Hyaluronan Synthases (HAS1-3) and Hyaluronidases (HYAL1-2) in The
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Evaluation of the Prognostic Potential of Hyaluronic Acid and Hyaluronidase (HYAL1) for Prostate Cancer1
[CANCER RESEARCH 63, 2638–2644, May 15, 2003] Evaluation of the Prognostic Potential of Hyaluronic Acid and Hyaluronidase (HYAL1) for Prostate Cancer1 J. Timothy Posey, Mark S. Soloway, Sinan Ekici, Mario Sofer, Francisco Civantos, Robert C. Duncan, and Vinata B. Lokeshwar2 Departments of Urology [J. T. P., M. S. S., S. E., M. S., F. C., V. B. L.], Department of Epidemiology [R. C. D.], and Cell Biology and Anatomy [V. B. L.], University of Miami School of Medicine, Miami, Florida 33101 ABSTRACT lapse), local/systemic recurrence] in ϳ10–50% of cases, depending on a variety of prognostic factors (5–7). Treatment failure may be Despite the development of nomograms designed to evaluate the prog- attributable to a local recurrence or distant metastasis. Existing pre- nosis of a patient with prostate cancer (CaP), the information has been operative indicators (i.e., PSA levels, clinical stage, biopsy Gleason limited to prostate-specific antigen (PSA), clinical stage, Gleason score, and tumor volume estimates. To improve our ability to predict prognosis, sum) or their combination in nomograms, as well as surgical and ϩ Ϫ information regarding the molecular properties of CaP is needed. Hyalu- pathologic parameters (i.e., prostatectomy Gleason sum, margin / , ronic acid (HA) is a glycosaminoglycan that promotes tumor metastasis. node status, seminal vesicle, and EPE), provide a limited estimate of Hyaluronidase (HAase) is an enzyme that degrades HA into angiogenic the prognosis for CaP (8, 9). Identifying molecules that are expressed fragments. We recently showed that in CaP tissues, whereas HA is local- in clinically localized CaP but associate with CaP invasion and ized mostly in the tumor-associated stroma, HYAL1 type HAase is exclu- metastasis might significantly improve the prognostic capabilities and sively localized in CaP cells (Lokeshwar et al. -
Critical Evaluation of Gene Expression Changes in Human Tissues In
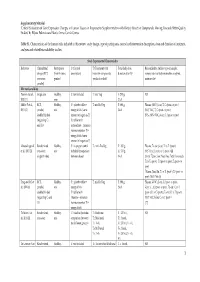
Supplementary Material ‘Critical Evaluation of Gene Expression Changes in Human Tissues in Response to Supplementation with Dietary Bioactive Compounds: Moving Towards Better-Quality Studies’ by Biljana Pokimica and María-Teresa García-Conesa Table S1. Characteristics of the human trials included in this review: study design, type of participants, control and intervention description, dose and duration of treatment, analyses and related bioavailability studies. Study Experimental Characteristics Reference Clinical trial Participants C (Control T (Treatment with Total daily dose, Bioavailability studies: type of sample, design (RCT, (health status, description) bioactive compounds, duration (d or h)1 compounds and (or) metabolites analysed, crossover, gender) products or diet) main results2 parallel) Mix meals and diets Persson I et al., Single arm Healthy, C: not included T: mix Veg T: 250 g, NR 2000 [1] men 21 d Møller P et al., RCT, Healthy, C1: placebo tablet + T: mix FruVeg T: 600 g, Plasma: (NS↑) β-car, T, C2 (post- vs pre-) 2003 [2] parallel, mix energy drink (same 24 d (NC) VitC, T, C2 (post- vs pre-) double blinded amount of sugars as T) (NS↓, 69%) VitC, β-car, C1 (post- vs pre-) (regarding C1 C2: tablet with and C2) antioxidants + minerals (same amount as T) + energy drink (same amount of sugars as T) Almendingen K Randomized, Healthy, C: no proper control T1,2: mix FruVeg T1: 300 g, Plasma: ↑α-car, β-car, T2 vs T1 (post-) et al., 2005 [3] crossover, mix included (comparison T2: 750 g, (NS↑) Lyc, Lut, T2 vs T1 (post-) [4] single -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Characterization of Visceral and Subcutaneous Adipose Tissue
J. Perinat. Med. 2016; 44(7): 813–835 Shali Mazaki-Tovi*, Adi L. Tarca, Edi Vaisbuch, Juan Pedro Kusanovic, Nandor Gabor Than, Tinnakorn Chaiworapongsa, Zhong Dong, Sonia S. Hassan and Roberto Romero* Characterization of visceral and subcutaneous adipose tissue transcriptome in pregnant women with and without spontaneous labor at term: implication of alternative splicing in the metabolic adaptations of adipose tissue to parturition DOI 10.1515/jpm-2015-0259 groups (unpaired analyses) and adipose tissue regions Received July 27, 2015. Accepted October 26, 2015. Previously (paired analyses). Selected genes were tested by quantita- published online March 19, 2016. tive reverse transcription-polymerase chain reaction. Abstract Results: Four hundred and eighty-two genes were differ- entially expressed between visceral and subcutaneous Objective: The aim of this study was to determine gene fat of pregnant women with spontaneous labor at term expression and splicing changes associated with par- (q-value < 0.1; fold change > 1.5). Biological processes turition and regions (visceral vs. subcutaneous) of the enriched in this comparison included tissue and vascu- adipose tissue of pregnant women. lature development as well as inflammatory and meta- Study design: The transcriptome of visceral and abdomi- bolic pathways. Differential splicing was found for 42 nal subcutaneous adipose tissue from pregnant women at genes [q-value < 0.1; differences in Finding Isoforms using term with (n = 15) and without (n = 25) spontaneous labor Robust Multichip Analysis scores > 2] between adipose was profiled with the Affymetrix GeneChip Human Exon tissue regions of women not in labor. Differential exon 1.0 ST array. Overall gene expression changes and the dif- usage associated with parturition was found for three ferential exon usage rate were compared between patient genes (LIMS1, HSPA5, and GSTK1) in subcutaneous tissues. -

HYAL1LUCA-1, a Candidate Tumor Suppressor Gene on Chromosome 3P21.3, Is Inactivated in Head and Neck Squamous Cell Carcinomas by Aberrant Splicing of Pre-Mrna
Oncogene (2000) 19, 870 ± 878 ã 2000 Macmillan Publishers Ltd All rights reserved 0950 ± 9232/00 $15.00 www.nature.com/onc HYAL1LUCA-1, a candidate tumor suppressor gene on chromosome 3p21.3, is inactivated in head and neck squamous cell carcinomas by aberrant splicing of pre-mRNA Gregory I Frost1,3, Gayatry Mohapatra2, Tim M Wong1, Antonei Benjamin Cso ka1, Joe W Gray2 and Robert Stern*,1 1Department of Pathology, School of Medicine, University of California, San Francisco, California, CA 94143, USA; 2Cancer Genetics Program, UCSF Cancer Center, University of California, San Francisco, California, CA 94115, USA The hyaluronidase ®rst isolated from human plasma, genes in the process of carcinogenesis (Sager, 1997; Hyal-1, is expressed in many somatic tissues. The Hyal- Baylin et al., 1998). Nevertheless, functionally 1 gene, HYAL1, also known as LUCA-1, maps to inactivating point mutations are generally viewed as chromosome 3p21.3 within a candidate tumor suppressor the critical `smoking gun' when de®ning a novel gene locus de®ned by homozygous deletions and by TSG. functional tumor suppressor activity. Hemizygosity in We recently mapped the HYAL1 gene to human this region occurs in many malignancies, including chromosome 3p21.3 (Cso ka et al., 1998), con®rming its squamous cell carcinomas of the head and neck. We identity with LUCA-1, a candidate tumor suppressor have investigated whether cell lines derived from such gene frequently deleted in small cell lung carcinomas malignancies expressed Hyal-1 activity, using normal (SCLC) (Wei et al., 1996). The HYAL1 gene resides human keratinocytes as controls. Hyal-1 enzyme activity within a commonly deleted region of 3p21.3 where a and protein were absent or markedly reduced in six of potentially informative 30 kb homozygous deletion has seven carcinoma cell lines examined. -
Figure S1. Reverse Transcription‑Quantitative PCR Analysis of ETV5 Mrna Expression Levels in Parental and ETV5 Stable Transfectants
Figure S1. Reverse transcription‑quantitative PCR analysis of ETV5 mRNA expression levels in parental and ETV5 stable transfectants. (A) Hec1a and Hec1a‑ETV5 EC cell lines; (B) Ishikawa and Ishikawa‑ETV5 EC cell lines. **P<0.005, unpaired Student's t‑test. EC, endometrial cancer; ETV5, ETS variant transcription factor 5. Figure S2. Survival analysis of sample clusters 1‑4. Kaplan Meier graphs for (A) recurrence‑free and (B) overall survival. Survival curves were constructed using the Kaplan‑Meier method, and differences between sample cluster curves were analyzed by log‑rank test. Figure S3. ROC analysis of hub genes. For each gene, ROC curve (left) and mRNA expression levels (right) in control (n=35) and tumor (n=545) samples from The Cancer Genome Atlas Uterine Corpus Endometrioid Cancer cohort are shown. mRNA levels are expressed as Log2(x+1), where ‘x’ is the RSEM normalized expression value. ROC, receiver operating characteristic. Table SI. Clinicopathological characteristics of the GSE17025 dataset. Characteristic n % Atrophic endometrium 12 (postmenopausal) (Control group) Tumor stage I 91 100 Histology Endometrioid adenocarcinoma 79 86.81 Papillary serous 12 13.19 Histological grade Grade 1 30 32.97 Grade 2 36 39.56 Grade 3 25 27.47 Myometrial invasiona Superficial (<50%) 67 74.44 Deep (>50%) 23 25.56 aMyometrial invasion information was available for 90 of 91 tumor samples. Table SII. Clinicopathological characteristics of The Cancer Genome Atlas Uterine Corpus Endometrioid Cancer dataset. Characteristic n % Solid tissue normal 16 Tumor samples Stagea I 226 68.278 II 19 5.740 III 70 21.148 IV 16 4.834 Histology Endometrioid 271 81.381 Mixed 10 3.003 Serous 52 15.616 Histological grade Grade 1 78 23.423 Grade 2 91 27.327 Grade 3 164 49.249 Molecular subtypeb POLE 17 7.328 MSI 65 28.017 CN Low 90 38.793 CN High 60 25.862 CN, copy number; MSI, microsatellite instability; POLE, DNA polymerase ε. -

Supplementary Table 1
Supplementary Table 1. 492 genes are unique to 0 h post-heat timepoint. The name, p-value, fold change, location and family of each gene are indicated. Genes were filtered for an absolute value log2 ration 1.5 and a significance value of p ≤ 0.05. Symbol p-value Log Gene Name Location Family Ratio ABCA13 1.87E-02 3.292 ATP-binding cassette, sub-family unknown transporter A (ABC1), member 13 ABCB1 1.93E-02 −1.819 ATP-binding cassette, sub-family Plasma transporter B (MDR/TAP), member 1 Membrane ABCC3 2.83E-02 2.016 ATP-binding cassette, sub-family Plasma transporter C (CFTR/MRP), member 3 Membrane ABHD6 7.79E-03 −2.717 abhydrolase domain containing 6 Cytoplasm enzyme ACAT1 4.10E-02 3.009 acetyl-CoA acetyltransferase 1 Cytoplasm enzyme ACBD4 2.66E-03 1.722 acyl-CoA binding domain unknown other containing 4 ACSL5 1.86E-02 −2.876 acyl-CoA synthetase long-chain Cytoplasm enzyme family member 5 ADAM23 3.33E-02 −3.008 ADAM metallopeptidase domain Plasma peptidase 23 Membrane ADAM29 5.58E-03 3.463 ADAM metallopeptidase domain Plasma peptidase 29 Membrane ADAMTS17 2.67E-04 3.051 ADAM metallopeptidase with Extracellular other thrombospondin type 1 motif, 17 Space ADCYAP1R1 1.20E-02 1.848 adenylate cyclase activating Plasma G-protein polypeptide 1 (pituitary) receptor Membrane coupled type I receptor ADH6 (includes 4.02E-02 −1.845 alcohol dehydrogenase 6 (class Cytoplasm enzyme EG:130) V) AHSA2 1.54E-04 −1.6 AHA1, activator of heat shock unknown other 90kDa protein ATPase homolog 2 (yeast) AK5 3.32E-02 1.658 adenylate kinase 5 Cytoplasm kinase AK7 -

Resveratrol Inhibits Cell Cycle Progression by Targeting Aurora Kinase a and Polo-Like Kinase 1 in Breast Cancer Cells
3696 ONCOLOGY REPORTS 35: 3696-3704, 2016 Resveratrol inhibits cell cycle progression by targeting Aurora kinase A and Polo-like kinase 1 in breast cancer cells RUBICELI MEDINA-AGUILAR1, LAURENCE A. Marchat2, ELENA ARECHAGA OCAMPO3, Patricio GARIGLIO1, JAIME GARCÍA MENA1, NICOLÁS VILLEGAS SEPÚlveda4, MACARIO MartÍNEZ CASTILLO4 and CÉSAR LÓPEZ-CAMARILLO5 1Department of Genetics and Molecular Biology, CINVESTAV-IPN, Mexico D.F.; 2Molecular Biomedicine Program and Biotechnology Network, National School of Medicine and Homeopathy, National Polytechnic Institute, Mexico D.F.; 3Natural Sciences Department, Metropolitan Autonomous University, Mexico D.F.; 4Department of Molecular Biomedicine, CINVESTAV-IPN, Mexico D.F.; 5Oncogenomics and Cancer Proteomics Laboratory, Universidad Autónoma de la Ciudad de México, Mexico D.F., Mexico Received December 4, 2015; Accepted January 8, 2016 DOI: 10.3892/or.2016.4728 Abstract. The Aurora protein kinase (AURKA) and the MDA-MB-231 and MCF-7 cells. By western blot assays, we Polo-like kinase-1 (PLK1) activate the cell cycle, and they confirmed that resveratrol suppressed AURKA, CCND1 and are considered promising druggable targets in cancer CCNB1 at 24 and 48 h. In summary, we showed for the first time therapy. However, resistance to chemotherapy and to specific that resveratrol regulates cell cycle progression by targeting small-molecule inhibitors is common in cancer patients; thus AURKA and PLK1. Our findings highlight the potential use of alternative therapeutic approaches are needed to overcome resveratrol as an adjuvant therapy for breast cancer. clinical resistance. Here, we showed that the dietary compound resveratrol suppressed the cell cycle by targeting AURKA Introduction and PLK1 kinases. First, we identified genes modulated by resveratrol using a genome-wide analysis of gene expression Cancer development results from the interaction between in MDA-MB-231 breast cancer cells. -

Usbiological Datasheet
HYAL1 (h.HYAL1 , FUS2, NAT6, LUCA1, Hyal-1, LuCa-1, Hyaluronidase-1, Hyaluronoglucosaminidase-1, Lung carcinoma protein 1) Catalog number 144710 Supplier United States Biological Hyaluronidase-1, also known as HYAL1 or LUCA1, is an enzyme that in humans is encoded by the HYAL1 gene. The gene is one of several related genes in a region of chromosome 3p21.3 associated with tumor suppression. This gene encodes a lysosomal hyaluronidase. Hyaluronidases intracellularly degrade hyaluronan, one of the major glycosaminoglycans of the extracellular matrix. Hyaluronan is thought to be involved in cell proliferation, migration and differentiation. This enzyme is active at an acidic pH and is the major hyaluronidase in plasma. Mutations in this gene are associated with mucopolysaccharidosis type IX, or hyaluronidase deficiency. UniProt Number Q12794 Gene ID HYAL1 Applications Suitable for use in Western Blot, Immunohistochemistry (Paraffin and Frozen), and Immunocytochemistry. Recommended Dilution Optimal dilutions to be determined by the researcher. Storage and Handling Store at -20˚C for one year. After reconstitution, store at 4˚C for one month. Can also be aliquoted and stored frozen at -20˚C for long term. Avoid repeated freezing and thawing. For maximum recovery of product, centrifuge the original vial after thawing and prior to removing the cap. Immunogen A synthetic peptide corresponding to a sequence at the N-terminus of human HYAL1. Formulation Supplied as a lyophilized powder. Each vial contains 5mg BSA, 0.9mg NaCl, 0.2mg Na2HPO4, 0.05mg Thimerosal, 0.05mg NaN3. Reconstitution: Add 0.2ml of distilled water will yield a concentration of 500ug/ml. Purity Purified by immunoaffinity chromatography. -

Mechanical Stress Induce PG-E2 in Murine Synovial Fibroblasts Originating from the Temporomandibular Joint
cells Article Mechanical Stress Induce PG-E2 in Murine Synovial Fibroblasts Originating from the Temporomandibular Joint Ute Nazet 1,* , Laura Feulner 1, Dominique Muschter 2 , Patrick Neubert 3, Valentin Schatz 3, Susanne Grässel 2 , Jonathan Jantsch 3, Peter Proff 1, Agnes Schröder 1,† and Christian Kirschneck 1,† 1 Department of Orthodontics, University Medical Centre of Regensburg, D-93053 Regensburg, Germany; [email protected] (L.F.); [email protected] (P.P.); [email protected] (A.S.); [email protected] (C.K.) 2 Centre for Medical Biotechnology, Department of Orthopaedic Surgery, Experimental Orthopaedics, University of Regensburg, D-93053 Regensburg, Germany; [email protected] (D.M.); [email protected] (S.G.) 3 Institute of Clinical Microbiology and Hygiene, University Hospital of Regensburg, D-93053 Regensburg, Germany; [email protected] (P.N.); [email protected] (V.S.); [email protected] (J.J.) * Correspondence: [email protected]; Tel.: +49-941-944-4990 † Contributed equally. Abstract: Genetic predisposition, traumatic events, or excessive mechanical exposure provoke arthritic changes in the temporomandibular joint (TMJ). We analysed the impact of mechanical stress that might be involved in the development and progression of TMJ osteoarthritis (OA) on murine synovial fibroblasts (SFs) of temporomandibular origin. SFs were subjected to different Citation: Nazet, U.; Feulner, L.; Muschter, D.; Neubert, P.; Schatz, V.; protocols of mechanical stress, either to a high-frequency tensile strain for 4 h or to a tensile strain of Grässel, S.; Jantsch, J.; Proff, P.; varying magnitude for 48 h. The TMJ OA induction was evaluated based on the gene and protein Schröder, A.; Kirschneck, C. -

Datasheet: 50309954
Datasheet: 50309954 Description: NATIVE BOVINE HYALURONIDASE Name: HYALURONIDASE Format: Purified Product Type: Purified Protein Quantity: 0.5 g Product Details Applications This product has been reported to work in the following applications. This information is derived from testing within our laboratories, peer-reviewed publications or personal communications from the originators. Please refer to references indicated for further information. For general protocol recommendations, please visit www.bio-rad-antibodies.com/protocols. Yes No Not Determined Suggested Dilution ELISA Where this product has not been tested for use in a particular technique this does not necessarily exclude its use in such procedures. Suggested working dilutions are given as a guide only. It is recommended that the user titrates the product for use in their own system using the appropriate negative/positive controls. Target Species Bovine Product Form Purified enzyme from bovine testes - liquid Preservative None present Stabilisers Approx. Protein Total protein concentration 0.2 g/ml Concentrations External Database UniProt: Links Q8SQG8 Related reagents Q5E985 Related reagents Entrez Gene: 281838 HYAL2 Related reagents 515397 HYAL1 Related reagents Product Information Hyaluronidase catalyzes the depolymerization of mucopolysaccharides, hyaluronic acid, and the chondroitin sulfates A and C. The enzyme is widely distributed in animal tissues but is found in great concentrations in the bovine and ovine testes. Molecular Weight MW: 55 kD Activity 315 U/mg protein 1 unit is defined as the amount of enzyme that liberates one micromole of N-acetylglucosamine Page 1 of 2 per minute at 37ºC and pH 4.0. EC 3.2.1.36 Storage Store at -20oC only. -

Molecular Signatures Differentiate Immune States in Type 1 Diabetes Families
Page 1 of 65 Diabetes Molecular signatures differentiate immune states in Type 1 diabetes families Yi-Guang Chen1, Susanne M. Cabrera1, Shuang Jia1, Mary L. Kaldunski1, Joanna Kramer1, Sami Cheong2, Rhonda Geoffrey1, Mark F. Roethle1, Jeffrey E. Woodliff3, Carla J. Greenbaum4, Xujing Wang5, and Martin J. Hessner1 1The Max McGee National Research Center for Juvenile Diabetes, Children's Research Institute of Children's Hospital of Wisconsin, and Department of Pediatrics at the Medical College of Wisconsin Milwaukee, WI 53226, USA. 2The Department of Mathematical Sciences, University of Wisconsin-Milwaukee, Milwaukee, WI 53211, USA. 3Flow Cytometry & Cell Separation Facility, Bindley Bioscience Center, Purdue University, West Lafayette, IN 47907, USA. 4Diabetes Research Program, Benaroya Research Institute, Seattle, WA, 98101, USA. 5Systems Biology Center, the National Heart, Lung, and Blood Institute, the National Institutes of Health, Bethesda, MD 20824, USA. Corresponding author: Martin J. Hessner, Ph.D., The Department of Pediatrics, The Medical College of Wisconsin, Milwaukee, WI 53226, USA Tel: 011-1-414-955-4496; Fax: 011-1-414-955-6663; E-mail: [email protected]. Running title: Innate Inflammation in T1D Families Word count: 3999 Number of Tables: 1 Number of Figures: 7 1 For Peer Review Only Diabetes Publish Ahead of Print, published online April 23, 2014 Diabetes Page 2 of 65 ABSTRACT Mechanisms associated with Type 1 diabetes (T1D) development remain incompletely defined. Employing a sensitive array-based bioassay where patient plasma is used to induce transcriptional responses in healthy leukocytes, we previously reported disease-specific, partially IL-1 dependent, signatures associated with pre and recent onset (RO) T1D relative to unrelated healthy controls (uHC).