Lokiarchaeota: Biologists Discover 'Missing Link' Microorganism
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Chapter 10: Classification of Microorganisms
Chapter 10: Classification of Microorganisms 1. The Taxonomic Hierarchy 2. Methods of Identification 1. The Taxonomic Hierarchy Phylogenetic Tree of the 3 Domains Taxonomic Hierarchy • 8 successive taxa are used to classify each species: Domain Kingdom Phylum Class Order Family Genus **species can also contain different strains** Species Scientific Nomenclature To avoid confusion, every type of organism must be referred to in a consistent way. The current system of nomenclature (naming) has been in use since the 18th century: • every type of organism is referred by its genus name followed by its specific epithet (i.e., species name) Homo sapiens (H. sapiens) Escherichia coli (E. coli) • name should be in italics and only the genus is capitalized which can also be abbreviated • names are Latin (or “Latinized” Greek) with the genus being a noun and the specific epithet an adjective **strain info can be listed after the specific epithet (e.g., E. coli DH5α)** 2. Methods of Identification Biochemical Testing In addition to morphological (i.e., appearance under the microscope) and differential staining characteristics, microorganisms can also be identified by their biochemical “signatures”: • the nutrient requirements and metabolic “by-products” of of a particular microorganism • different growth media can be used to test the physiological characteristics of a microorganism • e.g., medium with lactose only as energy source • e.g., medium that reveals H2S production **appearance on test medium reveals + or – result!** Commercial devices for rapid Identification Perform multiple tests simultaneously Enterotube II Such devices involve the simultaneous inoculation of various test media: • ~24 hrs later the panel of results reveals ID of organism! Use of Dichotomous Keys Series of “yes/no” biochemical tests to ID organism. -

Revised Glossary for AQA GCSE Biology Student Book
Biology Glossary amino acids small molecules from which proteins are A built abiotic factor physical or non-living conditions amylase a digestive enzyme (carbohydrase) that that affect the distribution of a population in an breaks down starch ecosystem, such as light, temperature, soil pH anaerobic respiration respiration without using absorption the process by which soluble products oxygen of digestion move into the blood from the small intestine antibacterial chemicals chemicals produced by plants as a defence mechanism; the amount abstinence method of contraception whereby the produced will increase if the plant is under attack couple refrains from intercourse, particularly when an egg might be in the oviduct antibiotic e.g. penicillin; medicines that work inside the body to kill bacterial pathogens accommodation ability of the eyes to change focus antibody protein normally present in the body acid rain rain water which is made more acidic by or produced in response to an antigen, which it pollutant gases neutralises, thus producing an immune response active site the place on an enzyme where the antimicrobial resistance (AMR) an increasing substrate molecule binds problem in the twenty-first century whereby active transport in active transport, cells use energy bacteria have evolved to develop resistance against to transport substances through cell membranes antibiotics due to their overuse against a concentration gradient antiretroviral drugs drugs used to treat HIV adaptation features that organisms have to help infections; they -

Marine Microorganisms: Evolution and Solution to Pollution Fu L Li1, Wang B1,2
COMMENTARY Marine microorganisms: Evolution and solution to pollution Fu L Li1, Wang B1,2 Li FL, Wang B. Marine microorganisms: Evolution and solution to pollution. J Mar Microbiol. 2018;2(1):4-5. nce ocean nurtured life, now she needs our care. Marine microorganism will be an opportunity to further understand ourselves and to seek for new Ois the host of ocean in all ages. We should learn from them humbly. methods of fighting old infections. Marine microorganism is tightly bond with human during the history of evolution and nowadays’ environment pollution. Along with industrial revolution, our marine ecosystem suffered serious pollutions. Microplastics are tiny plastic particles (<5 mm) (Figure 1B), Although the topic is still in debate, life is probably originated from which poison marine lives. Because these microplastics are very hard to be submarine in hydrothermal vent systems (1). In the journey of evolution, our degraded, it is predicted that there will be more microplastics than fish in biosphere was completely dominated by microbes for a very long time (Figure ocean by the year 2050 (7). Since marine sediments are considered as the sink 1A). Human being evolves with those microorganisms. Consequently, of microplastics and marine microbes are key dwellers of marine sediments, the influences of microorganisms can be found in all aspects of human more attention should be paid on the interactions between microplastics biology. More than 65% of our genes originated with bacteria, archaea, and and marine microbes. Actually, a call for this has been published in 2011 unicellular eukaryotes, including those genes responsible for host-microbe (8). -

Two Domains of Life, Not Three?
Asgard Archaea Two domains of life, not three? A picture of visual interpretation of the Wagner’s opera Das Rheingold, a part of Richard Wagner’s Der Ring des Nibelungen Loki's Castle is a field of five active hydrothermal vents in the mid-Atlantic Ocean, located at 73 degrees north on the Mid-Atlantic Ridge between Greenland and Norway at a depth of 2,352 metres The vents were discovered in 2008 by a multinational scientific expedition of the university of Bergen, and are the most northerly black smokers to date. The five active chimneys of Loki's Castle are venting water as hot as 300 °C and sit on a vast mound of sulfide minerals. The vent field was given the name Loki's Castle as its shape reminded its discoverers of a fantasy castle. The reference is to the ancient Norse god of trickery, Loki. The top three feet (1 m) of a vent chimney almost 40 feet (12 m) tall at Loki's Castle in mid-July 2008. Visible at left is the arm of a remotely operated vehicle, reaching in to take fluid samples. Loki’s Castle Wents Loki's Castle - a field of five active hydrothermal vents in the mid-Atlantic Ocean - at 73 degrees north on the Mid-Atlantic Ridge - at a depth of 2,352 meters In Norse mythology, Loki is a cunning trickster who has the ability to change his shape and sex. Loki is represented as the companion of the great gods Odin and Thor. Preliminary observations have shown the warm area around the Loki's Castle vents to be alive with diverse and apparently unique microorganisms, unlike vent communities observed elsewhere. -

Laboratory Exercises in Microbiology: Discovering the Unseen World Through Hands-On Investigation
City University of New York (CUNY) CUNY Academic Works Open Educational Resources Queensborough Community College 2016 Laboratory Exercises in Microbiology: Discovering the Unseen World Through Hands-On Investigation Joan Petersen CUNY Queensborough Community College Susan McLaughlin CUNY Queensborough Community College How does access to this work benefit ou?y Let us know! More information about this work at: https://academicworks.cuny.edu/qb_oers/16 Discover additional works at: https://academicworks.cuny.edu This work is made publicly available by the City University of New York (CUNY). Contact: [email protected] Laboratory Exercises in Microbiology: Discovering the Unseen World through Hands-On Investigation By Dr. Susan McLaughlin & Dr. Joan Petersen Queensborough Community College Laboratory Exercises in Microbiology: Discovering the Unseen World through Hands-On Investigation Table of Contents Preface………………………………………………………………………………………i Acknowledgments…………………………………………………………………………..ii Microbiology Lab Safety Instructions…………………………………………………...... iii Lab 1. Introduction to Microscopy and Diversity of Cell Types……………………......... 1 Lab 2. Introduction to Aseptic Techniques and Growth Media………………………...... 19 Lab 3. Preparation of Bacterial Smears and Introduction to Staining…………………...... 37 Lab 4. Acid fast and Endospore Staining……………………………………………......... 49 Lab 5. Metabolic Activities of Bacteria…………………………………………….…....... 59 Lab 6. Dichotomous Keys……………………………………………………………......... 77 Lab 7. The Effect of Physical Factors on Microbial Growth……………………………... 85 Lab 8. Chemical Control of Microbial Growth—Disinfectants and Antibiotics…………. 99 Lab 9. The Microbiology of Milk and Food………………………………………………. 111 Lab 10. The Eukaryotes………………………………………………………………........ 123 Lab 11. Clinical Microbiology I; Anaerobic pathogens; Vectors of Infectious Disease….. 141 Lab 12. Clinical Microbiology II—Immunology and the Biolog System………………… 153 Lab 13. Putting it all Together: Case Studies in Microbiology…………………………… 163 Appendix I. -

Marine Extremophiles: a Source of Hydrolases for Biotechnological Applications
Mar. Drugs 2015, 13, 1925-1965; doi:10.3390/md13041925 OPEN ACCESS marine drugs ISSN 1660-3397 www.mdpi.com/journal/marinedrugs Article Marine Extremophiles: A Source of Hydrolases for Biotechnological Applications Gabriel Zamith Leal Dalmaso 1,2, Davis Ferreira 3 and Alane Beatriz Vermelho 1,* 1 BIOINOVAR—Biotechnology laboratories: Biocatalysis, Bioproducts and Bioenergy, Institute of Microbiology Paulo de Góes, Federal University of Rio de Janeiro, Av. Carlos Chagas Filho, 373, 21941-902 Rio de Janeiro, Brazil; E-Mail: [email protected] 2 Graduate Program in Plant Biotechnology, Health and Science Centre, Federal University of Rio de Janeiro, Av. Carlos Chagas Filho, 373, 21941-902 Rio de Janeiro, Brazil 3 BIOINOVAR—Biotechnology Laboratories: Virus-Cell Interaction, Institute of Microbiology Paulo de Góes, Federal University of Rio de Janeiro, Av. Carlos Chagas Filho, 373, 21941-902 Rio de Janeiro, Brazil; E-Mail: [email protected] * Author to whom correspondence should be addressed; E-Mail: [email protected]; Tel.: +55-(21)-3936-6743; Fax: +55-(21)-2560-8344. Academic Editor: Kirk Gustafson Received: 1 December 2014 / Accepted: 25 March 2015 / Published: 3 April 2015 Abstract: The marine environment covers almost three quarters of the planet and is where evolution took its first steps. Extremophile microorganisms are found in several extreme marine environments, such as hydrothermal vents, hot springs, salty lakes and deep-sea floors. The ability of these microorganisms to support extremes of temperature, salinity and pressure demonstrates their great potential for biotechnological processes. Hydrolases including amylases, cellulases, peptidases and lipases from hyperthermophiles, psychrophiles, halophiles and piezophiles have been investigated for these reasons. -
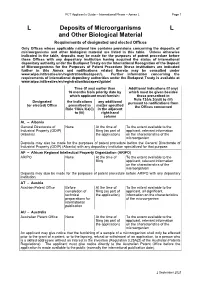
L Deposits of Microorganisms and Other Biological Material L
PCT Applicant’s Guide – International Phase – Annex L Page 1 L Deposits of Microorganisms L and Other Biological Material Requirements of designated and elected Offices Only Offices whose applicable national law contains provisions concerning the deposits of microorganisms and other biological material are listed in this table. Unless otherwise indicated in the table, deposits may be made for the purposes of patent procedure before these Offices with any depositary institution having acquired the status of international depositary authority under the Budapest Treaty on the International Recognition of the Deposit of Microorganisms for the Purposes of Patent Procedure (these institutions are indicated further in this Annex and notifications related thereto may be consulted under www.wipo.int/treaties/en/registration/budapest/). Further information concerning the requirements of international depository authorities under the Budapest Treaty is available at www.wipo.int/treaties/en/registration/budapest/guide/ Time (if any) earlier than Additional indications (if any) 16 months from priority date by which must be given besides which applicant must furnish: those prescribed in Rule 13bis.3(a)(i) to (iii) Designated the indications any additional pursuant to notifications from (or elected) Office prescribed in matter specified the Offices concerned Rule 13bis.3(a)(i) in the adjacent to (iii) right-hand column AL – Albania General Directorate of None At the time of To the extent available to the Industrial Property (GDIP) filing (as part of applicant, relevant information (Albania) the application) on the characteristics of the microorganism Deposits may also be made for the purposes of patent procedure before the General Directorate of Industrial Property (GDIP) (Albania) with any depositary institution specialized for that purpose. -

Origin and Evolution of Eukaryotic Compartmentalization
TESI DOCTORAL UPF 2016 Origin and evolution of eukaryotic compartmentalization Alexandros Pittis Director Dr. Toni Gabaldon´ Comparative Genomics Group Bioinformatics and Genomics Department Centre for Genomic Regulation (CRG) To my father Stavros for the PhD he never did Acknowledgments Looking back at the years of my PhD in the Comparative Genomics group at CRG, I feel it was equally a process of scientific, as well as personal development. All this time I was very privileged to be surrounded by people that offered me their generous support in both fronts. And they were available in the very moments that I was fighting more myself than the unsolvable (anyway) comparative genomics puzzles. I hope that in the future I will have many times the chance to express them my appreciation, way beyond these few words. First, to Toni, my supervisor, for all his support, and patience, and confidence, and respect, especially at the moments that things did not seem that promising. He offered me freedom, to try, to think, to fail, to learn, to achieve that few enjoy during their PhD, and I am very thankful to him. Then, to my good friends and colleagues, those that I found in the group already, and others that joined after me. A very special thanks to Marinita and Jaime, for all their valuable time, and guidance and paradigm, which nevertheless I never managed to follow. They have both marked my phylogenomics path so far and I cannot escape. To Les and Salvi, my fellow students and pals at the time for all that we shared; to Gab and Fran and Dam, for -

Antibiotics from Deep-Sea Microorganisms: Current Discoveries and Perspectives
marine drugs Review Antibiotics from Deep-Sea Microorganisms: Current Discoveries and Perspectives Emiliana Tortorella 1,†, Pietro Tedesco 1,2,†, Fortunato Palma Esposito 1,3,†, Grant Garren January 1,†, Renato Fani 4, Marcel Jaspars 5 and Donatella de Pascale 1,3,* 1 Institute of Protein Biochemistry, National Research Council, I-80131 Naples, Italy; [email protected] (E.T.); [email protected] (P.T.); [email protected] (F.P.E.); [email protected] (G.G.J.) 2 Laboratoire d’Ingénierie des Systèmes Biologiques et des Procédés, INSA, 31400 Toulouse, France 3 Stazione Zoologica “Anthon Dorn”, Villa Comunale, I-80121 Naples, Italy 4 Department of Biology, University of Florence, Sesto Fiorentino, I-50019 Florence, Italy; renato.fani@unifi.it 5 Marine Biodiscovery Centre, Department of Chemistry, University of Aberdeen, Aberdeen, Scotland AB24 3UE, UK; [email protected] * Correspondence: [email protected]; Tel.: +39-0816-132-314; Fax: +39-0816-132-277 † These authors equally contributed to the work. Received: 23 July 2018; Accepted: 27 September 2018; Published: 29 September 2018 Abstract: The increasing emergence of new forms of multidrug resistance among human pathogenic bacteria, coupled with the consequent increase of infectious diseases, urgently requires the discovery and development of novel antimicrobial drugs with new modes of action. Most of the antibiotics currently available on the market were obtained from terrestrial organisms or derived semisynthetically from fermentation products. The isolation of microorganisms from previously unexplored habitats may lead to the discovery of lead structures with antibiotic activity. The deep-sea environment is a unique habitat, and deep-sea microorganisms, because of their adaptation to this extreme environment, have the potential to produce novel secondary metabolites with potent biological activities. -

Role of the Cell-Wall Structure in the Retention of Bacteria by Microfiltration Membranes
Open Archive Toulouse Archive Ouverte (OATAO) OATAO is an open access repository that collects the work of Toulouse researchers and makes it freely available over the web where possible. This is an author-deposited version published in: http://oatao.univ-toulouse.fr/ Eprints ID: 5910 To link to this article: DOI:10.1016/J.MEMSCI.2008.09.049 URL: http://dx.doi.org/10.1016/J.MEMSCI.2008.09.049 To cite this version: Lebleu, Nathalie and Roques, Christine and Aimar, Pierre and Causserand, Christel (2009) Role of the cell-wall structure in the retention of bacteria by microfiltration membranes. Journal of Membrane Science, vol. 326 (n°1). pp. 178-185. ISSN 0376-7388 Any correspondence concerning this service should be sent to the repository administrator: [email protected] Role of the cell•wall structure in the retention of bacteria by microfiltration membranes Nathalie Lebleu a, Christine Roques b, Pierre Aimar a, Christel Causserand a,∗ a Laboratoire de Génie Chimique, Université Paul Sabatier, 31062 Toulouse Cedex 9, France b LU 49, Adhésion Bactérienne et Formation de Biofilms, Université Paul Sabatier, 31062 Toulouse Cedex 9, France a b s t r a c t This experimental study investigates the retention of bacteria by porous membranes. The transfer of bacteria larger than the nominal pore size of microfiltration track•etched membranes has been studied for several kinds of bacterial strains. This unexpected transfer does not correlate to the hydrophobicity, neither to the surface charge of the microorganism, as suggested in previous reports. We conclude that, in our conditions, the kind of bacteria (Gram•positive or Gram•negative) is finally the most important parameter. -

Spatial Separation of Ribosomes and DNA in Asgard Archaeal Cells ✉ ✉ Burak Avcı 1,2 , Jakob Brandt3, Dikla Nachmias4, Natalie Elia4, Mads Albertsen 3, Thijs J
www.nature.com/ismej BRIEF COMMUNICATION OPEN Spatial separation of ribosomes and DNA in Asgard archaeal cells ✉ ✉ Burak Avcı 1,2 , Jakob Brandt3, Dikla Nachmias4, Natalie Elia4, Mads Albertsen 3, Thijs J. G. Ettema 2, Andreas Schramm1,5 and Kasper Urup Kjeldsen1 © The Author(s) 2021 The origin of the eukaryotic cell is a major open question in biology. Asgard archaea are the closest known prokaryotic relatives of eukaryotes, and their genomes encode various eukaryotic signature proteins, indicating some elements of cellular complexity prior to the emergence of the first eukaryotic cell. Yet, microscopic evidence to demonstrate the cellular structure of uncultivated Asgard archaea in the environment is thus far lacking. We used primer-free sequencing to retrieve 715 almost full-length Loki- and Heimdallarchaeota 16S rRNA sequences and designed novel oligonucleotide probes to visualize their cells in marine sediments (Aarhus Bay, Denmark) using catalyzed reporter deposition-fluorescence in situ hybridization (CARD-FISH). Super-resolution microscopy revealed 1–2 µm large, coccoid cells, sometimes occurring as aggregates. Remarkably, the DNA staining was spatially separated from ribosome-originated FISH signals by 50–280 nm. This suggests that the genomic material is condensed and spatially distinct in a particular location and could indicate compartmentalization or membrane invagination in Asgard archaeal cells. The ISME Journal; https://doi.org/10.1038/s41396-021-01098-3 INTRODUCTION cells in marine sediments [8] yet methodical limitations did not The origin of the eukaryotic cell is a major unresolved puzzle in allow to discern single cells and to draw any in-depth conclusion the history of life. -

Archaea and the Origin of Eukaryotes
REVIEWS Archaea and the origin of eukaryotes Laura Eme, Anja Spang, Jonathan Lombard, Courtney W. Stairs and Thijs J. G. Ettema Abstract | Woese and Fox’s 1977 paper on the discovery of the Archaea triggered a revolution in the field of evolutionary biology by showing that life was divided into not only prokaryotes and eukaryotes. Rather, they revealed that prokaryotes comprise two distinct types of organisms, the Bacteria and the Archaea. In subsequent years, molecular phylogenetic analyses indicated that eukaryotes and the Archaea represent sister groups in the tree of life. During the genomic era, it became evident that eukaryotic cells possess a mixture of archaeal and bacterial features in addition to eukaryotic-specific features. Although it has been generally accepted for some time that mitochondria descend from endosymbiotic alphaproteobacteria, the precise evolutionary relationship between eukaryotes and archaea has continued to be a subject of debate. In this Review, we outline a brief history of the changing shape of the tree of life and examine how the recent discovery of a myriad of diverse archaeal lineages has changed our understanding of the evolutionary relationships between the three domains of life and the origin of eukaryotes. Furthermore, we revisit central questions regarding the process of eukaryogenesis and discuss what can currently be inferred about the evolutionary transition from the first to the last eukaryotic common ancestor. Sister groups Two descendants that split The pioneering work by Carl Woese and colleagues In this Review, we discuss how culture- independent from the same node; the revealed that all cellular life could be divided into three genomics has transformed our understanding of descendants are each other’s major evolutionary lines (also called domains): the archaeal diversity and how this has influenced our closest relative.