum, University of Missouri, Columbia, MO 65211, USA 3 Department of Entomology, Kasetsart University, Bangkok 10900 Thailand Corresponding author: Jerry L. Cook ([email protected]) Academic editor: L. Livermore | Received 7 May 2020 | Accepted 2 July 2020 | Published 5 October 2020 http://zoobank.org/0D8FEF96-BC1B-437E-81AB-D70BFCDB314A Citation: Cook JL, Sites RW, Vitheepradit A (2020) The Pleidae (Hemiptera, Heteroptera) of Thailand, with the descriptions of two new species and a discussion of species from Southeast Asia. ZooKeys 973: 35–68. https://doi. org/10.3897/zookeys.973.54026 Abstract The family Pleidae is represented in Thailand by four species in the genus Paraplea. Two of these species, P. frontalis and P. liturata, are widespread and relatively common in Southeast Asia. Two other species, P. lateromaculata Cook, sp. nov. and P. melanodera Cook, sp. nov., are described and only known from Thai- land. Full descriptions are provided for all four species. The distributions of these species are discussed, with an emphasis on Thailand.Paraplea frontalis, P. liturata, and P. lateromaculata Cook, sp. nov. are relatively widespread within Thailand and have overlapping distributions whereasP. melanodera Cook, sp.