ERK1/2 MAP Kinases: Structure, Function, and Regulation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Nuclear Pore Proteins and the Control of Genome Functions
Downloaded from genesdev.cshlp.org on September 30, 2021 - Published by Cold Spring Harbor Laboratory Press REVIEW Nuclear pore proteins and the control of genome functions Arkaitz Ibarra and Martin W. Hetzer Molecular and Cell Biology Laboratory, Salk Institute for Biological Studies, La Jolla, California 92037, USA Nuclear pore complexes (NPCs) are composed of several Cytoplasmic filaments are mainly formed by Nup358/ copies of ~30 different proteins called nucleoporins (Nups). RanBP2, Nup214, and Nup88, while the nuclear basket is NPCs penetrate the nuclear envelope (NE) and regulate the composed of Nup153 and Tpr (Fig. 1; for Nup othologs, see nucleocytoplasmic trafficking of macromolecules. Beyond Rothballer and Kutay 2012). this vital role, NPC components influence genome func- The selective access of regulatory factors into the tions in a transport-independent manner. Nups play an nucleus and export of specific RNA molecules mediated evolutionarily conserved role in gene expression regulation by the NPC is required for the accurate progression of most that, in metazoans, extends into the nuclear interior. major cellular processes. However, our perception of Additionally, in proliferative cells, Nups play a crucial role the NPC components is rapidly evolving, as accumulating in genome integrity maintenance and mitotic progression. evidence indicates that they can also directly impact Here we discuss genome-related functions of Nups and DNA metabolism by genome-related functions (Liang their impact on essential DNA metabolism processes such and Hetzer 2011). Among these, one of the most remark- as transcription, chromosome duplication, and segregation. able and well-conserved roles of Nups is to associate with specific target genes to regulate their transcriptional activity (Casolari et al. -

Casein Kinase 1 Isoforms in Degenerative Disorders
CASEIN KINASE 1 ISOFORMS IN DEGENERATIVE DISORDERS DISSERTATION Presented in Partial Fulfillment of the Requirements for the Degree Doctor of Philosophy in the Graduate School of The Ohio State University By Theresa Joseph Kannanayakal, M.Sc., M.S. * * * * * The Ohio State University 2004 Dissertation Committee: Approved by Professor Jeff A. Kuret, Adviser Professor John D. Oberdick Professor Dale D. Vandre Adviser Professor Mike X. Zhu Biophysics Graduate Program ABSTRACT Casein Kinase 1 (CK1) enzyme is one of the largest family of Serine/Threonine protein kinases. CK1 has a wide distribution spanning many eukaryotic families. In cells, its kinase activity has been found in various sub-cellular compartments enabling it to phosphorylate many proteins involved in cellular maintenance and disease pathogenesis. Tau is one such substrate whose hyperphosphorylation results in degeneration of neurons in Alzheimer’s disease (AD). AD is a slow neuroprogessive disorder histopathologically characterized by Granulovacuolar degeneration bodies (GVBs) and intraneuronal accumulation of tau in Neurofibrillary Tangles (NFTs). The level of CK1 isoforms, CK1α, CK1δ and CK1ε has been shown to be elevated in AD. Previous studies of the correlation of CK1δ with lesions had demonstrated its importance in tau hyperphosphorylation. Hence we investigated distribution of CK1α and CK1ε with the lesions to understand if they would play role in tau hyperphosphorylation similar to CK1δ. The kinase results were also compared with lesion correlation studies of peptidyl cis/trans prolyl isomerase (Pin1) and caspase-3. Our results showed that among the enzymes investigated, CK1 isoforms have the greatest extent of colocalization with the lesions. We have also investigated the distribution of CK1α with different stages of NFTs that follow AD progression. -

Product Data Sheet Purified Anti-NUP153
Version: 2 Revision Date: 2016-01-08 Product Data Sheet Purified anti-NUP153 Catalog # / Size: 906201 / 100 µl Previously: Covance Catalog# MMS-102P Clone: QE5 Isotype: Mouse IgG1 Immunogen: The QE5 monoclonal antibody was generated against rat liver nuclear envelope proteins. Reactivity: Eukaryote Preparation: The antibody was purified by affinity chromatography. Formulation: Phosphate-buffered solution + 0.03% thimerosal. Concentration: 1 mg/ml Storage: The antibody solution should be stored undiluted between 2°C and 8°C. Please note the storage condition for this antibody has been changed from -20°C to between 2°C and 8°C. You can also check your vial or your Methanol fixed HeLa stained with the CoA to find the most accurate storage condition for this antibody. antibody QE5. This antibody brilliantly highlights the nuclear membrane (green). The golgi is stained with the Applications: antibody to Giantin. Applications: ICC, WB, IF, IP IEM - Reported in literature Recommended Usage: Each lot of this antibody is quality control tested by Immunocytochemistry. The optimal working dilution should be determined for each specific assay condition. • WB: 1:500* • IF: 1:250 • IP: 1:50 Application Notes: This antibody is effective in immunoblotting, immunofluorescence (IF) and immunoprecipitation (IP). *Predicted MW = 250 kD This antibody recognizes NUP153 as well as two related nuclear pore complex proteins: NUP214 and p62. By immunofluorescence, QE5 labels the nuclear envelope of eukaryotic cells giving a punctate staining pattern. Application References: 1. Pare GC, Easlick JL, Mislow JM, McNally EM, Kapiloff MS. Nesprin-1alpha contributes to the targeting of mAKAP to the cardiac myocyte nuclear envelope. -
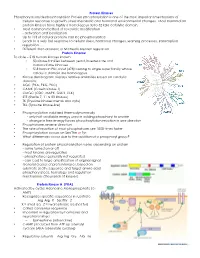
Protein Kinases Phosphorylation/Dephosphorylation Protein Phosphorylation Is One of the Most Important Mechanisms of Cellular Re
Protein Kinases Phosphorylation/dephosphorylation Protein phosphorylation is one of the most important mechanisms of cellular responses to growth, stress metabolic and hormonal environmental changes. Most mammalian protein kinases have highly a homologous 30 to 32 kDa catalytic domain. • Most common method of reversible modification - activation and localization • Up to 1/3 of cellular proteins can be phosphorylated • Leads to a very fast response to cellular stress, hormonal changes, learning processes, transcription regulation .... • Different than allosteric or Michealis Menten regulation Protein Kinome To date – 518 human kinases known • 50 kinase families between yeast, invertebrate and mammaliane kinomes • 518 human PKs, most (478) belong to single super family whose catalytic domain are homologous. • Kinase dendrogram displays relative similarities based on catalytic domains. • AGC (PKA, PKG, PKC) • CAMK (Casein kinase 1) • CMGC (CDC, MAPK, GSK3, CLK) • STE (Sterile 7, 11 & 20 kinases) • TK (Tryosine kinases memb and cyto) • TKL (Tyrosine kinase-like) • Phosphorylation stabilized thermodynamically - only half available energy used in adding phosphoryl to protein - change in free energy forces phosphorylation reaction in one direction • Phosphatases reverse direction • The rate of reaction of most phosphatases are 1000 times faster • Phosphorylation occurs on Ser/The or Tyr • What differences occur due to the addition of a phosphoryl group? • Regulation of protein phosphorylation varies depending on protein - some turned on or off -
HCC and Cancer Mutated Genes Summarized in the Literature Gene Symbol Gene Name References*
HCC and cancer mutated genes summarized in the literature Gene symbol Gene name References* A2M Alpha-2-macroglobulin (4) ABL1 c-abl oncogene 1, receptor tyrosine kinase (4,5,22) ACBD7 Acyl-Coenzyme A binding domain containing 7 (23) ACTL6A Actin-like 6A (4,5) ACTL6B Actin-like 6B (4) ACVR1B Activin A receptor, type IB (21,22) ACVR2A Activin A receptor, type IIA (4,21) ADAM10 ADAM metallopeptidase domain 10 (5) ADAMTS9 ADAM metallopeptidase with thrombospondin type 1 motif, 9 (4) ADCY2 Adenylate cyclase 2 (brain) (26) AJUBA Ajuba LIM protein (21) AKAP9 A kinase (PRKA) anchor protein (yotiao) 9 (4) Akt AKT serine/threonine kinase (28) AKT1 v-akt murine thymoma viral oncogene homolog 1 (5,21,22) AKT2 v-akt murine thymoma viral oncogene homolog 2 (4) ALB Albumin (4) ALK Anaplastic lymphoma receptor tyrosine kinase (22) AMPH Amphiphysin (24) ANK3 Ankyrin 3, node of Ranvier (ankyrin G) (4) ANKRD12 Ankyrin repeat domain 12 (4) ANO1 Anoctamin 1, calcium activated chloride channel (4) APC Adenomatous polyposis coli (4,5,21,22,25,28) APOB Apolipoprotein B [including Ag(x) antigen] (4) AR Androgen receptor (5,21-23) ARAP1 ArfGAP with RhoGAP domain, ankyrin repeat and PH domain 1 (4) ARHGAP35 Rho GTPase activating protein 35 (21) ARID1A AT rich interactive domain 1A (SWI-like) (4,5,21,22,24,25,27,28) ARID1B AT rich interactive domain 1B (SWI1-like) (4,5,22) ARID2 AT rich interactive domain 2 (ARID, RFX-like) (4,5,22,24,25,27,28) ARID4A AT rich interactive domain 4A (RBP1-like) (28) ARID5B AT rich interactive domain 5B (MRF1-like) (21) ASPM Asp (abnormal -

Antigen-Specific Memory CD4 T Cells Coordinated Changes in DNA
Downloaded from http://www.jimmunol.org/ by guest on September 24, 2021 is online at: average * The Journal of Immunology The Journal of Immunology published online 18 March 2013 from submission to initial decision 4 weeks from acceptance to publication http://www.jimmunol.org/content/early/2013/03/17/jimmun ol.1202267 Coordinated Changes in DNA Methylation in Antigen-Specific Memory CD4 T Cells Shin-ichi Hashimoto, Katsumi Ogoshi, Atsushi Sasaki, Jun Abe, Wei Qu, Yoichiro Nakatani, Budrul Ahsan, Kenshiro Oshima, Francis H. W. Shand, Akio Ametani, Yutaka Suzuki, Shuichi Kaneko, Takashi Wada, Masahira Hattori, Sumio Sugano, Shinichi Morishita and Kouji Matsushima J Immunol Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Author Choice option Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Freely available online through http://www.jimmunol.org/content/suppl/2013/03/18/jimmunol.120226 7.DC1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material Permissions Email Alerts Subscription Author Choice Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 24, 2021. Published March 18, 2013, doi:10.4049/jimmunol.1202267 The Journal of Immunology Coordinated Changes in DNA Methylation in Antigen-Specific Memory CD4 T Cells Shin-ichi Hashimoto,*,†,‡ Katsumi Ogoshi,* Atsushi Sasaki,† Jun Abe,* Wei Qu,† Yoichiro Nakatani,† Budrul Ahsan,x Kenshiro Oshima,† Francis H. -

Supplementary Table 1. in Vitro Side Effect Profiling Study for LDN/OSU-0212320. Neurotransmitter Related Steroids
Supplementary Table 1. In vitro side effect profiling study for LDN/OSU-0212320. Percent Inhibition Receptor 10 µM Neurotransmitter Related Adenosine, Non-selective 7.29% Adrenergic, Alpha 1, Non-selective 24.98% Adrenergic, Alpha 2, Non-selective 27.18% Adrenergic, Beta, Non-selective -20.94% Dopamine Transporter 8.69% Dopamine, D1 (h) 8.48% Dopamine, D2s (h) 4.06% GABA A, Agonist Site -16.15% GABA A, BDZ, alpha 1 site 12.73% GABA-B 13.60% Glutamate, AMPA Site (Ionotropic) 12.06% Glutamate, Kainate Site (Ionotropic) -1.03% Glutamate, NMDA Agonist Site (Ionotropic) 0.12% Glutamate, NMDA, Glycine (Stry-insens Site) 9.84% (Ionotropic) Glycine, Strychnine-sensitive 0.99% Histamine, H1 -5.54% Histamine, H2 16.54% Histamine, H3 4.80% Melatonin, Non-selective -5.54% Muscarinic, M1 (hr) -1.88% Muscarinic, M2 (h) 0.82% Muscarinic, Non-selective, Central 29.04% Muscarinic, Non-selective, Peripheral 0.29% Nicotinic, Neuronal (-BnTx insensitive) 7.85% Norepinephrine Transporter 2.87% Opioid, Non-selective -0.09% Opioid, Orphanin, ORL1 (h) 11.55% Serotonin Transporter -3.02% Serotonin, Non-selective 26.33% Sigma, Non-Selective 10.19% Steroids Estrogen 11.16% 1 Percent Inhibition Receptor 10 µM Testosterone (cytosolic) (h) 12.50% Ion Channels Calcium Channel, Type L (Dihydropyridine Site) 43.18% Calcium Channel, Type N 4.15% Potassium Channel, ATP-Sensitive -4.05% Potassium Channel, Ca2+ Act., VI 17.80% Potassium Channel, I(Kr) (hERG) (h) -6.44% Sodium, Site 2 -0.39% Second Messengers Nitric Oxide, NOS (Neuronal-Binding) -17.09% Prostaglandins Leukotriene, -

The Role of Casein Kinase 1 in Meiosis
e & Ch m rom ro o d s n o y m S Zhao and Liang, J Down Syndr Chr Abnorm 2016, 2:1 e n A Journal of Down Syndrome & w b o n DOI: 10.4172/2472-1115.1000106 D o f r m o l ISSN: 2472-1115a a l i n t r i e u s o J Chromosome Abnormalities Review Article Open Access The Role of Casein Kinase 1 in Meiosis Yue-Fang Zhao and Cheng-Guang Liang* The Key Laboratory of National Education Ministry for Mammalian Reproductive Biology and Biotechnology, The Research Center for Laboratory Animal Science, College of Life Science, Inner Mongolia University, Hohhot, Inner Mongolia, People’s Republic of China Abstract The casein kinase 1 (CK1) belongs to the serine/threonine protein kinases. CK1 members, which commonly exist in all eukaryotes, are involved in the regulation of many cellular processes linked to cell cycle progression, spindle- dynamics, and chromosome segregation. Additionally, CK1 regulate key signaling pathways such as Wnt (Wingless/ Int-1), Hh (Hedgehog), and Hippo, known to be critically involved in tumor progression. Considering the importance of CK1 for accurate cell division and regulation of tumor suppressor functions, it is not surprising scientific effort has enormously increased. In mammals, CK1 regulate the transition from interphase to metaphase in mitosis. In budding yeast and fission yeast, CK1 phosphorylate Rec8 subunits of cohesin complex and regulate chromosome segregation in meiosis. During meiosis, two rounds of chromosome segregation after a single round of DNA replication produce haploid gametes from diploid precursors. Any mistake in chromosome segregation may result in aneuploidy, which in meiosis are one of the main causes of infertility, abortion and many genetic diseases in humans. -

Identification of a Potential Mitotic Function for the Mammalian Nup50
Identification of a Potential Mitotic Function for the Mammalian Nup50 A Senior Thesis Presented in Partial Fulfillment of the Requirements for graduation with research distinction in Biology in the undergraduate colleges of The Ohio State University by Jessica El-Hallal The Ohio State University June 2011 Project Advisor: Dr. Stephen Osmani, Department of Molecular Genetics ABSTRACT Mitosis is a conserved process in which the genetic material, DNA, is equally segregated between two daughter cells. DNA is contained in the nucleus of the eukaryotic cell and surrounded by the nuclear envelope. Multi protein complexes known as the Nuclear Pore Complexes (NPCs) embed within the nuclear envelope and regulate the transport of molecules in and out of the nucleus. Surprisingly, in Aspergillus nidulans, the model system used in my study, a nuclear pore complex protein Nup2 undergoes a unique translocation to chromatin during mitosis and is essential for proper mitotic progression. Interestingly, the Nup2 homolog in higher eukaryotes, Nup50, undergoes the same translocation. Therefore, the purpose of this study is to test whether Nup50 can translocate onto chromatin in Aspergillus nidulans and complement the mitotic function of Nup2. In order to test this hypothesis, the Nup50 gene was integrated into A. nidulans using homologous recombination. Four way fusion PCR was used to generate a DNA cassette that contains the Nup50 gene fused to EGFP2 marker and its expression under control of the inducible promoter alcA. Once Nup50 was introduced into A. nidulans, Nup2 was deleted in the background. So far, we have discovered that Nup50 is present in the nucleus at interphase and disperses throughout the cell during mitosis in the absence or presence of the Aspergillus nidulans Nup2. -

PV3320 Certificate of Analysis for Lot 1322684A
Certificate of Analysis MAPK8 (JNK1), Inactive , 100 µg Mitogen-activated Protein Kinase 8 (MAP), Inactive, Histidine-tagged Part Number: PV3320 5781 Van Allen Way Lot Number: 1322684A Carlsbad, CA 92008 Immediate Storage: -80°C Phone: 760.603.7200 Shipping Conditions: dry ice www.thermofisher.com Description: Storage and Handling: Recombinant human full-length protein, inactive, Histidine-tagged, Store at -80°C. At first use, aliquot and store at -80°C to avoid multiple expressed in insect cells. freeze-thaws. If properly stored at -80°C, this product is guaranteed for 6 Specific Activity: months from date of purchase. Approximately 2% activity as compared to the active form of the kinase. Storage Buffer: 50 mM Tris (pH 7.5), 150 mM NaCl, 0.5 mM EDTA, 0.05% Triton® X–100, Can be activated by the upstream kinase MAP2K7, mutant. 2 mM DTT and 50% Glycerol. Concentration: 0.87 mg/mL total protein as measured using the Bradford protein assay with BSA as a standard. Calculated 18,300 nM. Aliases: JNK, JNK1, SAPK1 QUALITY ASSURANCE Activation Test: Gel Information for MAPK8 (JNK1), Inactive The coupled MAPK8 activation assay uses active MAP2K7, mutant to Page Description: The SDS- phosphorylate ATF2 substrate. The assay was setup in two states: 1st PAGE and/or Native PAGE MAPK8 phosphorylation by MAP2K7, mutant without 32P-ATP; and 2nd were run on 4-20% Tris-Glycine ATF2 substrate phosphorylation by activated MAPK8 in the presence of Novex® gels (Catalog #: 32P-ATP. The basal activity of MAPK8 was also assayed in the absence of EC6025BOX). MAP2K7, mutant. Lane 1: Invitrogen™ BenchMark™ Protein Ladder (Catalog #: 10747-012). -

LIM and Cysteine-Rich Domains 1 (LMCD1) Regulates Skeletal Muscle Hypertrophy, Calcium Handling, and Force Duarte M
Ferreira et al. Skeletal Muscle (2019) 9:26 https://doi.org/10.1186/s13395-019-0214-1 RESEARCH Open Access LIM and cysteine-rich domains 1 (LMCD1) regulates skeletal muscle hypertrophy, calcium handling, and force Duarte M. S. Ferreira1, Arthur J. Cheng2,3, Leandro Z. Agudelo1,4, Igor Cervenka1, Thomas Chaillou2,5, Jorge C. Correia1, Margareta Porsmyr-Palmertz1, Manizheh Izadi1,6, Alicia Hansson1, Vicente Martínez-Redondo1, Paula Valente-Silva1, Amanda T. Pettersson-Klein1, Jennifer L. Estall7, Matthew M. Robinson8, K. Sreekumaran Nair8, Johanna T. Lanner2 and Jorge L. Ruas1* Abstract Background: Skeletal muscle mass and strength are crucial determinants of health. Muscle mass loss is associated with weakness, fatigue, and insulin resistance. In fact, it is predicted that controlling muscle atrophy can reduce morbidity and mortality associated with diseases such as cancer cachexia and sarcopenia. Methods: We analyzed gene expression data from muscle of mice or human patients with diverse muscle pathologies and identified LMCD1 as a gene strongly associated with skeletal muscle function. We transiently expressed or silenced LMCD1 in mouse gastrocnemius muscle or in mouse primary muscle cells and determined muscle/cell size, targeted gene expression, kinase activity with kinase arrays, protein immunoblotting, and protein synthesis levels. To evaluate force, calcium handling, and fatigue, we transduced the flexor digitorum brevis muscle with a LMCD1-expressing adenovirus and measured specific force and sarcoplasmic reticulum Ca2+ release in individual fibers. Finally, to explore the relationship between LMCD1 and calcineurin, we ectopically expressed Lmcd1 in the gastrocnemius muscle and treated those mice with cyclosporine A (calcineurin inhibitor). In addition, we used a luciferase reporter construct containing the myoregulin gene promoter to confirm the role of a LMCD1- calcineurin-myoregulin axis in skeletal muscle mass control and calcium handling. -

Persistent Transcription-Blocking DNA Lesions Trigger Somatic Growth Attenuation Associated with Longevity
ARTICLES Persistent transcription-blocking DNA lesions trigger somatic growth attenuation associated with longevity George A. Garinis1,2, Lieneke M. Uittenboogaard1, Heike Stachelscheid3,4, Maria Fousteri5, Wilfred van Ijcken6, Timo M. Breit7, Harry van Steeg8, Leon H. F. Mullenders5, Gijsbertus T. J. van der Horst1, Jens C. Brüning4,9, Carien M. Niessen3,9,10, Jan H. J. Hoeijmakers1 and Björn Schumacher1,9,11 The accumulation of stochastic DNA damage throughout an organism’s lifespan is thought to contribute to ageing. Conversely, ageing seems to be phenotypically reproducible and regulated through genetic pathways such as the insulin-like growth factor-1 (IGF-1) and growth hormone (GH) receptors, which are central mediators of the somatic growth axis. Here we report that persistent DNA damage in primary cells from mice elicits changes in global gene expression similar to those occurring in various organs of naturally aged animals. We show that, as in ageing animals, the expression of IGF-1 receptor and GH receptor is attenuated, resulting in cellular resistance to IGF-1. This cell-autonomous attenuation is specifically induced by persistent lesions leading to stalling of RNA polymerase II in proliferating, quiescent and terminally differentiated cells; it is exacerbated and prolonged in cells from progeroid mice and confers resistance to oxidative stress. Our findings suggest that the accumulation of DNA damage in transcribed genes in most if not all tissues contributes to the ageing-associated shift from growth to somatic maintenance that triggers stress resistance and is thought to promote longevity. Ageing represents the progressive functional decline that is exempted levels as a result of pituitary dysfunction (Snell and Ames mice) — have an from evolutionary selection because it largely occurs after reproduc- extended lifespan17–20.