Analysis of Map3k-Dependent Molecular Mechanisms Regulating Signaling
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplemental Information to Mammadova-Bach Et Al., “Laminin Α1 Orchestrates VEGFA Functions in the Ecosystem of Colorectal Carcinogenesis”
Supplemental information to Mammadova-Bach et al., “Laminin α1 orchestrates VEGFA functions in the ecosystem of colorectal carcinogenesis” Supplemental material and methods Cloning of the villin-LMα1 vector The plasmid pBS-villin-promoter containing the 3.5 Kb of the murine villin promoter, the first non coding exon, 5.5 kb of the first intron and 15 nucleotides of the second villin exon, was generated by S. Robine (Institut Curie, Paris, France). The EcoRI site in the multi cloning site was destroyed by fill in ligation with T4 polymerase according to the manufacturer`s instructions (New England Biolabs, Ozyme, Saint Quentin en Yvelines, France). Site directed mutagenesis (GeneEditor in vitro Site-Directed Mutagenesis system, Promega, Charbonnières-les-Bains, France) was then used to introduce a BsiWI site before the start codon of the villin coding sequence using the 5’ phosphorylated primer: 5’CCTTCTCCTCTAGGCTCGCGTACGATGACGTCGGACTTGCGG3’. A double strand annealed oligonucleotide, 5’GGCCGGACGCGTGAATTCGTCGACGC3’ and 5’GGCCGCGTCGACGAATTCACGC GTCC3’ containing restriction site for MluI, EcoRI and SalI were inserted in the NotI site (present in the multi cloning site), generating the plasmid pBS-villin-promoter-MES. The SV40 polyA region of the pEGFP plasmid (Clontech, Ozyme, Saint Quentin Yvelines, France) was amplified by PCR using primers 5’GGCGCCTCTAGATCATAATCAGCCATA3’ and 5’GGCGCCCTTAAGATACATTGATGAGTT3’ before subcloning into the pGEMTeasy vector (Promega, Charbonnières-les-Bains, France). After EcoRI digestion, the SV40 polyA fragment was purified with the NucleoSpin Extract II kit (Machery-Nagel, Hoerdt, France) and then subcloned into the EcoRI site of the plasmid pBS-villin-promoter-MES. Site directed mutagenesis was used to introduce a BsiWI site (5’ phosphorylated AGCGCAGGGAGCGGCGGCCGTACGATGCGCGGCAGCGGCACG3’) before the initiation codon and a MluI site (5’ phosphorylated 1 CCCGGGCCTGAGCCCTAAACGCGTGCCAGCCTCTGCCCTTGG3’) after the stop codon in the full length cDNA coding for the mouse LMα1 in the pCIS vector (kindly provided by P. -

1 Tumor Suppressor PLK2 May Serve As a Biomarker in Triple-Negative Breast Cancer for Improved Response to PLK1 Therapeutics
bioRxiv preprint doi: https://doi.org/10.1101/2021.06.16.448722; this version posted June 16, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Tumor suppressor PLK2 may serve as a biomarker in triple-negative breast cancer for improved response to PLK1 therapeutics Yang Gao1, 2, 3, Elena B. Kabotyanski1, 2, Elizabeth Villegas7, Jonathan H. Shepherd8, Deanna Acosta1, 2, Clark Hamor1, 2, Tingting Sun2,4,5, Celina Montmeyor-Garcia9, Xiaping He8, Lacey E. Dobrolecki1, 2, 3, Thomas F. Westbrook2, 4, 5, Michael T. Lewis1, 2, 3, Susan G. Hilsenbeck2, 3, Xiang H.-F. Zhang1, 2, 3, 6, Charles M. Perou8 and Jeffrey M. Rosen1, 2 1Department of Molecular and Cellular Biology 2Dan L. Duncan Cancer Center 3Lester and Sue Smith Breast Center 4Department of Molecular and Human Genetics 5Verna & Marrs McLean Department of Biochemistry and Molecular Biology 6McNair Medical Institute Baylor College of Medicine, One Baylor Plaza, Houston, TX 77030, USA 7University of Houston-Downtown, Houston, TX 77002, USA 8The University of North Carolina at Chapel Hill, Chapel Hill, NC 27599, USA 9 Canadian Blood Services, Toronto, ON M5G 2M1, Canada Correspondence to Jeffrey M. Rosen (Mail Stop: BCM130, Room: BCM-M638a, Baylor College of Medicine, 1 Baylor Plaza, Houston, TX 77030. Office: 713-798-6210. Fax: 713-898-8012. Email: [email protected]) 1 bioRxiv preprint doi: https://doi.org/10.1101/2021.06.16.448722; this version posted June 16, 2021. -

TAK1 Mediates Convergence of Cellular Signals for Death and Survival
Apoptosis (2019) 24:3–20 https://doi.org/10.1007/s10495-018-1490-7 REVIEW TAK1 mediates convergence of cellular signals for death and survival Sabreena Aashaq1 · Asiya Batool1 · Khurshid I. Andrabi1 Published online: 4 October 2018 © Springer Science+Business Media, LLC, part of Springer Nature 2018 Abstract TGF-β activated kinase 1, a MAPK kinase kinase family serine threonine kinase has been implicated in regulating diverse range of cellular processes that include embryonic development, differentiation, autophagy, apoptosis and cell survival. TAK1 along with its binding partners TAB1, TAB2 and TAB3 displays a complex pattern of regulation that includes serious crosstalk with major signaling pathways including the C-Jun N-terminal kinase (JNK), p38 MAPK, and I-kappa B kinase complex (IKK) involved in establishing cellular commitments for death and survival. This review also highlights how TAK1 orchestrates regulation of energy homeostasis via AMPK and its emerging role in influencing mTORC1 pathway to regulate death or survival in tandem. Keywords Apoptosis · Autophagy · Cytokine · Inflammatory · Smad TAK1, a multifunctional kinase regulate a wide array of downstream cellular responses [2, 7, 8]. Various branches of MAP kinase pathways including Transforming growth factor-β is a versatile cytokine, regu- the extracellular signal regulated kinase (Erk) ½ [3, 9], p38 lating a wide variety of intracellular signaling pathways. MAPK [10, 11], c-Jun N-Terminal kinase (JNK) [12, 13], The Smad dependent signaling pathway is conventionally phosphatidylinositol-3-kinase/AKT pathway [14, 15] and acknowledged as the traditional pathway promoted by TGF- Rho-like GTPase [16, 17] signaling pathways are included β1 [1]. However, the Smad dependent signaling pathway among the Smad independent pathways. -

Genome-Wide Association Study to Identify Genomic Regions And
www.nature.com/scientificreports OPEN Genome‑wide association study to identify genomic regions and positional candidate genes associated with male fertility in beef cattle H. Sweett1, P. A. S. Fonseca1, A. Suárez‑Vega1, A. Livernois1,2, F. Miglior1 & A. Cánovas1* Fertility plays a key role in the success of calf production, but there is evidence that reproductive efciency in beef cattle has decreased during the past half‑century worldwide. Therefore, identifying animals with superior fertility could signifcantly impact cow‑calf production efciency. The objective of this research was to identify candidate regions afecting bull fertility in beef cattle and positional candidate genes annotated within these regions. A GWAS using a weighted single‑step genomic BLUP approach was performed on 265 crossbred beef bulls to identify markers associated with scrotal circumference (SC) and sperm motility (SM). Eight windows containing 32 positional candidate genes and fve windows containing 28 positional candidate genes explained more than 1% of the genetic variance for SC and SM, respectively. These windows were selected to perform gene annotation, QTL enrichment, and functional analyses. Functional candidate gene prioritization analysis revealed 14 prioritized candidate genes for SC of which MAP3K1 and VIP were previously found to play roles in male fertility. A diferent set of 14 prioritized genes were identifed for SM and fve were previously identifed as regulators of male fertility (SOD2, TCP1, PACRG, SPEF2, PRLR). Signifcant enrichment results were identifed for fertility and body conformation QTLs within the candidate windows. Gene ontology enrichment analysis including biological processes, molecular functions, and cellular components revealed signifcant GO terms associated with male fertility. -

High Expression of LINC01268 Is Positively Associated with Hepatocellular Carcinoma Progression Via Regulating MAP3K7
OncoTargets and Therapy Dovepress open access to scientific and medical research Open Access Full Text Article ORIGINAL RESEARCH High Expression of LINC01268 is Positively Associated with Hepatocellular Carcinoma Progression via Regulating MAP3K7 This article was published in the following Dove Press journal: OncoTargets and Therapy Xiuli Jin,1 Weixin Fu,2 Dan Li,1 Objective: As one of the most common neoplastic diseases, hepatocellular carcinoma Ningning Wang,1 Jiayu Chen,1 (HCC) has a high morbidity and mortality, which seriously threatens human health and Zilu Zeng,1 Jiaqi Guo,1 Hao places a heavy burden on society and medical care. At present, effective early diagnosis, Liu,3 Xinping Zhong,3 Hu prognosis and treatment of HCC are limited. Altered gene expression patterns of lncRNA are Peng,4 Xin Yu,5 Jing Sun,1 associated with the occurrence, development and prognosis of various malignancies, includ- ing HCC. The aim of this study was to investigate the correlation between the expression of Xinhe Zhang,1 Xue Wang,1 LINC01268 and HCC, and to elucidate the potential underlying molecular mechanism. Beibei Xu,1 Yingbo Lin,6 4 Methods: Expression level and localization of LINC01268 in human liver cancer cells and Jianping Liu, Claudia HCC tissues were investigated using RT-qPCR and fluorescent in situ hybridization (FISH), 7 1 Kutter, Yiling Li respectively. Correlation of expression levels of LINC01268 and MAP3K7 with differentia- 1Department of Gastroenterology, First tion and poor overall patient survival of HCC were analyzed using in house collected and AffiliatedHospital of China Medical publicly available HCC tissue data. RT-qPCR and Western blot were applied to inspect the University, Shenyang, 110001, People’s Republic of China; 2Science Experiment effects of depletion and overexpression of LINC01268 on MAP3K7 expression. -

Kinase Profiling Book
Custom and Pre-Selected Kinase Prof iling to f it your Budget and Needs! As of July 1, 2021 19.8653 mm 128 196 12 Tyrosine Serine/Threonine Lipid Kinases Kinases Kinases Carna Biosciences, Inc. 2007 Carna Biosciences, Inc. Profiling Assays available from Carna Biosciences, Inc. As of July 1, 2021 Page Kinase Name Assay Platform Page Kinase Name Assay Platform 4 ABL(ABL1) MSA 21 EGFR[T790M/C797S/L858R] MSA 4 ABL(ABL1)[E255K] MSA 21 EGFR[T790M/L858R] MSA 4 ABL(ABL1)[T315I] MSA 21 EPHA1 MSA 4 ACK(TNK2) MSA 21 EPHA2 MSA 4 AKT1 MSA 21 EPHA3 MSA 5 AKT2 MSA 22 EPHA4 MSA 5 AKT3 MSA 22 EPHA5 MSA 5 ALK MSA 22 EPHA6 MSA 5 ALK[C1156Y] MSA 22 EPHA7 MSA 5 ALK[F1174L] MSA 22 EPHA8 MSA 6 ALK[G1202R] MSA 23 EPHB1 MSA 6 ALK[G1269A] MSA 23 EPHB2 MSA 6 ALK[L1196M] MSA 23 EPHB3 MSA 6 ALK[R1275Q] MSA 23 EPHB4 MSA 6 ALK[T1151_L1152insT] MSA 23 Erk1(MAPK3) MSA 7 EML4-ALK MSA 24 Erk2(MAPK1) MSA 7 NPM1-ALK MSA 24 Erk5(MAPK7) MSA 7 AMPKα1/β1/γ1(PRKAA1/B1/G1) MSA 24 FAK(PTK2) MSA 7 AMPKα2/β1/γ1(PRKAA2/B1/G1) MSA 24 FER MSA 7 ARG(ABL2) MSA 24 FES MSA 8 AurA(AURKA) MSA 25 FGFR1 MSA 8 AurA(AURKA)/TPX2 MSA 25 FGFR1[V561M] MSA 8 AurB(AURKB)/INCENP MSA 25 FGFR2 MSA 8 AurC(AURKC) MSA 25 FGFR2[V564I] MSA 8 AXL MSA 25 FGFR3 MSA 9 BLK MSA 26 FGFR3[K650E] MSA 9 BMX MSA 26 FGFR3[K650M] MSA 9 BRK(PTK6) MSA 26 FGFR3[V555L] MSA 9 BRSK1 MSA 26 FGFR3[V555M] MSA 9 BRSK2 MSA 26 FGFR4 MSA 10 BTK MSA 27 FGFR4[N535K] MSA 10 BTK[C481S] MSA 27 FGFR4[V550E] MSA 10 BUB1/BUB3 MSA 27 FGFR4[V550L] MSA 10 CaMK1α(CAMK1) MSA 27 FGR MSA 10 CaMK1δ(CAMK1D) MSA 27 FLT1 MSA 11 CaMK2α(CAMK2A) MSA 28 -

Modulation of NF-Κb Signalling by Microbial Pathogens
REVIEWS Modulation of NF‑κB signalling by microbial pathogens Masmudur M. Rahman and Grant McFadden Abstract | The nuclear factor-κB (NF‑κB) family of transcription factors plays a central part in the host response to infection by microbial pathogens, by orchestrating the innate and acquired host immune responses. The NF‑κB proteins are activated by diverse signalling pathways that originate from many different cellular receptors and sensors. Many successful pathogens have acquired sophisticated mechanisms to regulate the NF‑κB signalling pathways by deploying subversive proteins or hijacking the host signalling molecules. Here, we describe the mechanisms by which viruses and bacteria micromanage the host NF‑κB signalling circuitry to favour the continued survival of the pathogen. The nuclear factor-κB (NF-κB) family of transcription Signalling targets upstream of NF‑κB factors regulates the expression of hundreds of genes that NF-κB proteins are tightly regulated in both the cyto- are associated with diverse cellular processes, such as pro- plasm and the nucleus6. Under normal physiological liferation, differentiation and death, as well as innate and conditions, NF‑κB complexes remain inactive in the adaptive immune responses. The mammalian NF‑κB cytoplasm through a direct interaction with proteins proteins are members of the Rel domain-containing pro- of the inhibitor of NF-κB (IκB) family, including IκBα, tein family: RELA (also known as p65), RELB, c‑REL, IκBβ and IκBε (also known as NF-κBIα, NF-κBIβ and the NF-κB p105 subunit (also known as NF‑κB1; which NF-κBIε, respectively); IκB proteins mask the nuclear is cleaved into the p50 subunit) and the NF-κB p100 localization domains in the NF‑κB complex, thus subunit (also known as NF‑κB2; which is cleaved into retaining the transcription complex in the cytoplasm. -

Comparing Signaling Networks Between Normal and Transformed Hepatocytes Using Discrete Logical Models
Published OnlineFirst July 8, 2011; DOI: 10.1158/0008-5472.CAN-10-4453 Cancer Integrated Systems and Technologies: Mathematical Oncology Research Comparing Signaling Networks between Normal and Transformed Hepatocytes Using Discrete Logical Models Julio Saez-Rodriguez1,2, Leonidas G. Alexopoulos1,2, MingSheng Zhang1, Melody K. Morris2, Douglas A. Lauffenburger2, and Peter K. Sorger1,2 Abstract Substantial effort in recent years has been devoted to constructing and analyzing large-scale gene and protein networks on the basis of "omic" data and literature mining. These interaction graphs provide valuable insight into the topologies of complex biological networks but are rarely context specific and cannot be used to predict the responses of cell signaling proteins to specific ligands or drugs. Conversely, traditional approaches to analyzing cell signaling are narrow in scope and cannot easily make use of network-level data. Here, we combine network analysis and functional experimentation by using a hybrid approach in which graphs are converted into simple mathematical models that can be trained against biochemical data. Specifically, we created Boolean logic models of immediate-early signaling in liver cells by training a literature-based prior knowledge network against biochemical data obtained from primary human hepatocytes and 4 hepatocellular carcinoma cell lines exposed to combinations of cytokines and small-molecule kinase inhibitors. Distinct families of models were recovered for each cell type, and these families clustered topologically into normal and diseased sets. Cancer Res; 71(16); 5400–11. Ó2011 AACR. Introduction Major Findings The availability of high-throughput interaction data has led Clustering arises from systematic differences in signaling to the creation of methods for summarizing and exploring logic in three regions of the network. -

MAP Kinase Kinase Kinase, MEKK, MAPKKK
MAP3K MAP kinase kinase kinase, MEKK, MAPKKK MAP3Ks (Mitogen-activated protein kinase kinase kinases), the top components of MAPK cascades, provide specificity for stimulus-dependent activation of MAP2K-MAPK pathways through unique protein-protein interactions and phosphorylation of signaling effectors. The MAP3Ks are highly divergent in gene numbers and structure, including TAK1, ASK1, A-Raf and C-Raf. MAPK system is a three-step sequential phosphorylation cascade which is composed of MAPK, MAP2K, and MAP3K. ERK, p38 MAPK, and JNK, which are known to be activated by mechanical stimuli, belong to the MAPK family. MAP3Ks function as “platforms to integrate MAPK signaling, and activation of multiple MAP3Ks provides the spatiotemporal regulation of the MAPK pathways, which induces a wide range of physiological responses required for determining cell fate, such as cytokine production, survival, differentiation and apoptosis”. www.MedChemExpress.com 1 MAP3K Inhibitors 5Z-7-Oxozeaenol ASK1-IN-2 (FR148083; L783279; LL-Z 1640-2) Cat. No.: HY-12686 Cat. No.: HY-131969 5Z-7-Oxozeaenol is a natural anti-protozoan ASK1-IN-2 is a potent and orally active inhibitor compound from fungal origin, acting as a potent of apoptosis signal-regulating kinase 1 (ASK1), irreversible and selective inhibitor of TAK1 and with an IC50 of 32.8 nM. ASK1-IN-2 can be used for VEGF-R2, with IC50s of 8 nM and 52 nM, the research of ulcerative colitis. respectively. Purity: ≥99.0% Purity: 98.49% Clinical Data: No Development Reported Clinical Data: No Development Reported Size: 1 mg Size: 10 mM × 1 mL, 5 mg, 10 mg, 25 mg, 50 mg, 100 mg Cot inhibitor-1 Cot inhibitor-2 Cat. -

MAP3K7 Antibody Purified Mouse Monoclonal Antibody Catalog # Ao2028a
10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 MAP3K7 Antibody Purified Mouse Monoclonal Antibody Catalog # AO2028a Specification MAP3K7 Antibody - Product Information Application WB, E Primary Accession O43318 Reactivity Human Host Mouse Clonality Monoclonal Isotype IgG2a Calculated MW 67.2kDa KDa Description The protein encoded by this gene is a member of the serine/threonine protein kinase family. This kinase mediates the signaling transduction induced by TGF beta and morphogenetic protein (BMP), and controls a variety of cell functions including transcription regulation and apoptosis. In response to IL-1, this protein forms a kinase complex including TRAF6, MAP3K7P1/TAB1 and MAP3K7P2/TAB2; this complex is required for the activation of nuclear factor kappa B. This kinase can also activate MAPK8/JNK, MAP2K4/MKK4, and thus plays a role in the cell response to environmental stresses. Four alternatively spliced transcript variants encoding distinct isoforms have been reported. Immunogen Purified recombinant fragment of human MAP3K7 (AA: 471-579) expressed in E. Coli. Formulation Purified antibody in PBS with 0.05% sodium azide and 0.5% protein stabilizer MAP3K7 Antibody - Additional Information Gene ID 6885 Other Names Mitogen-activated protein kinase kinase kinase 7, 2.7.11.25, Transforming growth factor-beta-activated kinase 1, TGF-beta-activated kinase 1, MAP3K7, TAK1 Page 1/3 10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 Dilution WB~~1/500 - 1/2000 E~~1/10000 Storage Maintain refrigerated at 2-8°C for up to 6 months. -
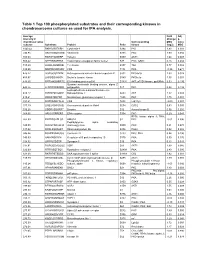
Table 1 Top 100 Phosphorylated Substrates and Their Corresponding Kinases in Chondrosarcoma Cultures As Used for IPA Analysis
Table 1 Top 100 phosphorylated substrates and their corresponding kinases in chondrosarcoma cultures as used for IPA analysis. Average Fold Adj intensity in Change p- chondrosarcoma Corresponding MSC value cultures Substrate Protein Psite kinase (log2) MSC 1043.42 RKKKVSSTKRH Cytohesin-1 S394 PKC 1.83 0.001 746.95 RKGYRSQRGHS Vitronectin S381 PKC 1.00 0.056 709.03 RARSTSLNERP Tuberin S939 AKT1 1.64 0.008 559.42 SPPRSSLRRSS Transcription elongation factor A-like1 S37 PKC; GSK3 0.18 0.684 515.29 LRRSLSRSMSQ Telethonin S157 Titin 0.77 0.082 510.00 MQPDNSSDSDY CD5 T434 PKA -0.35 0.671 476.27 GGRGGSRARNL Heterogeneous nuclear ribonucleoprotein K S302 PKCdelta 1.03 0.028 455.97 LKPGSSHRKTK Bruton's tyrosine kinase S180 PKCbeta 1.55 0.001 444.65 RRRMASMQRTG E1A binding protein p300 S1834 AKT; p70S6 kinase; pp90Rsk 0.53 0.195 Guanine nucleotide binding protein, alpha Z 440.26 HLRSESQRQRR polypeptide S27 PKC 0.88 0.199 6-phosphofructo-2-kinase/fructose-2,6- 424.12 RPRNYSVGSRP biphosphatase 2 S483 AKT 1.32 0.003 419.61 KKKIATRKPRF Metabotropic glutamate receptor 1 T695 PKC 1.75 0.001 391.21 DNSSDSDYDLH CD5 T453 Lck; Fyn -2.09 0.001 377.39 LRQLRSPRRAQ Ras associated protein Rab4 S204 CDC2 0.63 0.091 376.28 SSQRVSSYRRT Desmin S12 Aurora kinase B 0.56 0.255 369.05 ARIGGSRRERS EP4 receptor S354 PKC 0.29 0.543 RPS6 kinase alpha 3; PKA; 367.99 EPKRRSARLSA HMG14 S7 PKC -0.01 0.996 Peptidylglycine alpha amidating 349.08 SRKGYSRKGFD monooxygenase S930 PKC 0.21 0.678 347.92 RRRLSSLRAST Ribosomal protein S6 S236 PAK2 0.02 0.985 346.84 RSNPPSRKGSG Connexin -

Molecular Signatures of Membrane Protein Complexes Underlying Muscular Dystrophy*□S
crossmark Research Author’s Choice © 2016 by The American Society for Biochemistry and Molecular Biology, Inc. This paper is available on line at http://www.mcponline.org Molecular Signatures of Membrane Protein Complexes Underlying Muscular Dystrophy*□S Rolf Turk‡§¶ʈ**, Jordy J. Hsiao¶, Melinda M. Smits¶, Brandon H. Ng¶, Tyler C. Pospisil‡§¶ʈ**, Kayla S. Jones‡§¶ʈ**, Kevin P. Campbell‡§¶ʈ**, and Michael E. Wright¶‡‡ Mutations in genes encoding components of the sar- The muscular dystrophies are hereditary diseases charac- colemmal dystrophin-glycoprotein complex (DGC) are re- terized primarily by the progressive degeneration and weak- sponsible for a large number of muscular dystrophies. As ness of skeletal muscle. Most are caused by deficiencies in such, molecular dissection of the DGC is expected to both proteins associated with the cell membrane (i.e. the sarco- reveal pathological mechanisms, and provides a biologi- lemma in skeletal muscle), and typical features include insta- cal framework for validating new DGC components. Es- bility of the sarcolemma and consequent death of the myofi- tablishment of the molecular composition of plasma- ber (1). membrane protein complexes has been hampered by a One class of muscular dystrophies is caused by mutations lack of suitable biochemical approaches. Here we present in genes that encode components of the sarcolemmal dys- an analytical workflow based upon the principles of pro- tein correlation profiling that has enabled us to model the trophin-glycoprotein complex (DGC). In differentiated skeletal molecular composition of the DGC in mouse skeletal mus- muscle, this structure links the extracellular matrix to the cle. We also report our analysis of protein complexes in intracellular cytoskeleton.