Aberrant Sialylation in Cancer: Biomarker and Potential Target for Therapeutic Intervention?
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Screening and Identification of Key Biomarkers in Clear Cell Renal Cell Carcinoma Based on Bioinformatics Analysis
bioRxiv preprint doi: https://doi.org/10.1101/2020.12.21.423889; this version posted December 23, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Screening and identification of key biomarkers in clear cell renal cell carcinoma based on bioinformatics analysis Basavaraj Vastrad1, Chanabasayya Vastrad*2 , Iranna Kotturshetti 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karanataka, India. 3. Department of Ayurveda, Rajiv Gandhi Education Society`s Ayurvedic Medical College, Ron, Karnataka 562209, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India bioRxiv preprint doi: https://doi.org/10.1101/2020.12.21.423889; this version posted December 23, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract Clear cell renal cell carcinoma (ccRCC) is one of the most common types of malignancy of the urinary system. The pathogenesis and effective diagnosis of ccRCC have become popular topics for research in the previous decade. In the current study, an integrated bioinformatics analysis was performed to identify core genes associated in ccRCC. An expression dataset (GSE105261) was downloaded from the Gene Expression Omnibus database, and included 26 ccRCC and 9 normal kideny samples. Assessment of the microarray dataset led to the recognition of differentially expressed genes (DEGs), which was subsequently used for pathway and gene ontology (GO) enrichment analysis. -

Download (PDF)
Tomono et al.: Glycan evolution based on phylogenetic profiling 1 Supplementary Table S1. List of 173 enzymes that are composed of glycosyltransferases and functionally-linked glycan synthetic enzymes UniProt ID Protein Name Categories of Glycan Localization CAZy Class 1 Q8N5D6 Globoside -1,3-N -acetylgalactosaminyltransferase 1 Glycosphingolipid Golgi apparatus GT6 P16442 Histo-blood group ABO system transferase Glycosphingolipid Golgi apparatus GT6 P19526 Galactoside 2--L-fucosyltransferase 1 Glycosphingolipid Golgi apparatus GT11 Q10981 Galactoside 2--L-fucosyltransferase 2 Glycosphingolipid Golgi apparatus GT11 Q00973 -1,4 N -acetylgalactosaminyltransferase 1 Glycosphingolipid Golgi apparatus GT12 Q8NHY0 -1,4 N -acetylgalactosaminyltransferase 2 O -Glycan, N -Glycan, Glycosphingolipid Golgi apparatus GT12 Q09327 -1,4-mannosyl-glycoprotein 4--N -acetylglucosaminyltransferase N -Glycan Golgi apparatus GT17 Q09328 -1,6-mannosylglycoprotein 6--N -acetylglucosaminyltransferase A N -Glycan Golgi apparatus GT18 Q3V5L5 -1,6-mannosylglycoprotein 6--N -acetylglucosaminyltransferase B O -Glycan, N -Glycan Golgi apparatus GT18 Q92186 -2,8-sialyltransferase 8B (SIAT8-B) (ST8SiaII) (STX) N -Glycan Golgi apparatus GT29 O15466 -2,8-sialyltransferase 8E (SIAT8-E) (ST8SiaV) Glycosphingolipid Golgi apparatus GT29 P61647 -2,8-sialyltransferase 8F (SIAT8-F) (ST8SiaVI) O -Glycan Golgi apparatus GT29 Q9NSC7 -N -acetylgalactosaminide -2,6-sialyltransferase 1 (ST6GalNAcI) (SIAT7-A) O -Glycan Golgi apparatus GT29 Q9UJ37 -N -acetylgalactosaminide -2,6-sialyltransferase -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Two Arabidopsis Proteins Synthesize Acetylated Xylan Invitro
The Plant Journal (2014) 80, 197–206 doi: 10.1111/tpj.12643 FEATURED ARTICLE Two Arabidopsis proteins synthesize acetylated xylan in vitro Breeanna R. Urbanowicz, Maria J. Pena*,~ Heather A. Moniz, Kelley W. Moremen and William S. York* Complex Carbohydrate Research Center, University of Georgia, 315 Riverbend Road, Athens, GA 30602, USA Received 4 June 2014; revised 18 July 2014; accepted 1 August 2014; published online 21 August 2014. *For correspondence (e-mails [email protected]; [email protected]). SUMMARY Xylan is the third most abundant glycopolymer on earth after cellulose and chitin. As a major component of wood, grain and forage, this natural biopolymer has far-reaching impacts on human life. This highly acetylated cell wall polysaccharide is a vital component of the plant cell wall, which functions as a molecular scaffold, pro- viding plants with mechanical strength and flexibility. Mutations that impair synthesis of the xylan backbone give rise to plants that fail to grow normally because of collapsed xylem cells in the vascular system. Phenotypic analysis of these mutants has implicated many proteins in xylan biosynthesis; however, the enzymes directly responsible for elongation and acetylation of the xylan backbone have not been unambiguously identified. Here we provide direct biochemical evidence that two Arabidopsis thaliana proteins, IRREGULAR XYLEM 10–L (IRX10-L) and ESKIMO1/TRICOME BIREFRINGENCE 29 (ESK1/TBL29), catalyze these respective processes in vi- tro. By identifying the elusive xylan synthase and establishing ESK1/TBL29 as the archetypal plant polysaccha- ride O-acetyltransferase, we have resolved two long-standing questions in plant cell wall biochemistry. -

Multiplexed Engineering Glycosyltransferase Genes in CHO Cells Via Targeted Integration for Producing Antibodies with Diverse Complex‑Type N‑Glycans Ngan T
www.nature.com/scientificreports OPEN Multiplexed engineering glycosyltransferase genes in CHO cells via targeted integration for producing antibodies with diverse complex‑type N‑glycans Ngan T. B. Nguyen, Jianer Lin, Shi Jie Tay, Mariati, Jessna Yeo, Terry Nguyen‑Khuong & Yuansheng Yang* Therapeutic antibodies are decorated with complex‑type N‑glycans that signifcantly afect their biodistribution and bioactivity. The N‑glycan structures on antibodies are incompletely processed in wild‑type CHO cells due to their limited glycosylation capacity. To improve N‑glycan processing, glycosyltransferase genes have been traditionally overexpressed in CHO cells to engineer the cellular N‑glycosylation pathway by using random integration, which is often associated with large clonal variations in gene expression levels. In order to minimize the clonal variations, we used recombinase‑mediated‑cassette‑exchange (RMCE) technology to overexpress a panel of 42 human glycosyltransferase genes to screen their impact on antibody N‑linked glycosylation. The bottlenecks in the N‑glycosylation pathway were identifed and then released by overexpressing single or multiple critical genes. Overexpressing B4GalT1 gene alone in the CHO cells produced antibodies with more than 80% galactosylated bi‑antennary N‑glycans. Combinatorial overexpression of B4GalT1 and ST6Gal1 produced antibodies containing more than 70% sialylated bi‑antennary N‑glycans. In addition, antibodies with various tri‑antennary N‑glycans were obtained for the frst time by overexpressing MGAT5 alone or in combination with B4GalT1 and ST6Gal1. The various N‑glycan structures and the method for producing them in this work provide opportunities to study the glycan structure‑and‑function and develop novel recombinant antibodies for addressing diferent therapeutic applications. -

Induced Structural Changes in a Multifunctional Sialyltransferase
Biochemistry 2006, 45, 2139-2148 2139 Cytidine 5′-Monophosphate (CMP)-Induced Structural Changes in a Multifunctional Sialyltransferase from Pasteurella multocida†,‡ Lisheng Ni,§ Mingchi Sun,§ Hai Yu,§ Harshal Chokhawala,§ Xi Chen,*,§ and Andrew J. Fisher*,§,| Department of Chemistry and the Section of Molecular and Cellular Biology, UniVersity of California, One Shields AVenue, DaVis, California 95616 ReceiVed NoVember 23, 2005; ReVised Manuscript ReceiVed December 19, 2005 ABSTRACT: Sialyltransferases catalyze reactions that transfer a sialic acid from CMP-sialic acid to an acceptor (a structure terminated with galactose, N-acetylgalactosamine, or sialic acid). They are key enzymes that catalyze the synthesis of sialic acid-containing oligosaccharides, polysaccharides, and glycoconjugates that play pivotal roles in many critical physiological and pathological processes. The structures of a truncated multifunctional Pasteurella multocida sialyltransferase (∆24PmST1), in the absence and presence of CMP, have been determined by X-ray crystallography at 1.65 and 2.0 Å resolutions, respectively. The ∆24PmST1 exists as a monomer in solution and in crystals. Different from the reported crystal structure of a bifunctional sialyltransferase CstII that has only one Rossmann domain, the overall structure of the ∆24PmST1 consists of two separate Rossmann nucleotide-binding domains. The ∆24PmST1 structure, thus, represents the first sialyltransferase structure that belongs to the glycosyltransferase-B (GT-B) structural group. Unlike all other known GT-B structures, however, there is no C-terminal extension that interacts with the N-terminal domain in the ∆24PmST1 structure. The CMP binding site is located in the deep cleft between the two Rossmann domains. Nevertheless, the CMP only forms interactions with residues in the C-terminal domain. -

Sialyltransferase of the 13762 Rat Mammary Ascites Tumor Cells1
[CANCER RESEARCH 44, 1148-1152, March 1984] Sialyltransferase of the 13762 Rat Mammary Ascites Tumor Cells1 ThérèsePrattand Anne P. Sherblom2 Department of Biochemistry, University of Maine, Orano, Maine 04469 ABSTRACT The MAT-B1 and MAT-C1 sublines of the 13762 rat mammary adenocarcinoma are a suitable system for studying sialic acid The MAT-B1 and MAT-C1 ascites sublines of the 13762 rat metabolism. The 2 cell lines, originally derived from the same mammary adenocarcinoma differ in morphology, agglutinability solid tumor, show marked differences in ability to be transplanted with concanavalin A, and xenotransplantability. Both cell lines into mice, agglutinability with concanavalin A, and total sialic acid contain a major mucin-type glycoprotein, but the MAT-C1 (xen- content (19). Greater than 70% of the protein-bound sialic acid otransplantable) subline contains a 3-fold-greater content of sialic in both cell lines is due to a high-molecular-weight mucin-type acid on the glycoprotein than does the MAT-B1 (nonxeno- glycoprotein, ASGP-1 (16). The 0-linked chains have a core transplantable) subline. structure Gal(01-»4)GlcNAc(01-»6)[Gal(|31-»3)]GalNAc3 where The present work indicates that whole cells of both lines both galactose residues may be substituted with sialic acids incorporate radioactivity from labeled CMP-sialic acid into a linked («2—>3).4TheMAT-C1 subline contains much more of component which comigrates with the major glycoprotein by disialylated hexasaccharide than does the MAT-B1 subline,4 sodium dodecyl sulfate polyacrylamide gel electrophoresis, and whereas the MAT-B1 oligosaccharides are predominantly neutral that label incorporated by MAT-B1 cells is released by alkaline- but may contain sulfate as well as sialic acid (17). -

Downloaded 18 July 2014 with a 1% False Discovery Rate (FDR)
UC Berkeley UC Berkeley Electronic Theses and Dissertations Title Chemical glycoproteomics for identification and discovery of glycoprotein alterations in human cancer Permalink https://escholarship.org/uc/item/0t47b9ws Author Spiciarich, David Publication Date 2017 Peer reviewed|Thesis/dissertation eScholarship.org Powered by the California Digital Library University of California Chemical glycoproteomics for identification and discovery of glycoprotein alterations in human cancer by David Spiciarich A dissertation submitted in partial satisfaction of the requirements for the degree Doctor of Philosophy in Chemistry in the Graduate Division of the University of California, Berkeley Committee in charge: Professor Carolyn R. Bertozzi, Co-Chair Professor David E. Wemmer, Co-Chair Professor Matthew B. Francis Professor Amy E. Herr Fall 2017 Chemical glycoproteomics for identification and discovery of glycoprotein alterations in human cancer © 2017 by David Spiciarich Abstract Chemical glycoproteomics for identification and discovery of glycoprotein alterations in human cancer by David Spiciarich Doctor of Philosophy in Chemistry University of California, Berkeley Professor Carolyn R. Bertozzi, Co-Chair Professor David E. Wemmer, Co-Chair Changes in glycosylation have long been appreciated to be part of the cancer phenotype; sialylated glycans are found at elevated levels on many types of cancer and have been implicated in disease progression. However, the specific glycoproteins that contribute to cell surface sialylation are not well characterized, specifically in bona fide human cancer. Metabolic and bioorthogonal labeling methods have previously enabled enrichment and identification of sialoglycoproteins from cultured cells and model organisms. The goal of this work was to develop technologies that can be used for detecting changes in glycoproteins in clinical models of human cancer. -

LN-EPC Vs CEPC List
Supplementary Information Table 5. List of genes upregulated on LN-EPC (LCB represents the variation of gene expression comparing LN-EPC with CEPC) Gene dystrophin (muscular dystrophy, Duchenne and Becker types) regulator of G-protein signalling 13 chemokine (C-C motif) ligand 8 vascular cell adhesion molecule 1 matrix metalloproteinase 9 (gelatinase B, 92kDa gelatinase, 92kDa type IV collagenase) chemokine (C-C motif) ligand 2 solute carrier family 2 (facilitated glucose/fructose transporter), member 5 eukaryotic translation initiation factor 1A, Y-linked regulator of G-protein signalling 1 ubiquitin D chemokine (C-X-C motif) ligand 3 transcription factor 4 chemokine (C-X-C motif) ligand 13 (B-cell chemoattractant) solute carrier family 7, (cationic amino acid transporter, y+ system) member 11 transcription factor 4 apolipoprotein D RAS guanyl releasing protein 3 (calcium and DAG-regulated) matrix metalloproteinase 1 (interstitial collagenase) DEAD (Asp-Glu-Ala-Asp) box polypeptide 3, Y-linked /// DEAD (Asp-Glu-Ala-Asp) box polypeptide 3, Y-linked transcription factor 4 regulator of G-protein signalling 1 B-cell linker interleukin 8 POU domain, class 2, associating factor 1 CD24 antigen (small cell lung carcinoma cluster 4 antigen) Consensus includes gb:AK000168.1 /DEF=Homo sapiens cDNA FLJ20161 fis, clone COL09252, highly similar to L33930 Homo sapiens CD24 signal transducer mRNA. /FEA=mRNA /DB_XREF=gi:7020079 /UG=Hs.332045 Homo sapiens cDNA FLJ20161 fis, clone COL09252, highly similar to L33930 Homo sapiens CD24 signal transducer mRNA -
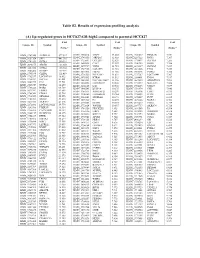
(A) Up-Regulated Genes in HCC827-GR-High2 Compared to Parental HCC827
Table S2. Results of expression profiling analysis (A) Up-regulated genes in HCC827-GR-high2 compared to parental HCC827 Fold Fold Fold Unique ID Symbol Unique ID Symbol Unique ID Symbol change* change* change* ILMN_1709348 ALDH1A1 577.587 ILMN_2310814 MAPT 13.003 ILMN_1741017 PIP4K2B 7.331 ILMN_1651354 SPP1 441.316 ILMN_1748650 MRPL45 12.988 ILMN_3237623 RNY1 7.297 ILMN_1701831 GSTA1 260.591 ILMN_1755897 UGT2B7 12.629 ILMN_1734897 SLC4A4 7.285 ILMN_1658835 CAV2 12.357 ILMN_1746359 RERG 7.280 ILMN_2094875 ABCB1 183.050 ILMN_1678939 VNN2 11.935 ILMN_1671337 SLC2A5 7.257 ILMN_3251540 GSTA2 145.982 ILMN_1729905 GAL3ST1 11.910 ILMN_1691606 LYG2 7.254 ILMN_2062468 IGFBP7 127.721 ILMN_1672536 FBLN1 11.716 ILMN_1785646 PMP22 7.246 ILMN_1795190 CLDN2 111.439 ILMN_1796339 PLEKHA2 11.631 ILMN_1737387 LOC728441 7.207 ILMN_1782937 LOC647169 98.612 ILMN_1676563 HTRA1 11.592 ILMN_1684401 FMO1 7.117 ILMN_1754247 SLC3A1 81.001 ILMN_3263423 LOC100129027 11.346 ILMN_1687035 ADAMTSL4 7.098 ILMN_1662795 CA2 79.581 ILMN_1694898 LOC653857 10.906 ILMN_2153572 MAGEA3 7.086 ILMN_2168747 GSTA2 66.250 ILMN_2404625 LAT 10.560 ILMN_1784283 USH1C 7.079 ILMN_1764228 DAB2 64.709 ILMN_1666546 DUSP14 10.375 ILMN_1731374 CPE 7.046 ILMN_1675797 EPDR1 63.605 ILMN_1764571 ARHGAP23 10.299 ILMN_1765446 EMP3 6.933 ILMN_1708341 PDZK1 59.714 ILMN_3200140 LOC645638 10.284 ILMN_1754002 IL1F8 6.863 ILMN_1713529 SEMA6A 52.575 ILMN_3244343 SNORA21 10.171 ILMN_1878007 FUT9 6.835 ILMN_1708391 NR1H4 43.218 ILMN_1671489 PC 10.075 ILMN_1699208 NAP1L1 6.763 ILMN_2412336 AKR1C2 42.826 ILMN_2404688 -

Supplementary Table 1
Supplementary Table 1. 492 genes are unique to 0 h post-heat timepoint. The name, p-value, fold change, location and family of each gene are indicated. Genes were filtered for an absolute value log2 ration 1.5 and a significance value of p ≤ 0.05. Symbol p-value Log Gene Name Location Family Ratio ABCA13 1.87E-02 3.292 ATP-binding cassette, sub-family unknown transporter A (ABC1), member 13 ABCB1 1.93E-02 −1.819 ATP-binding cassette, sub-family Plasma transporter B (MDR/TAP), member 1 Membrane ABCC3 2.83E-02 2.016 ATP-binding cassette, sub-family Plasma transporter C (CFTR/MRP), member 3 Membrane ABHD6 7.79E-03 −2.717 abhydrolase domain containing 6 Cytoplasm enzyme ACAT1 4.10E-02 3.009 acetyl-CoA acetyltransferase 1 Cytoplasm enzyme ACBD4 2.66E-03 1.722 acyl-CoA binding domain unknown other containing 4 ACSL5 1.86E-02 −2.876 acyl-CoA synthetase long-chain Cytoplasm enzyme family member 5 ADAM23 3.33E-02 −3.008 ADAM metallopeptidase domain Plasma peptidase 23 Membrane ADAM29 5.58E-03 3.463 ADAM metallopeptidase domain Plasma peptidase 29 Membrane ADAMTS17 2.67E-04 3.051 ADAM metallopeptidase with Extracellular other thrombospondin type 1 motif, 17 Space ADCYAP1R1 1.20E-02 1.848 adenylate cyclase activating Plasma G-protein polypeptide 1 (pituitary) receptor Membrane coupled type I receptor ADH6 (includes 4.02E-02 −1.845 alcohol dehydrogenase 6 (class Cytoplasm enzyme EG:130) V) AHSA2 1.54E-04 −1.6 AHA1, activator of heat shock unknown other 90kDa protein ATPase homolog 2 (yeast) AK5 3.32E-02 1.658 adenylate kinase 5 Cytoplasm kinase AK7 -

Protein & Peptide Letters
696 Send Orders for Reprints to [email protected] Protein & Peptide Letters, 2017, 24, 696-709 REVIEW ARTICLE ISSN: 0929-8665 eISSN: 1875-5305 Impact Factor: 1.068 Glycan Phosphorylases in Multi-Enzyme Synthetic Processes BENTHAM Editor-in-Chief: SCIENCE Ben M. Dunn Giulia Pergolizzia, Sakonwan Kuhaudomlarpa, Eeshan Kalitaa,b and Robert A. Fielda,* aDepartment of Biological Chemistry, John Innes Centre, Norwich Research Park, Norwich NR4 7UH, UK; bDepartment of Molecular Biology and Biotechnology, Tezpur University, Napaam, Tezpur, Assam -784028, India Abstract: Glycoside phosphorylases catalyse the reversible synthesis of glycosidic bonds by glyco- A R T I C L E H I S T O R Y sylation with concomitant release of inorganic phosphate. The equilibrium position of such reac- tions can render them of limited synthetic utility, unless coupled with a secondary enzymatic step Received: January 17, 2017 Revised: May 24, 2017 where the reaction lies heavily in favour of product. This article surveys recent works on the com- Accepted: June 20, 2017 bined use of glycan phosphorylases with other enzymes to achieve synthetically useful processes. DOI: 10.2174/0929866524666170811125109 Keywords: Phosphorylase, disaccharide, α-glucan, cellodextrin, high-value products, biofuel. O O 1. INTRODUCTION + HO OH Glycoside phosphorylases (GPs) are carbohydrate-active GH enzymes (CAZymes) (URL: http://www.cazy.org/) [1] in- H2O O GP volved in the formation/cleavage of glycosidic bond together O O GT O O + HO O + HO with glycosyltransferase (GT) and glycoside hydrolase (GH) O -- NDP OPO3 NDP -- families (Figure 1) [2-6]. GT reactions favour the thermody- HPO4 namically more stable glycoside product [7]; however, these GS R enzymes can be challenging to work with because of their O O + HO current limited availability and relative instability, along R with the expense of sugar nucleotide substrates [7].