Lessons from the Proteome and S-Nitrosoproteome in Human Preterm Labor
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Glycoproteomics-Based Signatures for Tumor Subtyping and Clinical Outcome Prediction of High-Grade Serous Ovarian Cancer
ARTICLE https://doi.org/10.1038/s41467-020-19976-3 OPEN Glycoproteomics-based signatures for tumor subtyping and clinical outcome prediction of high-grade serous ovarian cancer Jianbo Pan 1,2,3, Yingwei Hu1,3, Shisheng Sun 1,3, Lijun Chen1, Michael Schnaubelt1, David Clark1, ✉ Minghui Ao1, Zhen Zhang1, Daniel Chan1, Jiang Qian2 & Hui Zhang 1 1234567890():,; Inter-tumor heterogeneity is a result of genomic, transcriptional, translational, and post- translational molecular features. To investigate the roles of protein glycosylation in the heterogeneity of high-grade serous ovarian carcinoma (HGSC), we perform mass spectrometry-based glycoproteomic characterization of 119 TCGA HGSC tissues. Cluster analysis of intact glycoproteomic profiles delineates 3 major tumor clusters and 5 groups of intact glycopeptides. It also shows a strong relationship between N-glycan structures and tumor molecular subtypes, one example of which being the association of fucosylation with mesenchymal subtype. Further survival analysis reveals that intact glycopeptide signatures of mesenchymal subtype are associated with a poor clinical outcome of HGSC. In addition, we study the expression of mRNAs, proteins, glycosites, and intact glycopeptides, as well as the expression levels of glycosylation enzymes involved in glycoprotein biosynthesis pathways in each tumor. The results show that glycoprotein levels are mainly controlled by the expression of their individual proteins, and, furthermore, that the glycoprotein-modifying glycans cor- respond to the protein levels of glycosylation enzymes. The variation in glycan types further shows coordination to the tumor heterogeneity. Deeper understanding of the glycosylation process and glycosylation production in different subtypes of HGSC may provide important clues for precision medicine and tumor-targeted therapy. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

The Biomarkers of Key Mirnas and Target Genes Associated with Acute Myocardial Infarction
The biomarkers of key miRNAs and target genes associated with acute myocardial infarction Qi Wang1, Bingyan Liu2,3, Yuanyong Wang4, Baochen Bai1, Tao Yu3 and Xian–ming Chu1,5 1 Department of Cardiology, The Affiliated hospital of Qingdao University, Qingdao, China 2 School of Basic Medicine, Qingdao University, Qingdao, China 3 Institute for Translational Medicine, Qingdao University, Qingdao, China 4 Department of Thoracic Surgery, Affiliated Hospital of Qingdao University, Qingdao, China 5 Department of Cardiology, The Affiliated Cardiovascular Hospital of Qingdao University, Qingdao, China ABSTRACT Background. Acute myocardial infarction (AMI) is considered one of the most prominent causes of death from cardiovascular disease worldwide. Knowledge of the molecular mechanisms underlying AMI remains limited. Accurate biomarkers are needed to predict the risk of AMI and would be beneficial for managing the incidence rate. The gold standard for the diagnosis of AMI, the cardiac troponin T (cTnT) assay, requires serial testing, and the timing of measurement with respect to symptoms affects the results. As attractive candidate diagnostic biomarkers in AMI, circulating microRNAs (miRNAs) are easily detectable, generally stable and tissue specific. Methods. The Gene Expression Omnibus (GEO) database was used to compare miRNA expression between AMI and control samples, and the interactions between miRNAs and mRNAs were analysed for expression and function. Furthermore, a protein-protein interaction (PPI) network was constructed. The miRNAs identified in the bioinformatic analysis were verified by RT-qPCR in an H9C2 cell line. The miRNAs in plasma samples from patients with AMI (n D 11) and healthy controls (n D 11) were used to construct Submitted 23 December 2019 receiver operating characteristic (ROC) curves to evaluate the clinical prognostic value Accepted 14 April 2020 of the identified miRNAs. -

HNRPH1 (HNRNPH1) (NM 005520) Human Tagged ORF Clone Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for RC201834 HNRPH1 (HNRNPH1) (NM_005520) Human Tagged ORF Clone Product data: Product Type: Expression Plasmids Product Name: HNRPH1 (HNRNPH1) (NM_005520) Human Tagged ORF Clone Tag: Myc-DDK Symbol: HNRNPH1 Synonyms: hnRNPH; HNRPH; HNRPH1 Vector: pCMV6-Entry (PS100001) E. coli Selection: Kanamycin (25 ug/mL) Cell Selection: Neomycin This product is to be used for laboratory only. Not for diagnostic or therapeutic use. View online » ©2021 OriGene Technologies, Inc., 9620 Medical Center Drive, Ste 200, Rockville, MD 20850, US 1 / 6 HNRPH1 (HNRNPH1) (NM_005520) Human Tagged ORF Clone – RC201834 ORF Nucleotide >RC201834 ORF sequence Sequence: Red=Cloning site Blue=ORF Green=Tags(s) TTTTGTAATACGACTCACTATAGGGCGGCCGGGAATTCGTCGACTGGATCCGGTACCGAGGAGATCTGCC GCCGCGATCGCC ATGATGTTGGGCACGGAAGGTGGAGAGGGATTCGTGGTGAAGGTCCGGGGCTTGCCCTGGTCTTGCTCGG CCGATGAAGTGCAGAGGTTTTTTTCTGACTGCAAAATTCAAAATGGGGCTCAAGGTATTCGTTTCATCTA CACCAGAGAAGGCAGACCAAGTGGCGAGGCTTTTGTTGAACTTGAATCAGAAGATGAAGTCAAATTGGCC CTGAAAAAAGACAGAGAAACTATGGGACACAGATATGTTGAAGTATTCAAGTCAAACAACGTTGAAATGG ATTGGGTGTTGAAGCATACTGGTCCAAATAGTCCTGACACGGCCAATGATGGCTTTGTACGGCTTAGAGG ACTTCCCTTTGGATGTAGCAAGGAAGAAATTGTTCAGTTCTTCTCAGGGTTGGAAATCGTGCCAAATGGG ATAACATTGCCGGTGGACTTCCAGGGGAGGAGTACGGGGGAGGCCTTCGTGCAGTTTGCTTCACAGGAAA TAGCTGAAAAGGCTCTAAAGAAACACAAGGAAAGAATAGGGCACAGGTATATTGAAATCTTTAAGAGCAG TAGAGCTGAAGTTAGAACTCATTATGATCCACCACGAAAGCTTATGGCCATGCAGCGGCCAGGTCCTTAT -
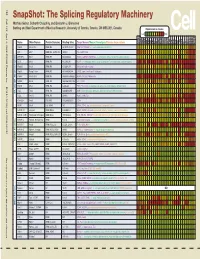
Snapshot: the Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J
192 Cell SnapShot: The Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J. Blencowe 133 Banting and Best Department of Medical Research, University of Toronto, Toronto, ON M5S 3E1, Canada Expression in mouse , April4, 2008©2008Elsevier Inc. Low High Name Other Names Protein Domains Binding Sites Target Genes/Mouse Phenotypes/Disease Associations Amy Ceb Hip Hyp OB Eye SC BM Bo Ht SM Epd Kd Liv Lu Pan Pla Pro Sto Spl Thy Thd Te Ut Ov E6.5 E8.5 E10.5 SRp20 Sfrs3, X16 RRM, RS GCUCCUCUUC SRp20, CT/CGRP; −/− early embryonic lethal E3.5 9G8 Sfrs7 RRM, RS, C2HC Znf (GAC)n Tau, GnRH, 9G8 ASF/SF2 Sfrs1 RRM, RS RGAAGAAC HipK3, CaMKIIδ, HIV RNAs; −/− embryonic lethal, cond. KO cardiomyopathy SC35 Sfrs2 RRM, RS UGCUGUU AChE; −/− embryonic lethal, cond. KO deficient T-cell maturation, cardiomyopathy; LS SRp30c Sfrs9 RRM, RS CUGGAUU Glucocorticoid receptor SRp38 Fusip1, Nssr RRM, RS ACAAAGACAA CREB, type II and type XI collagens SRp40 Sfrs5, HRS RRM, RS AGGAGAAGGGA HipK3, PKCβ-II, Fibronectin SRp55 Sfrs6 RRM, RS GGCAGCACCUG cTnT, CD44 DOI 10.1016/j.cell.2008.03.010 SRp75 Sfrs4 RRM, RS GAAGGA FN1, E1A, CD45; overexpression enhances chondrogenic differentiation Tra2α Tra2a RRM, RS GAAARGARR GnRH; overexpression promotes RA-induced neural differentiation SR and SR-Related Proteins Tra2β Sfrs10 RRM, RS (GAA)n HipK3, SMN, Tau SRm160 Srrm1 RS, PWI AUGAAGAGGA CD44 SWAP Sfrs8 RS, SWAP ND SWAP, CD45, Tau; possible asthma susceptibility gene hnRNP A1 Hnrnpa1 RRM, RGG UAGGGA/U HipK3, SMN2, c-H-ras; rheumatoid arthritis, systemic lupus -

Global Mapping of Herpesvirus-‐Host Protein Complexes Reveals a Novel Transcription
Global mapping of herpesvirus-host protein complexes reveals a novel transcription strategy for late genes By Zoe Hartman Davis A dissertation submitted in partial satisfaction of the Requirements for the degree of Doctor of Philosophy in Infectious Disease and Immunity in the Graduate Division of the University of California, Berkeley Committee in charge: Professor Britt A. Glaunsinger, Chair Professor Laurent Coscoy Professor Qiang Zhou Spring 2015 Abstract Global mapping of herpesvirus-host protein complexes reveals a novel transcription strategy for late genes By Zoe Hartman Davis Doctor of Philosophy in Infectious Diseases and Immunity University of California, Berkeley Professor Britt A. Glaunsinger, Chair Mapping host-pathogen interactions has proven instrumental for understanding how viruses manipulate host machinery and how numerous cellular processes are regulated. DNA viruses such as herpesviruses have relatively large coding capacity and thus can target an extensive network of cellular proteins. To identify the host proteins hijacked by this pathogen, we systematically affinity tagged and purified all 89 proteins of Kaposi’s sarcoma-associated herpesvirus (KSHV) from human cells. Mass spectrometry of this material identified over 500 high-confidence virus-host interactions. KSHV causes AIDS-associated cancers and its interaction network is enriched for proteins linked to cancer and overlaps with proteins that are also targeted by HIV-1. This work revealed many new interactions between viral and host proteins. I have focused on one interaction in particular, that of a previously uncharacterized KSHV protein, ORF24, with cellular RNA polymerase II (RNAP II). All DNA viruses encode a class of genes that are expressed only late in the infectious cycle, following replication of the viral genome. -

Genetic Complexity of Autosomal Dominant Polycystic Kidney and Liver Diseases
BRIEF REVIEW www.jasn.org Genetic Complexity of Autosomal Dominant Polycystic Kidney and Liver Diseases Emilie Cornec-Le Gall,1,2 Vicente E. Torres,1 and Peter C. Harris1 1Division of Nephrology and Hypertension, Mayo Clinic, Rochester, Minnesota; and 2Department of Nephrology, University Hospital, European University of Brittany, and National Institute of Health and Medical Sciences, INSERM U1078, Brest, France ABSTRACT Data indicate significant phenotypic and genotypic overlap, plus a common patho- ADPLD (Table 1).15–20 The difference in genesis, between two groups of inherited disorders, autosomal dominant polycystic renal survival between PKD1 and PKD2 kidney diseases (ADPKD), a significant cause of ESRD, and autosomal dominant patients has been highlighted in multiple polycystic liver diseases (ADPLD), which result in significant PLD with minimal studies (Table 2).3,21 In addition, PKD1 PKD. Eight genes have been associated with ADPKD (PKD1 and PKD2), ADPLD patients have a larger height-adjusted total (PRKCSH, SEC63, LRP5, ALG8,andSEC61B), or both (GANAB). Although genetics kidney volume (HtTKV; an early measure is only infrequently used for diagnosing these diseases and prognosing the associ- of the severity of renal disease in ADPKD) ated outcomes, its value is beginning to be appreciated, and the genomics revolu- and lower eGFR than PKD2 patients.14,22 tion promises more reliable and less expensive molecular diagnostic tools for these A further difference is the number of kid- diseases. We therefore propose categorization of patients with a phenotypic and ney cysts, with fewer in PKD2 than PKD1 genotypic descriptor that will clarify etiology, provide prognostic information, and (Figure 1, A and C), although the rate of better describe atypical cases. -

Individual Protomers of a G Protein-Coupled Receptor Dimer Integrate Distinct Functional Modules
OPEN Citation: Cell Discovery (2015) 1, 15011; doi:10.1038/celldisc.2015.11 © 2015 SIBS, CAS All rights reserved 2056-5968/15 ARTICLE www.nature.com/celldisc Individual protomers of a G protein-coupled receptor dimer integrate distinct functional modules Nathan D Camp1, Kyung-Soon Lee2, Jennifer L Wacker-Mhyre2, Timothy S Kountz2, Ji-Min Park2, Dorathy-Ann Harris2, Marianne Estrada2, Aaron Stewart2, Alejandro Wolf-Yadlin1, Chris Hague2 1Department of Genome Sciences, University of Washington School of Medicine, Seattle, WA, USA; 2Department of Pharmacology, University of Washington School of Medicine, Seattle, WA, USA Recent advances in proteomic technology reveal G-protein-coupled receptors (GPCRs) are organized as large, macromolecular protein complexes in cell membranes, adding a new layer of intricacy to GPCR signaling. We previously reported the α1D-adrenergic receptor (ADRA1D)—a key regulator of cardiovascular, urinary and CNS function—binds the syntrophin family of PDZ domain proteins (SNTA, SNTB1, and SNTB2) through a C-terminal PDZ ligand inter- action, ensuring receptor plasma membrane localization and G-protein coupling. To assess the uniqueness of this novel GPCR complex, 23 human GPCRs containing Type I PDZ ligands were subjected to TAP/MS proteomic analysis. Syntrophins did not interact with any other GPCRs. Unexpectedly, a second PDZ domain protein, scribble (SCRIB), was detected in ADRA1D complexes. Biochemical, proteomic, and dynamic mass redistribution analyses indicate syntrophins and SCRIB compete for the PDZ ligand, simultaneously exist within an ADRA1D multimer, and impart divergent pharmacological properties to the complex. Our results reveal an unprecedented modular dimeric architecture for the ADRA1D in the cell membrane, providing unexpected opportunities for fine-tuning receptor function through novel protein interactions in vivo, and for intervening in signal transduction with small molecules that can stabilize or disrupt unique GPCR:PDZ protein interfaces. -
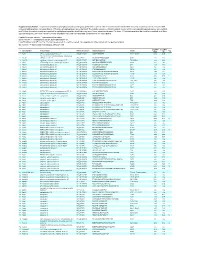
Supplementary Table 1
Supplementary Table 1. Large-scale quantitative phosphoproteomic profiling was performed on paired vehicle- and hormone-treated mTAL-enriched suspensions (n=3). A total of 654 unique phosphopeptides corresponding to 374 unique phosphoproteins were identified. The peptide sequence, phosphorylation site(s), and the corresponding protein name, gene symbol, and RefSeq Accession number are reported for each phosphopeptide identified in any one of three experimental pairs. For those 414 phosphopeptides that could be quantified in all three experimental pairs, the mean Hormone:Vehicle abundance ratio and corresponding standard error are also reported. Peptide Sequence column: * = phosphorylated residue Site(s) column: ^ = ambiguously assigned phosphorylation site Log2(H/V) Mean and SE columns: H = hormone-treated, V = vehicle-treated, n/a = peptide not observable in all 3 experimental pairs Sig. column: * = significantly changed Log 2(H/V), p<0.05 Log (H/V) Log (H/V) # Gene Symbol Protein Name Refseq Accession Peptide Sequence Site(s) 2 2 Sig. Mean SE 1 Aak1 AP2-associated protein kinase 1 NP_001166921 VGSLT*PPSS*PK T622^, S626^ 0.24 0.95 PREDICTED: ATP-binding cassette, sub-family A 2 Abca12 (ABC1), member 12 XP_237242 GLVQVLS*FFSQVQQQR S251^ 1.24 2.13 3 Abcc10 multidrug resistance-associated protein 7 NP_001101671 LMT*ELLS*GIRVLK T464, S468 -2.68 2.48 4 Abcf1 ATP-binding cassette sub-family F member 1 NP_001103353 QLSVPAS*DEEDEVPVPVPR S109 n/a n/a 5 Ablim1 actin-binding LIM protein 1 NP_001037859 PGSSIPGS*PGHTIYAK S51 -3.55 1.81 6 Ablim1 actin-binding -

MALE Protein Name Accession Number Molecular Weight CP1 CP2 H1 H2 PDAC1 PDAC2 CP Mean H Mean PDAC Mean T-Test PDAC Vs. H T-Test
MALE t-test t-test Accession Molecular H PDAC PDAC vs. PDAC vs. Protein Name Number Weight CP1 CP2 H1 H2 PDAC1 PDAC2 CP Mean Mean Mean H CP PDAC/H PDAC/CP - 22 kDa protein IPI00219910 22 kDa 7 5 4 8 1 0 6 6 1 0.1126 0.0456 0.1 0.1 - Cold agglutinin FS-1 L-chain (Fragment) IPI00827773 12 kDa 32 39 34 26 53 57 36 30 55 0.0309 0.0388 1.8 1.5 - HRV Fab 027-VL (Fragment) IPI00827643 12 kDa 4 6 0 0 0 0 5 0 0 - 0.0574 - 0.0 - REV25-2 (Fragment) IPI00816794 15 kDa 8 12 5 7 8 9 10 6 8 0.2225 0.3844 1.3 0.8 A1BG Alpha-1B-glycoprotein precursor IPI00022895 54 kDa 115 109 106 112 111 100 112 109 105 0.6497 0.4138 1.0 0.9 A2M Alpha-2-macroglobulin precursor IPI00478003 163 kDa 62 63 86 72 14 18 63 79 16 0.0120 0.0019 0.2 0.3 ABCB1 Multidrug resistance protein 1 IPI00027481 141 kDa 41 46 23 26 52 64 43 25 58 0.0355 0.1660 2.4 1.3 ABHD14B Isoform 1 of Abhydrolase domain-containing proteinIPI00063827 14B 22 kDa 19 15 19 17 15 9 17 18 12 0.2502 0.3306 0.7 0.7 ABP1 Isoform 1 of Amiloride-sensitive amine oxidase [copper-containing]IPI00020982 precursor85 kDa 1 5 8 8 0 0 3 8 0 0.0001 0.2445 0.0 0.0 ACAN aggrecan isoform 2 precursor IPI00027377 250 kDa 38 30 17 28 34 24 34 22 29 0.4877 0.5109 1.3 0.8 ACE Isoform Somatic-1 of Angiotensin-converting enzyme, somaticIPI00437751 isoform precursor150 kDa 48 34 67 56 28 38 41 61 33 0.0600 0.4301 0.5 0.8 ACE2 Isoform 1 of Angiotensin-converting enzyme 2 precursorIPI00465187 92 kDa 11 16 20 30 4 5 13 25 5 0.0557 0.0847 0.2 0.4 ACO1 Cytoplasmic aconitate hydratase IPI00008485 98 kDa 2 2 0 0 0 0 2 0 0 - 0.0081 - 0.0 -
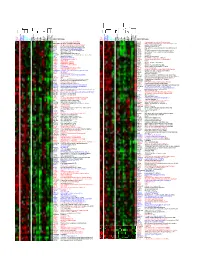
Supporting Figure 8
HUGO ID Name HUGO ID Name Nalm-6 TOM-1 Reh Karpas-422 DoHH-2 SU-DHL-5 Nama lwa DG-75 Ramos Raji BEL EHEB BONNA-12 L-428 DEL BCP-1 BC-3 BCBL-1 JSC-1 PEL -SY HBL-6 DS-1 RPMI-8226 NCI-H929 L-363 SK-MM-2 Nalm-6 TOM-1 Reh Karpas-422 DoHH-2 SU-DHL-5 Nama lwa DG-75 Ramos Raji BEL EHEB BONNA-12 L-428 DEL BCP-1 BC-3 BCBL-1 JSC-1 PEL -SY HBL-6 DS-1 RPMI-8226 NCI-H929 L-363 SK-MM-2 CDC5L Cdc5-related protein (PCDC5RP) RAD51 Replication protein A (E coli RecA homolog) HSU53209 transformer-2 alpha (htra-2 alpha) HMG2 High-mobility group (nonhistone chromosomal) protein 2 CDC10 similar to TR:G560623 G560623 HCDC10 SPS selenium donor protein (selD) NFYB CAAT-box DNA binding protein subunit B CES2 Carboxylestease 2 (liver) GTF2I Bruton's tyrosine kinase-associated protein NOE1 clone 23876 neuronal olfactomedin-related ER localized CCT4 stimulator of TAR RNA binding (SRB) ZNF22 HKR-T1 RY1 RY-1 putative nucleic acid binding protein FKBP5 54 kDa progesterone receptor-associated FKBP54 DVL2 dishevelled 2 (DVL2) FKBP5 54 kDa progesterone receptor-associated FKBP54 RPL13 60S RIBOSOMAL PROTEIN L13 CU L4 A Hs-cul-4A C14ORF3 Hydroxyacyl-Coenzyme A dehydrogenase, beta subuit CU L4 A Hs-cul-4A SM S spermine synthase CLCN6 putative chloride channel SF3A1 splicing factor SF3a120 FALZ fetal Alz-50-reactive clone 1 (FAC1) KNSL4 oriP binding protein (OBP-1) GTF2H3 General transcription factor IIH, polypeptide 3 TUBA1 alpha-tubulin TRIP hT RIP TUBA1 Alpha-tubulin (K00558) GM2A glucose transporter pseudogene TUBA1 Alpha-tubulin (K00558) RBBP6 RBQ-1 TUBA1 TUBULIN ALPHA-4 -

Evolutionary Fate of Retroposed Gene Copies in the Human Genome
Evolutionary fate of retroposed gene copies in the human genome Nicolas Vinckenbosch*, Isabelle Dupanloup*†, and Henrik Kaessmann*‡ *Center for Integrative Genomics, University of Lausanne, Ge´nopode, 1015 Lausanne, Switzerland; and †Computational and Molecular Population Genetics Laboratory, Zoological Institute, University of Bern, 3012 Bern, Switzerland Communicated by Wen-Hsiung Li, University of Chicago, Chicago, IL, December 30, 2005 (received for review December 14, 2005) Given that retroposed copies of genes are presumed to lack the and rodent genomes (7–12). In addition, three recent studies regulatory elements required for their expression, retroposition using EST data (13, 14) and tiling-microarray data from chro- has long been considered a mechanism without functional rele- mosome 22 (15) indicated that retrocopy transcription may be vance. However, through an in silico assay for transcriptional widespread, although these surveys were limited, and potential activity, we identify here >1,000 transcribed retrocopies in the functional implications were not addressed. human genome, of which at least Ϸ120 have evolved into bona To further explore the functional significance of retroposition fide genes. Among these, Ϸ50 retrogenes have evolved functions in the human genome, we systematically screened for signatures in testes, more than half of which were recruited as functional of selection related to retrocopy transcription. Our results autosomal counterparts of X-linked genes during spermatogene- suggest that retrocopy transcription is not rare and that the sis. Generally, retrogenes emerge ‘‘out of the testis,’’ because they pattern of transcription of human retrocopies has been pro- are often initially transcribed in testis and later evolve stronger and foundly shaped by natural selection, acting both for and against sometimes more diverse spatial expression patterns.