Phylogenetic Analyses and Comparative Genomics of Vitamin B6 (Pyridoxine) and Pyridoxal Phosphate Biosynthesis Pathways
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Genome-Resolved Meta-Analysis of the Microbiome in Oil Reservoirs Worldwide
microorganisms Article Genome-Resolved Meta-Analysis of the Microbiome in Oil Reservoirs Worldwide Kelly J. Hidalgo 1,2,* , Isabel N. Sierra-Garcia 3 , German Zafra 4 and Valéria M. de Oliveira 1 1 Microbial Resources Division, Research Center for Chemistry, Biology and Agriculture (CPQBA), University of Campinas–UNICAMP, Av. Alexandre Cazellato 999, 13148-218 Paulínia, Brazil; [email protected] 2 Graduate Program in Genetics and Molecular Biology, Institute of Biology, University of Campinas (UNICAMP), Rua Monteiro Lobato 255, Cidade Universitária, 13083-862 Campinas, Brazil 3 Biology Department & CESAM, University of Aveiro, Aveiro, Portugal, Campus de Santiago, Avenida João Jacinto de Magalhães, 3810-193 Aveiro, Portugal; [email protected] 4 Grupo de Investigación en Bioquímica y Microbiología (GIBIM), Escuela de Microbiología, Universidad Industrial de Santander, Cra 27 calle 9, 680002 Bucaramanga, Colombia; [email protected] * Correspondence: [email protected]; Tel.: +55-19981721510 Abstract: Microorganisms inhabiting subsurface petroleum reservoirs are key players in biochemical transformations. The interactions of microbial communities in these environments are highly complex and still poorly understood. This work aimed to assess publicly available metagenomes from oil reservoirs and implement a robust pipeline of genome-resolved metagenomics to decipher metabolic and taxonomic profiles of petroleum reservoirs worldwide. Analysis of 301.2 Gb of metagenomic information derived from heavily flooded petroleum reservoirs in China and Alaska to non-flooded petroleum reservoirs in Brazil enabled us to reconstruct 148 metagenome-assembled genomes (MAGs) of high and medium quality. At the phylum level, 74% of MAGs belonged to bacteria and 26% to archaea. The profiles of these MAGs were related to the physicochemical parameters and recovery management applied. -

Archaeoglobus Profundus Type Strain (AV18T)
Standards in Genomic Sciences (2010) 2:327-346 DOI:10.4056/sigs.942153 Complete genome sequence of Archaeoglobus profundus type strain (AV18T) Mathias von Jan1, Alla Lapidus2, Tijana Glavina Del Rio2, Alex Copeland2, Hope Tice2, Jan-Fang Cheng2, Susan Lucas2, Feng Chen2, Matt Nolan2, Lynne Goodwin2,3, Cliff Han2,3, Sam Pitluck2, Konstantinos Liolios2, Natalia Ivanova2, Konstantinos Mavromatis2, Galina Ovchinnikova2, Olga Chertkov2, Amrita Pati2, Amy Chen4, Krishna Palaniappan4, Miriam Land2,5, Loren Hauser2,5, Yun-Juan Chang2,5, Cynthia D. Jeffries2,5, Elizabeth Saunders2, Thomas Brettin2,3, John C. Detter2,3, Patrick Chain2,4, Konrad Eichinger6, Harald Huber6, Ste- fan Spring1, Manfred Rohde7, Markus Göker1, Reinhard Wirth6, Tanja Woyke2, Jim Bristow2, Jonathan A. Eisen2,8, Victor Markowitz4, Philip Hugenholtz2, Nikos C Kyrpides2, and Hans-Peter Klenk1* 1 DSMZ - German Collection of Microorganisms and Cell Cultures GmbH, Braunschweig, Germany 2 DOE Joint Genome Institute, Walnut Creek, California, USA 3 Los Alamos National Laboratory, Bioscience Division, Los Alamos, New Mexico, USA 4 Biological Data Management and Technology Center, Lawrence Berkeley National Laboratory, Berkeley, California, USA 5 Oak Ridge National Laboratory, Oak Ridge, Tennessee, USA 6 University of Regensburg, Microbiology – Archaeenzentrum, Regensburg, Germany 7 HZI – Helmholtz Centre for Infection Research, Braunschweig, Germany 8 University of California Davis Genome Center, Davis, California, USA *Corresponding author: Hans-Peter Klenk Keywords: hyperthermophilic, marine, strictly anaerobic, sulfate respiration, hydrogen utili- zation, hydrothermal systems, Archaeoglobaceae, GEBA Archaeoglobus profundus (Burggraf et al. 1990) is a hyperthermophilic archaeon in the eu- ryarchaeal class Archaeoglobi, which is currently represented by the single family Archaeog- lobaceae, containing six validly named species and two strains ascribed to the genus 'Geoglobus' which is taxonomically challenged as the corresponding type species has no va- lidly published name. -
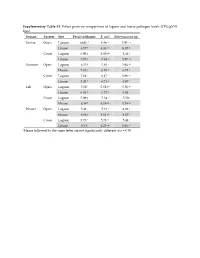
Supplementary Table S1. Fisher Pairwise Comparisons of Lagoon and House Pathogen Levels (CFU/Gvss Log10) Season System Site Fecal Coliforms E
Supplementary Table S1. Fisher pairwise comparisons of lagoon and house pathogen levels (CFU/gVSS log10) Season System Site Fecal coliforms E. coli Enterococcus sp. Spring Open Lagoon 6.00 g1 5.46 hi 5.91 cd House 6.57 e 6.33 cd 6.35 b Cover Lagoon 6.00 g 5.60 gh 5.44 f House 7.73 a 7.63 a 5.87 cd Summer Open Lagoon 6.35 f 5.83 f 5.94 cd House 7.83 a 6.10 e 6.73 a Cover Lagoon 7.04 c 6.47 c 5.86 cd House 7.42 b 6.71 b 6.07 c Fall Open Lagoon 5.58 i 5.58 gh 5.56 ef House 6.81 d 6.72 b 6.01 c Cover Lagoon 5.99 g 5.34 i 5.50 f House 6.38 f 6.19 de 5.74 de Winter Open Lagoon 5.41 j 5.13 j 4.88 g House 6.06 g 5.61 gh 6.67 a Cover Lagoon 5.73 h 5.73 fg 5.44 f House 6.34 f 6.25 de 5.86 cd 1Means followed by the same letter are not significantly different at p = 0.05 Supplementary Table S2. Relative abundances of OTUs identified using universal bacterial primer set, presented as relative abundances (%) Only bacterial families are counted in Figure 5 and discussion involving bacterial family identification. # Identified in # Identified from all 8 All 16 All #OTU Table SpOLRA SuOLRA FOLRA WOLRA SpCLRA SuCLRA FCLRA WCLRA SpOHRA SuOHRA FOHRA WOHRA SpCHRA SuCHRA FCHRA WCHRA all samples lagoon samples Samples Lagoons Notes Key Patulibacteraceae 0 9.93739E‐05* 0.000134 0.000116 0 0 0 0 0 0 0.000452 0 0 0 0 0 4 3 Sp = Spring O = Open Ruaniaceae 0 0 0 0 0 0 0 0 0 0 0 0 0 0.000221577 0 7.73E‐05 2 0 Su = Summer C = Covered Thermoactinomycetaceae 0 0 0.000267 0.000116 0 0.00037092 0.000529 0.000526 0 0.000342922 0.000271 0.000197 0 0 0 0 8 5 F = Fall L = Lagoon Beutenbergiaceae 0.000226334 0.000331246 0.000579 0.001663 0.000620176 0 0.000106 0.000807 0.002320743 0.000571537 0.000723 0.000591 0.000169731 0 0.000606 0 13 7 W = Winter H = House Thermoanaerobacterales Family III. -

Electrical Current Generation in Microbial Electrolysis Cells by Hyperthermophilic Archaea Ferroglobus Placidus and Geoglobus Ahangari
Bioelectrochemistry 119 (2018) 142–149 Contents lists available at ScienceDirect Bioelectrochemistry journal homepage: www.elsevier.com/locate/bioelechem Electrical current generation in microbial electrolysis cells by hyperthermophilic archaea Ferroglobus placidus and Geoglobus ahangari Yasemin D. Yilmazel a,b,⁎, Xiuping Zhu b,c,Kyoung-YeolKimb,DawnE.Holmesd, Bruce E. Logan b a Department of Chemical Engineering, Rochester Institute of Technology, Rochester, NY, USA b Department of Civil and Environmental Engineering, The Pennsylvania State University, University Park, PA, USA c Department of Civil and Environmental Engineering, Louisiana State University, Baton Rouge, LA, USA d Department of Biology, Western New England University, Springfield, MA, USA article info abstract Article history: Few microorganisms have been examined for current generation under thermophilic (40–65 °C) or hyperther- Received 5 May 2017 mophilic temperatures (≥80 °C) in microbial electrochemical systems. Two iron-reducing archaea from the fam- Received in revised form 27 September 2017 ily Archaeoglobaceae, Ferroglobus placidus and Geoglobus ahangari, showed electro-active behavior leading to Accepted 28 September 2017 current generation at hyperthermophilic temperatures in single-chamber microbial electrolysis cells (MECs). A Available online 02 October 2017 current density (j) of 0.68 ± 0.11 A/m2 was attained in F. placidus MECs at 85 °C, and 0.57 ± 0.10 A/m2 in G. ahangari MECs at 80 °C, with an applied voltage of 0.7 V. Cyclic voltammetry (CV) showed that both strains pro- Keywords: − − Hyperthermophilic archaea duced a sigmoidal catalytic wave, with a mid-point potential of 0.39 V (vs. Ag/AgCl) for F. placidus and 0.37 V Ferroglobus placidus for G. -

University of Florida Thesis Or Dissertation Formatting
CHARACTERIZING THE MOLECULAR MECHANISMS CONTRIBUTING TO BIOLOGICALLY INDUCED CARBONATE MINERALIZATION AND THROMBOLITE FORMATION By ARTEMIS S. LOUYAKIS A DISSERTATION PRESENTED TO THE GRADUATE SCHOOL OF THE UNIVERSITY OF FLORIDA IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE OF DOCTOR OF PHILOSOPHY UNIVERSITY OF FLORIDA 2017 © 2017 Artemis S. Louyakis To my mother, for supporting every single goal I’ve ever had, the memory of my father, for keeping me focused, and my partner, for all he’s done ACKNOWLEDGMENTS I would like to begin by acknowledging and thanking my mentor, Dr. Jamie Foster, for all her guidance throughout this Ph.D. I thank my committee members for all of their advice and support - Drs. Eric Triplett, Julie Maupin, Nian Wang, and Eric McLamore. I’d like to thank the rest of the Department of Microbiology and Cell Science, staff for always keeping my academic life in order, faculty for never turning me away when I came to use equipment or ask for help, especially Drs. K.T. Shanmugan and Wayne Nicholson, as well as Dr. Andy Schuerger from the Dept. of Plant Pathology for his advice over the years. I’d also like to acknowledge those lab members and extended lab members who made themselves readily available to talk through any problems I came up against and celebrate when all went well, including Drs. Rafael Oliveira, Jennifer Mobberley, and Giorgio Casaburi, and Lexi Duscher, Rachelle Banjawo, Maddie Vroom, Hadrien Gourlé, and so many more. I’d also like to profusely thank my family and friends who have never been anything less than completely supportive of me, specifically my partner Nathan Prince, my mother and siblings Denise Louyakis, Bobbi Louyakis, Nick Newman, Cori Sergi, extended parents and siblings Carol Prince, Barry Prince, Aaron Prince, my nieces and nephew Bailey O’Regan, Bella O’Regan, Layla Newman, Colton Prince, and Summer Prince, and my dearest friends Tina Pontbriand, Tom Pontbriand, Karen Chan, Dalal Haouchar, Alexi Casaburi, and Eloise Stikeman. -

Systematic Bacteriology Second Edition
BERGEY'S MANUAL® OF Systematic Bacteriology Second Edition Volume One The Archaea and the Deeply Branching and Phototrophic Bacteria Springer New York Berlin Heidelberg Barcelona HongKong London Milan Paris Singapore Tokyo BERGEY'S MANUAL® OF Systematic Bacteriology Second Edition Volume One The Archaea and the Deeply Branching and Phototrophic Bacteria David R. Boone Richard W. Castenholz EDITORS, VOLUME ONE George M. Garrity EDITOR-IN-CHIEF EDITORIAL BOARD James T. Staley, Chairman, David R. Boone, Vice Chairman, Don J. Brenner, Richard W. Castenholz, George M. Garrity, Michael Goodfellow, Noel R. Krieg, Fred A. Rainey, Karl-Heinz Schleifer WITH CONTRIBUTIONS FROM 105 COLLEAGUES Springer George M. Garrity Department of Microbiology and Molecular Genetics Bergey's Manual Trust Michigan State University East Lansing, MI 48824-1101 USA Library of Congress Cataloging-in-Publication Data Bergey's manual of systematic bacteriology / David R. Boone, Richard W. Castenholz, editors, volume 1 ; George M. Garrity, editor-in-chief.-2nd ed. p.cm. Includes bibliographical references and index. Contents: v.I. The archaea and the deeply branching and phototrophic bacteria. ISBN 0-387-98771-1 (alk. paper) 1. Bacteria-Classification. I. Title: Systematic bacteriology. II. Boone, David R. III. Castenholz, Richard W. IV. Garrity, George M. QR81.B462001 579.3'01 '2-dc21 2001020400 With 330 illustrations The following proprietary names of products are used in this volume: Casamino@ acids; Vector NTI®; XLI0-Gold®; Gelrite®; Tryptone@l; Phytagel@; bio-Trypcasew; Trypticasev; Oxoid® purified agar. Printed on acid-free paper. First edition published 1984-1989 by Bergey's Manual Trust and Williams & Wilkins, Baltimore. © 2001 Bergey's Manual Trust All rights reserved. -

1 This Manuscript Is Submitted for Publication in Renewable and Sustainable Energy
1 This manuscript is submitted for publication in Renewable and Sustainable Energy 2 Reviews. Please note that, this manuscript has not undergone peer-review and is not yet 3 formally accepted for publication. Subsequent versions of this manuscript may have 4 slightly different content. If accepted, the final version of this manuscript will be 5 available via the ‘Peer-reviewed Publication DOI’ link on the right-hand side of this 6 webpage. 7 Manuscript Title: Estimating Microbial Hydrogen Consumption in Hydrogen Storage in 8 Porous Media as a Basis for Site Selection 9 Authors 10 Eike Marie Thaysena: [email protected] 11 Sean McMahona, b: [email protected] 12 Gion Strobelc: [email protected] 13 Ian Butlera: [email protected] 14 Bryne Ngwenyaa: [email protected] 15 Niklas Heinemanna: [email protected] 16 Mark Wilkinsona: [email protected] 17 Aliakbar Hassanpouryouzbanda: [email protected] 18 Chris McDermotta: [email protected] 19 Katriona Edlmanna: [email protected] 20 21 Affiliations: 22 a) School of Geosciences, University of Edinburgh, Grant Institute, West Main Road, 23 Edinburgh, EH9 3JW, UK 24 b) School of Physics and Astronomy, James Clerk Maxwell Building, University of 25 Edinburgh, EH9 3FD, United Kingdom 26 c) Department of Petroleum Engineering, Clausthal University of Technology, Germany 27 28 1 29 Estimating Microbial Hydrogen Consumption in 30 Hydrogen Storage in Porous Media as a Basis for 31 Site Selection 32 Eike M. Thaysen1*, Sean McMahon1,3, Gion Strobel2, -

Supplement of Biogeosciences, 16, 4229–4241, 2019 © Author(S) 2019
Supplement of Biogeosciences, 16, 4229–4241, 2019 https://doi.org/10.5194/bg-16-4229-2019-supplement © Author(s) 2019. This work is distributed under the Creative Commons Attribution 4.0 License. Supplement of Identifying the core bacterial microbiome of hydrocarbon degradation and a shift of dominant methanogenesis pathways in the oil and aqueous phases of petroleum reservoirs of different temperatures from China Zhichao Zhou et al. Correspondence to: Ji-Dong Gu ([email protected]) and Bo-Zhong Mu ([email protected]) The copyright of individual parts of the supplement might differ from the CC BY 4.0 License. 1 Supplementary Data 1.1 Characterization of geographic properties of sampling reservoirs Petroleum fluids samples were collected from eight sampling sites across China covering oilfields of different geological properties. The reservoir and crude oil properties together with the aqueous phase chemical concentration characteristics were listed in Table 1. P1 represents the sample collected from Zhan3-26 well located in Shengli Oilfield. Zhan3 block region in Shengli Oilfield is located in the coastal area from the Yellow River Estuary to the Bohai Sea. It is a medium-high temperature reservoir of fluvial face, made of a thin layer of crossed sand-mudstones, pebbled sandstones and fine sandstones. P2 represents the sample collected from Ba-51 well, which is located in Bayindulan reservoir layer of Erlian Basin, east Inner Mongolia Autonomous Region. It is a reservoir with highly heterogeneous layers, high crude oil viscosity and low formation fluid temperature. It was dedicated to water-flooding, however, due to low permeability and high viscosity of crude oil, displacement efficiency of water-flooding driving process was slowed down along the increase of water-cut rate. -

Electrotrophy As Potential Primary Metabolism for Colonization Of
bioRxiv preprint doi: https://doi.org/10.1101/2020.11.11.377697; this version posted November 11, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 Electrotrophy as potential primary metabolism for colonization of 2 conductive surfaces in deep-sea hydrothermal chimneys. 3 Guillaume Pillot1, Sylvain Davidson1, Laetitia Shintu3, Oulfat Amin AliX, Anne Godfroy2, Yannick Combet-Blanc1, Patricia 4 Bonin1, Pierre-Pol Liebgott1* 5 1 Aix Marseille Univ., Université de Toulon, IRD, CNRS, MIO UM 110, 13288, Marseille, France 6 2 IFREMER, CNRS, Université de Bretagne Occidentale, Laboratoire de Microbiologie des Environnements Extrêmes – 7 UMR6197, Ifremer, Centre de Brest CS10070, Plouzané, France. 8 3Aix Marseille Univ, CNRS, Centrale Marseille, iSm2, Marseille, France 9 * To whom correspondence may be addressed. Email: [email protected]; Mediterranean Institute 10 of Oceanography, Campus de Luminy, Bâtiment OCEANOMED, 13288 Marseille Cedex 09. 11 https://orcid.org/0000-0002-2559-1738 12 Key-words 13 Electrotrophy, hyperthermophiles, microbial electrochemical system, deep-sea hydrothermal 14 vent, electrosynthesis 15 Summary 16 Deep-sea hydrothermal vents are extreme and complex ecosystems based on a trophic chain. We 17 are still unsure of the first colonizers of these environments and their metabolism, but they are 18 thought to be (hyper)thermophilic autotrophs. Here we investigate whether the electric potential 19 observed across hydrothermal chimneys could serve as an energy source for these first colonizers. 20 Experiments were performed in a two-chamber microbial electrochemical system inoculated with 21 deep-sea hydrothermal chimney samples, with a cathode as sole electron donor, CO2 as sole 22 carbon source, and three different electron acceptors (nitrate, sulfate, and oxygen). -

Physiological and Genomic Characterization of a Hyperthermophilic Archaeon Archaeoglobus Neptunius Sp
Physiological and Genomic Characterization of a Hyperthermophilic Archaeon Archaeoglobus neptunius sp. nov. Isolated From a Deep-Sea Hydrothermal Vent Warrants the Reclassification of the Genus Archaeoglobus Galina Slobodkina, Maxime Allioux, Alexander Merkel, Marie-Anne Cambon-Bonavita, Karine Alain, Mohamed Jebbar, Alexander Slobodkin To cite this version: Galina Slobodkina, Maxime Allioux, Alexander Merkel, Marie-Anne Cambon-Bonavita, Karine Alain, et al.. Physiological and Genomic Characterization of a Hyperthermophilic Archaeon Ar- chaeoglobus neptunius sp. nov. Isolated From a Deep-Sea Hydrothermal Vent Warrants the Re- classification of the Genus Archaeoglobus. Frontiers in Microbiology, Frontiers Media, 2021,12, 10.3389/fmicb.2021.679245. hal-03322688 HAL Id: hal-03322688 https://hal.archives-ouvertes.fr/hal-03322688 Submitted on 19 Aug 2021 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. ORIGINAL RESEARCH published: 16 July 2021 doi: 10.3389/fmicb.2021.679245 Physiological and Genomic Characterization of a Hyperthermophilic Archaeon Archaeoglobus -

Complete Genome Sequence of Ferroglobus Placidus AEDII12DO
Standards in Genomic Sciences (2011) 5:50-60 DOI:10.4056/sigs.2225018 Complete genome sequence of Ferroglobus placidus AEDII12DO Iain Anderson,1* Carla Risso,2 Dawn Holmes,2 Susan Lucas,1 Alex Copeland,1 Alla Lapidus,1 Jan-Fang Cheng,1 David Bruce,1,3 Lynne Goodwin,1,3 Samuel Pitluck,1 Elizabeth Saunders,1,3 Thomas Brettin,1,3 John C. Detter,1,3 Cliff Han,1,3 Roxanne Tapia,1,3 Frank Larimer,4 Miriam Land,4 Loren Hauser,4 Tanja Woyke,1 Derek Lovley,2 Nikos Kyrpides,1 and Natalia Ivanova1 1DOE Joint Genome Institute, Walnut Creek, California, USA 2Microbiology Department, University of Massachusetts, Amherst, Massachusetts, USA 3Los Alamos National Laboratory, Los Alamos, New Mexico, USA 4Biosciences Division, Oak Ridge National Laboratory, Oak Ridge, Tennessee, USA *Corresponding author: [email protected] Keywords: Archaea, Euryarchaeota, Archaeoglobales, hydrothermal vent, hyperthermophile, anaerobe Ferroglobus placidus belongs to the order Archaeoglobales within the archaeal phylum Euryar- chaeota. Strain AEDII12DO is the type strain of the species and was isolated from a shallow marine hydrothermal system at Vulcano, Italy. It is a hyperthermophilic, anaerobic chemoli- thoautotroph, but it can also use a variety of aromatic compounds as electron donors. Here we describe the features of this organism together with the complete genome sequence and anno- tation. The 2,196,266 bp genome with its 2,567 protein-coding and 55 RNA genes was se- quenced as part of a DOE Joint Genome Institute Laboratory Sequencing Program (LSP) project. Introduction Strain AEDII12DO (=DSM 10642) is the type strain Organism information of the species Ferroglobus placidus. -

Metabolic Roles of Uncultivated Bacterioplankton Lineages in the Northern Gulf of Mexico 2 “Dead Zone” 3 4 J
bioRxiv preprint doi: https://doi.org/10.1101/095471; this version posted June 12, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. 1 Metabolic roles of uncultivated bacterioplankton lineages in the northern Gulf of Mexico 2 “Dead Zone” 3 4 J. Cameron Thrash1*, Kiley W. Seitz2, Brett J. Baker2*, Ben Temperton3, Lauren E. Gillies4, 5 Nancy N. Rabalais5,6, Bernard Henrissat7,8,9, and Olivia U. Mason4 6 7 8 1. Department of Biological Sciences, Louisiana State University, Baton Rouge, LA, USA 9 2. Department of Marine Science, Marine Science Institute, University of Texas at Austin, Port 10 Aransas, TX, USA 11 3. School of Biosciences, University of Exeter, Exeter, UK 12 4. Department of Earth, Ocean, and Atmospheric Science, Florida State University, Tallahassee, 13 FL, USA 14 5. Department of Oceanography and Coastal Sciences, Louisiana State University, Baton Rouge, 15 LA, USA 16 6. Louisiana Universities Marine Consortium, Chauvin, LA USA 17 7. Architecture et Fonction des Macromolécules Biologiques, CNRS, Aix-Marseille Université, 18 13288 Marseille, France 19 8. INRA, USC 1408 AFMB, F-13288 Marseille, France 20 9. Department of Biological Sciences, King Abdulaziz University, Jeddah, Saudi Arabia 21 22 *Correspondence: 23 JCT [email protected] 24 BJB [email protected] 25 26 27 28 Running title: Decoding microbes of the Dead Zone 29 30 31 Abstract word count: 250 32 Text word count: XXXX 33 34 Page 1 of 31 bioRxiv preprint doi: https://doi.org/10.1101/095471; this version posted June 12, 2017.