Gingival Transcriptomics of Follicular T Cell Footprints in Progressing
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
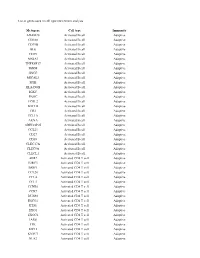
List of Genes Used in Cell Type Enrichment Analysis
List of genes used in cell type enrichment analysis Metagene Cell type Immunity ADAM28 Activated B cell Adaptive CD180 Activated B cell Adaptive CD79B Activated B cell Adaptive BLK Activated B cell Adaptive CD19 Activated B cell Adaptive MS4A1 Activated B cell Adaptive TNFRSF17 Activated B cell Adaptive IGHM Activated B cell Adaptive GNG7 Activated B cell Adaptive MICAL3 Activated B cell Adaptive SPIB Activated B cell Adaptive HLA-DOB Activated B cell Adaptive IGKC Activated B cell Adaptive PNOC Activated B cell Adaptive FCRL2 Activated B cell Adaptive BACH2 Activated B cell Adaptive CR2 Activated B cell Adaptive TCL1A Activated B cell Adaptive AKNA Activated B cell Adaptive ARHGAP25 Activated B cell Adaptive CCL21 Activated B cell Adaptive CD27 Activated B cell Adaptive CD38 Activated B cell Adaptive CLEC17A Activated B cell Adaptive CLEC9A Activated B cell Adaptive CLECL1 Activated B cell Adaptive AIM2 Activated CD4 T cell Adaptive BIRC3 Activated CD4 T cell Adaptive BRIP1 Activated CD4 T cell Adaptive CCL20 Activated CD4 T cell Adaptive CCL4 Activated CD4 T cell Adaptive CCL5 Activated CD4 T cell Adaptive CCNB1 Activated CD4 T cell Adaptive CCR7 Activated CD4 T cell Adaptive DUSP2 Activated CD4 T cell Adaptive ESCO2 Activated CD4 T cell Adaptive ETS1 Activated CD4 T cell Adaptive EXO1 Activated CD4 T cell Adaptive EXOC6 Activated CD4 T cell Adaptive IARS Activated CD4 T cell Adaptive ITK Activated CD4 T cell Adaptive KIF11 Activated CD4 T cell Adaptive KNTC1 Activated CD4 T cell Adaptive NUF2 Activated CD4 T cell Adaptive PRC1 Activated -

Markers of T Cell Senescence in Humans
International Journal of Molecular Sciences Review Markers of T Cell Senescence in Humans Weili Xu 1,2 and Anis Larbi 1,2,3,4,5,* 1 Biology of Aging Program and Immunomonitoring Platform, Singapore Immunology Network (SIgN), Agency for Science Technology and Research (A*STAR), Immunos Building, Biopolis, Singapore 138648, Singapore; [email protected] 2 School of Biological Sciences, Nanyang Technological University, Singapore 637551, Singapore 3 Department of Microbiology, National University of Singapore, Singapore 117597, Singapore 4 Department of Geriatrics, Faculty of Medicine, University of Sherbrooke, Sherbrooke, QC J1K 2R1, Canada 5 Faculty of Sciences, University ElManar, Tunis 1068, Tunisia * Correspondence: [email protected]; Tel.: +65-6407-0412 Received: 31 May 2017; Accepted: 26 July 2017; Published: 10 August 2017 Abstract: Many countries are facing the aging of their population, and many more will face a similar obstacle in the near future, which could be a burden to many healthcare systems. Increased susceptibility to infections, cardiovascular and neurodegenerative disease, cancer as well as reduced efficacy of vaccination are important matters for researchers in the field of aging. As older adults show higher prevalence for a variety of diseases, this also implies higher risk of complications, including nosocomial infections, slower recovery and sequels that may reduce the autonomy and overall quality of life of older adults. The age-related effects on the immune system termed as “immunosenescence” can be exemplified by the reported hypo-responsiveness to influenza vaccination of the elderly. T cells, which belong to the adaptive arm of the immune system, have been extensively studied and the knowledge gathered enables a better understanding of how the immune system may be affected after acute/chronic infections and how this matters in the long run. -

Human Β‑1,3‑Glucuronyltransferase 1/B3GAT1 Antibody Monoclonal Mouse Igg2a Clone # 1002707 Catalog Number: MAB8560
Human β‑1,3‑Glucuronyltransferase 1/B3GAT1 Antibody Monoclonal Mouse IgG2A Clone # 1002707 Catalog Number: MAB8560 DESCRIPTION Species Reactivity Human Specificity Detects human β-1,3-Glucuronyltransferase 1/B3GAT1 in direct ELISAs. Source Monoclonal Mouse IgG2A Clone # 1002707 Purification Protein A or G purified from hybridoma culture supernatant Immunogen Chinese Hamster Ovary cell line, CHO-derived human β‑1,3‑Glucuronyltransferase 1/B3GAT1 His25-Ile334 Accession # Q9P2W7 Formulation Lyophilized from a 0.2 μm filtered solution in PBS with Trehalose. See Certificate of Analysis for details. *Small pack size (-SP) is supplied either lyophilized or as a 0.2 μm filtered solution in PBS. APPLICATIONS Please Note: Optimal dilutions should be determined by each laboratory for each application. General Protocols are available in the Technical Information section on our website. Recommended Sample Concentration Immunohistochemistry 5-25 µg/mL See Below DATA Immunohistochemistry β-1,3-Glucuronyltransferase 1/B3GAT1 in Human Brain. β-1,3-Glucuronyltransferase 1/B3GAT1 was detected in immersion fixed paraffin-embedded sections of human brain (cortex) using Mouse Anti-Human β-1,3- Glucuronyltransferase 1/B3GAT1 Monoclonal Antibody (Catalog # MAB8560) at 5 µg/mL for 1 hour at room temperature followed by incubation with the Anti-Mouse IgG VisUCyte™ HRP Polymer Antibody (Catalog # VC001). Before incubation with the primary antibody, tissue was subjected to heat-induced epitope retrieval using Antigen Retrieval Reagent-Basic (Catalog # CTS013). Tissue was stained using DAB (brown) and counterstained with hematoxylin (blue). Specific staining was localized to cytoplasm in neurons. View our protocol for IHC Staining with VisUCyte HRP Polymer Detection Reagents. PREPARATION AND STORAGE Reconstitution Reconstitute at 0.5 mg/mL in sterile PBS. -

Differential Integrin Adhesome Expression Defines Human Natural
bioRxiv preprint doi: https://doi.org/10.1101/2020.12.01.404806; this version posted December 1, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. Differential integrin adhesome expression defines human natural killer cell residency and developmental stage Everardo Hegewisch Solloa1, Seungmae Seo1, Bethany L. Mundy-Bosse2, Anjali Mishra3,a, Erik Waldman4,b, Sarah Maurrasse4,b, Eli Grunstein4, Thomas J. Connors5, Aharon G. Freud2,6, and Emily M. Mace1 1Department of Pediatrics, Columbia University College of Physicians and Surgeons, New York NY 10032 2Division of Hematology, Department of Internal Medicine, The Ohio State University, Columbus, OH 43210, USA; Comprehensive Cancer Center and The James Cancer Hospital and Solove Research Institute, The Ohio State University, Columbus, OH 43210 33Division of Dermatology, Department of Internal Medicine, The Ohio State University, Columbus, OH 43210, USA; Comprehensive Cancer Center and The James Cancer Hospital and Solove Research Institute, The Ohio State University, Columbus, OH 43210 4Department of Otolaryngology - Head and Neck Surgery, Columbia University Medical Center, New York, New York 10032 5Department of Pediatrics, Division of Pediatric Critical Care and Hospital Medicine, Columbia University Irving Medical Center, New York, NY 10024 6Department of Pathology, The Ohio State University, Columbus, -

(P -Value<0.05, Fold Change≥1.4), 4 Vs. 0 Gy Irradiation
Table S1: Significant differentially expressed genes (P -Value<0.05, Fold Change≥1.4), 4 vs. 0 Gy irradiation Genbank Fold Change P -Value Gene Symbol Description Accession Q9F8M7_CARHY (Q9F8M7) DTDP-glucose 4,6-dehydratase (Fragment), partial (9%) 6.70 0.017399678 THC2699065 [THC2719287] 5.53 0.003379195 BC013657 BC013657 Homo sapiens cDNA clone IMAGE:4152983, partial cds. [BC013657] 5.10 0.024641735 THC2750781 Ciliary dynein heavy chain 5 (Axonemal beta dynein heavy chain 5) (HL1). 4.07 0.04353262 DNAH5 [Source:Uniprot/SWISSPROT;Acc:Q8TE73] [ENST00000382416] 3.81 0.002855909 NM_145263 SPATA18 Homo sapiens spermatogenesis associated 18 homolog (rat) (SPATA18), mRNA [NM_145263] AA418814 zw01a02.s1 Soares_NhHMPu_S1 Homo sapiens cDNA clone IMAGE:767978 3', 3.69 0.03203913 AA418814 AA418814 mRNA sequence [AA418814] AL356953 leucine-rich repeat-containing G protein-coupled receptor 6 {Homo sapiens} (exp=0; 3.63 0.0277936 THC2705989 wgp=1; cg=0), partial (4%) [THC2752981] AA484677 ne64a07.s1 NCI_CGAP_Alv1 Homo sapiens cDNA clone IMAGE:909012, mRNA 3.63 0.027098073 AA484677 AA484677 sequence [AA484677] oe06h09.s1 NCI_CGAP_Ov2 Homo sapiens cDNA clone IMAGE:1385153, mRNA sequence 3.48 0.04468495 AA837799 AA837799 [AA837799] Homo sapiens hypothetical protein LOC340109, mRNA (cDNA clone IMAGE:5578073), partial 3.27 0.031178378 BC039509 LOC643401 cds. [BC039509] Homo sapiens Fas (TNF receptor superfamily, member 6) (FAS), transcript variant 1, mRNA 3.24 0.022156298 NM_000043 FAS [NM_000043] 3.20 0.021043295 A_32_P125056 BF803942 CM2-CI0135-021100-477-g08 CI0135 Homo sapiens cDNA, mRNA sequence 3.04 0.043389246 BF803942 BF803942 [BF803942] 3.03 0.002430239 NM_015920 RPS27L Homo sapiens ribosomal protein S27-like (RPS27L), mRNA [NM_015920] Homo sapiens tumor necrosis factor receptor superfamily, member 10c, decoy without an 2.98 0.021202829 NM_003841 TNFRSF10C intracellular domain (TNFRSF10C), mRNA [NM_003841] 2.97 0.03243901 AB002384 C6orf32 Homo sapiens mRNA for KIAA0386 gene, partial cds. -

Identification of Key Pathways and Genes in Dementia Via Integrated Bioinformatics Analysis
bioRxiv preprint doi: https://doi.org/10.1101/2021.04.18.440371; this version posted July 19, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Identification of Key Pathways and Genes in Dementia via Integrated Bioinformatics Analysis Basavaraj Vastrad1, Chanabasayya Vastrad*2 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karnataka, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India bioRxiv preprint doi: https://doi.org/10.1101/2021.04.18.440371; this version posted July 19, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract To provide a better understanding of dementia at the molecular level, this study aimed to identify the genes and key pathways associated with dementia by using integrated bioinformatics analysis. Based on the expression profiling by high throughput sequencing dataset GSE153960 derived from the Gene Expression Omnibus (GEO), the differentially expressed genes (DEGs) between patients with dementia and healthy controls were identified. With DEGs, we performed a series of functional enrichment analyses. Then, a protein–protein interaction (PPI) network, modules, miRNA-hub gene regulatory network and TF-hub gene regulatory network was constructed, analyzed and visualized, with which the hub genes miRNAs and TFs nodes were screened out. Finally, validation of hub genes was performed by using receiver operating characteristic curve (ROC) analysis. -

B3GAT1 Antibody Cat
B3GAT1 Antibody Cat. No.: 64-189 B3GAT1 Antibody Flow cytometric analysis of K562 cells (right histogram) compared to a negative control cell (left histogram).FITC-conjugated goat-anti- rabbit secondary antibodies were used for the analysis. Specifications HOST SPECIES: Rabbit SPECIES REACTIVITY: Human This B3GAT1 antibody is generated from rabbits immunized with a KLH conjugated IMMUNOGEN: synthetic peptide between 21-48 amino acids from the N-terminal region of human B3GAT1. TESTED APPLICATIONS: Flow, WB For WB starting dilution is: 1:1000 APPLICATIONS: For FACS starting dilution is: 1:10~50 PREDICTED MOLECULAR 38 kDa WEIGHT: September 26, 2021 1 https://www.prosci-inc.com/b3gat1-antibody-64-189.html Properties This antibody is purified through a protein A column, followed by peptide affinity PURIFICATION: purification. CLONALITY: Polyclonal ISOTYPE: Rabbit Ig CONJUGATE: Unconjugated PHYSICAL STATE: Liquid BUFFER: Supplied in PBS with 0.09% (W/V) sodium azide. CONCENTRATION: batch dependent Store at 4˚C for three months and -20˚C, stable for up to one year. As with all antibodies STORAGE CONDITIONS: care should be taken to avoid repeated freeze thaw cycles. Antibodies should not be exposed to prolonged high temperatures. Additional Info OFFICIAL SYMBOL: B3GAT1 Galactosylgalactosylxylosylprotein 3-beta-glucuronosyltransferase 1, Beta-1,3- ALTERNATE NAMES: glucuronyltransferase 1, Glucuronosyltransferase P, GlcAT-P, UDP-GlcUA:glycoprotein beta-1,3-glucuronyltransferase, GlcUAT-P, B3GAT1, GLCATP ACCESSION NO.: Q9P2W7 PROTEIN GI NO.: 205830910 GENE ID: 27087 USER NOTE: Optimal dilutions for each application to be determined by the researcher. Background and References The protein encoded by this gene is a member of the glucuronyltransferase gene family. -

Efficient Generation of Human Natural Killer Cell Lines by Viral Transformation
Letters to the Editor 192 2 Department of Immunology, Erasmus MC, Erasmus University selected children with hypoplastic refractory cytopenia. Haematologica 2007; 92: Medical Center, Rotterdam, The Netherlands; 397–400. 3Department of Pediatrics and Adolescent Medicine, Division of 4 Dunn DE, Tanawattanacharoen P, Boccuni P, Nagakura S, Green SW, Kirby MR Pediatric Hematology and Oncology, University of Freiburg, et al. Paroxysmal nocturnal hemoglobinuria cells in patients with bone marrow Freiburg, Germany; failure syndromes. Ann Intern Med 1999; 131: 401–408. 4Department of Pathology, Clinical Centre South West, Bo¨blingen 5 Maciejewski JP, Follmann D, Nakamura R, Saunthararajah Y, Rivera CE, Simonis T et al. Increased frequency of HLA-DR2 in patients with paroxysmal nocturnal Clinics, Bo¨blingen, Germany; 5 hemoglobinuria and the PNH/aplastic anemia syndrome. Blood 2001; 98: Department of Clinical Genetics, Erasmus MC, Erasmus University 3513–3519. Medical Center, Rotterdam, The Netherlands; 6 Wang H, Chuhjo T, Yasue S, Omine M, Nakao S. Clinical significance of a minor 6 St. Anna Children’s Hospital and Children’s Cancer Research Institute, population of paroxysmal nocturnal hemoglobinuria-type cells in bone marrow Department of Pediatrics, Medical University of Vienna, failure syndrome. Blood 2002; 100: 3897–3902. Vienna, Austria; 7 Yoshida N, Yagasaki H, Takahashi Y, Yamamoto T, Liang J, Wang Y et al. 7Department of Pediatrics, Aarhus University Hospital Skejby, Clinical impact of HLA-DR15, a minor population of paroxysmal Aarhus, Denmark; nocturnal haemoglobinuria-type cells, and an aplastic anaemia-associated auto- 8Department of Pediatric Hematology-Oncology, IRCCS Ospedale antibody in children with acquired aplastic anaemia. Br J Haematol 2008; 142: 427–435. -

Table S1. Detailed Gene Lists Related with Glaucoma, Sclera and Extracellular Matrix (ECM) Remodeling from Text Mining
Table S1. Detailed gene lists related with Glaucoma, Sclera and Extracellular matrix (ECM) remodeling from text mining. Glaucoma related genes Sclera related genes ECM remodeling related genes Common genes GSTM1 NID1 AKT3 NID1 TAP2 EPRS NID1 TGFB2 LGALS8 TGFB2 ACTA1 HSD11B1 NID1 PROX1 PARP1 CD34 TBCE IRF6 MIXL1 REN MIXL1 HSD11B1 TGFB2 DES LBR PLXNA2 PROX1 MPZ EPRS CD34 HSD11B1 ABCB6 TGFB2 CD46 CD34 FN1 IRF6 CR2 IL10 RPE HSD11B1 CD55 REN SST PLXNA2 REN FMOD KNG1 CD34 FMOD CHI3L1 VCAM1 CD46 PRELP CHIT1 F3 IL10 ADORA1 AGXT SOD2 NFASC CFH GPC1 RHO REN PRG4 PTPRC SHH OPTC MYOC COL6A3 NOS3 CHI3L1 SELE PTGS2 GJA1 RNPEP SLC4A3 LAMC2 TIMP2 LMOD1 DES SOAT1 MKI67 PER2 MPZ FASLG APOA1 PTPRC ABCB6 SERPINE2 CALD1 PRLH USF1 SELP C2 CRB1 CD244 POU2F1 VAMP8 CFH VANGL2 DES MMP3 PTGS2 FN1 DDR2 MMP1 PDC NES MPZ GH1 C1orf27 RPE ABCB6 SLC17A5 DHX9 TFPI ADAMTS4 NOVA2 LAMC1 NBPF14 IHH DMPK SOAT1 SST CYP27A1 NOS1 PSMD1 OLFML3 NHLH1 ACE PRDX6 KNG1 CRP PVR TNFSF4 CRYGS IGFBP5 SERPINE1 FASLG ADORA3 NTRK1 ACHE MYOC GCG FN1 ENG COL4A3 VCAM1 NES BGN SELE F3 RPE CD79A SELL MBNL1 CREB1 GALNS F5 PLG MUC1 TGFB1 SELP SOD2 NRP2 VEGFA ATP1B1 RHO SHC1 HSPA5 DPT VIPR2 BMPR2 COL1A1 SLC4A3 B3GAT1 SUMO1 IGF1 DES NLRP5 CASP8 GSN PBX1 SHH S100A1 COL1A2 MPZ LIMS1 S100A4 DCN ABCB6 NOS3 FLG LUM USF1 HGS CTSK TNC CD244 HGD CTSS GFAP VANGL2 GJA1 ECM1 CDH1 PEX19 TIMP2 TFPI CDKN1A ATP1A2 MKI67 CALCRL PLAU PNKD KLK1 ITGAV MFAP2 CRP APOA1 MFI2 CTNNB1 DARC TFG TTN C10orf27 IGFBP2 CALD1 ATF2 ELN NTRK1 C2 NGF PTCH1 FN1 PAEP TP63 MMP2 LMNA TSPAN33 SST CNOT8 RPE CNTN3 ITGA6 EDN1 MAP2 DBH ADIPOQ -

Same Role but Different Actors: Genetic Regulation of Post-Translational Modification of Two Distinct Proteins
bioRxiv preprint doi: https://doi.org/10.1101/2021.05.04.442584; this version posted May 4, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-ND 4.0 International license. Same role but different actors: genetic regulation of post-translational modification of two distinct proteins Arianna Landini*1, Irena Trbojević-Akmačić*2, Pau Navarro3, Yakov A. Tsepilov4,5, Sodbo Z. Sharapov4, Frano Vučković2, Ozren Polašek6,7 Caroline Hayward3, Tea Petrovic2, Marija Vilaj2, Yurii S. Aulchenko4, Gordan Lauc*2,8, James F. Wilson*1,3, Lucija Klarić*3 1 Centre for Global Health Research, Usher Institute, University of Edinburgh, Edinburgh, United Kingdom 2 Genos Glycoscience Research Laboratory, Zagreb, Croatia 3 MRC Human Genetics Unit, Institute for Genetics and Cancer, University of Edinburgh, Edinburgh, United Kingdom 4 Laboratory of Glycogenomics, Institute of Cytology and Genetics, Novosibirsk, Russia 5 Laboratory of Theoretical and Applied Functional Genomics, Novosibirsk State University, Novosibirsk, Russia 6 Department of Public Health, School of Medicine, University of Split, Split, Croatia 7 Gen-info Ltd., Zagreb, Croatia 8 Faculty of Pharmacy and Biochemistry, University of Zagreb, Zagreb, Croatia * Authors contributed equally. Correspondence to: J.F.W ([email protected]) or L.K. ([email protected]) Post-translational modifications (PTMs) diversify protein functions and dynamically coordinate their signalling networks, influencing most aspects of cell physiology. Nevertheless, their genetic regulation or influence on complex traits is not fully understood. -

Clinical Impact of Inherited and Acquired Genetic Variants in Mastocytosis
International Journal of Molecular Sciences Review Clinical Impact of Inherited and Acquired Genetic Variants in Mastocytosis Boguslaw Nedoszytko 1,* , Michel Arock 2,3 , Jonathan J. Lyons 4, Guillaume Bachelot 2, Lawrence B. Schwartz 5, Andreas Reiter 6, Mohamad Jawhar 6 , Juliana Schwaab 6 , Magdalena Lange 1, Georg Greiner 7,8,9 , Gregor Hoermann 8,10 , Marek Niedoszytko 11, Dean D. Metcalfe 4 and Peter Valent 8,12 1 Department of Dermatology, Allergology and Venereology, Medical University of Gdansk, 80-211 Gdansk, Poland; [email protected] 2 Department of Hematology, APHP, Hôpital Pitié-Salpêtrière and Sorbonne University, 75013 Paris, France; [email protected] (M.A.); [email protected] (G.B.) 3 Centre de Recherche des Cordeliers, INSERM, Sorbonne Université, Université de Paris, Cell Death and Drug Resistance in Hematological Disorders Team, 75006 Paris, France 4 Laboratory of Allergic Diseases, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, MD 20892-188, USA; [email protected] (J.J.L.); [email protected] (D.D.M.) 5 Department of Internal Medicine, Division of Rheumatology, Allergy & Immunology, Virginia Commonwealth University, Richmond, VA 23298, USA; [email protected] 6 University Hospital Mannheim, Heidelberg University, 68167 Mannheim, Germany; [email protected] (A.R.); [email protected] (M.J.); [email protected] (J.S.) 7 Department of Laboratory Medicine, Medical -

B3GAT1 Monoclonal Antibody, Clone NK-1
B3GAT1 monoclonal antibody, Gene Alias: CD57, GLCATP, GlcAT-P, GlcUAT-P, clone NK-1 (PE) HNK-1, HNK1, LEU7, NK-1 Catalog Number: MAB6074 Gene Summary: The protein encoded by this gene is a member of the glucuronyltransferase gene family. These Regulatory Status: For research use only (RUO) enzymes exhibit strict acceptor specificity, recognizing nonreducing terminal sugars and their anomeric Product Description: Mouse monoclonal antibody linkages. This gene product functions as the key enzyme raised against native B3GAT1. in a glucuronyl transfer reaction during the biosynthesis of the carbohydrate epitope HNK-1 (human natural Clone Name: NK-1 killer-1, also known as CD57 and LEU7). Alternate transcriptional splice variants have been characterized. Immunogen: Native purified from human peripheral [provided by RefSeq] blood mononuclear cells. References: Host: Mouse 1. CD57+ T lymphocytes are derived from CD57- precursors by differentiation occurring in late immune Reactivity: Human responses. d'Angeac AD, Monier S, Pilling D, Travaglio-Encinoza A, Reme T, Salmon M. Eur J Applications: Flow Cyt, IHC-Fr Immunol. 1994 Jul;24(7):1503-11. (See our web site product page for detailed applications 2. Characterization of HNK-1+ (Leu-7) human information) lymphocytes. I. Two distinct phenotypes of human NK Protocols: See our web site at cells with different cytotoxic capability. Abo T, Cooper http://www.abnova.com/support/protocols.asp or product MD, Balch CM. J Immunol. 1982 Oct;129(4):1752-7. page for detailed protocols 3. A differentiation antigen of human NK and K cells identified by a monoclonal antibody (HNK-1). Abo T, Specificity: Specificity human CD57 Balch CM.