A Study of Protein Dynamics and Cofactor Interactions in Photosystem I
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Theoretical Study of the Tryptophan Synthase Enzyme Reaction Network
A Theoretical Study of the Tryptophan Synthase Enzyme Reaction Network Dissertation zur Erlangung des akademischen Grades doctor rerum naturalium (Dr. rer. nat.) im Fach Chemie Spezialisierung: Physikalische und theoretische Chemie Eingereicht an der Mathematisch-Naturwissenschaftlichen Fakult¨at der Humboldt-Universit¨at zu Berlin von Dimitri Loutchko Pr¨asidentin der Humboldt-Universit¨atzu Berlin Prof. Dr.-Ing. Dr. Sabine Kunst Dekan der Mathematisch-Naturwissenschaftlichen Fakult¨at Prof. Dr. Elmar Kulke 1. Gutachter: Prof. Dr. Gerhard Ertl 2. Gutachter: Prof. Dr. Klaus Rademann 3. Gutachter: Prof. Dr. Yannick de Decker Tag der m¨undlichen Pr¨ufung:09.07.2018 ii Abstract iii Abstract The channeling enzyme tryptophan synthase provides a paradigmatic example of a chemical nanomachine. It catalyzes the biosynthesis of tryptophan from serine and indole glycerol phos- phate. As a single macromolecule, it possesses two distinct catalytic subunits and implements 13 different elementary reaction steps. A complex pattern of allosteric regulation is involved in its operation. The catalytic activity in a subunit is enhanced or inhibited depending on the state of the other subunit. The gates controlling arrival and release of the ligands can become open or closed depending on the chemical states. The intermediate product indole is directly channeled within the protein from one subunit to another, so that it is never released into the solution around it. In this thesis, the first single-molecule kinetic model of the enzyme is proposed and analyzed. All its transition rate constants are extracted from available experimental data, and thus, no fitting parameters are employed. Numerical simulations reveal strong correlations in the states of the active centers and the emergent synchronization of intramolecular processes in tryptophan synthase. -

Directed Evolution of the Tryptophan Synthase Β-Subunit for Stand-Alone Function Recapitulates Allosteric Activation
Directed evolution of the tryptophan synthase β-subunit for stand-alone function recapitulates allosteric activation Andrew R. Buller1, Sabine Brinkmann-Chen1, David K. Romney, Michael Herger, Javier Murciano-Calles, and Frances H. Arnold2 Division of Chemistry and Chemical Engineering, California Institute of Technology, Pasadena, CA 91125 Edited by Alan R. Fersht, Medical Research Council Laboratory of Molecular Biology, Cambridge, United Kingdom, and approved October 16, 2015 (received for review August 17, 2015) Enzymes in heteromeric, allosterically regulated complexes cata- associated with open, partially closed, and fully closed states lyze a rich array of chemical reactions. Separating the subunits of during the catalytic cycle (2, 4, 6). such complexes, however, often severely attenuates their catalytic TrpS is a naturally promiscuous enzyme complex: the model activities, because they can no longer be activated by their protein system from S. typhimurium catalyzes its β-substitution reaction partners. We used directed evolution to explore allosteric regula- with most haloindoles, methylindoles, and aminoindoles, along tion as a source of latent catalytic potential using the β-subunit of with an assortment of nonindole nucleophiles for C–S, C–N, and – tryptophan synthase from Pyrococcus furiosus (PfTrpB). As part of C C bond formation (7). Such noncanonical amino acids (NCAAs) its native αββα complex, TrpB efficiently produces tryptophan and have diverse applications in chemical biology (8), serve as inter- tryptophan analogs; activity drops considerably when it is used as mediates in the synthesis of natural products (9, 10), and are priv- a stand-alone catalyst without the α-subunit. Kinetic, spectro- ileged scaffolds for the development of pharmaceuticals (11). -

(12) United States Patent (10) Patent No.: US 7,094,568 B2 Kozlowski Et Al
US007094568B2 (12) United States Patent (10) Patent No.: US 7,094,568 B2 Kozlowski et al. (45) Date of Patent: Aug. 22, 2006 (54) METHOD FOR PRODUCING PROTEINS WO WO O1, 57198 A3 2, 2002 TAGGED AT THE N- OR C-TERMINUS (75) Inventors: Roland Kozlowski, Babraham (GB); OTHER PUBLICATIONS Michael B. McAndrew, Babraham (GB); Jonathan Michael Blackburn, Bordini, E., and Hamdan, M., “Investigation of Some Covalent and Cambridge (GB); Michelle Anne Noncovalent Complexes by Matrix-assisted Laser Desorption/Ion Mulder, Capetown (ZA); Mitali ization Time-of-flight and Electrospray Mass Spectrometry.” Rapid Commun. Mass Spectrom. 13:1143-1151, John Wiley & Sons, Ltd. Samaddar, Hyderabad (IN) (1999). Cai, J., et al., “Functional Expression of Multidrug Resistance (73) Assignee: Sense Proteomic Ltd., (GB) Protein 1 in Pichia pastoris.” Biochemistry 40:8307-8316, Ameri can Chemical Society (Jun. 2001). (*) Notice: Subject to any disclaimer, the term of this DeRisi, J., et al., “Use of a cDNA microarray to analyse gene patent is extended or adjusted under 35 expression patterns in human cancer,” Nat. Genet. 14:457-460, U.S.C. 154(b) by 477 days. Nature Publishing Co. (1996). Doellgast, G.J., et al., “Sensitive Enzyme-Linked Immunosorbent (21) Appl. No.: 10/114,334 Assay for Detection of Clostridium botulinum Neurotoxins A, B, and E Using Signal Amplification via Enzyme-Linked Coagulation (22) Filed: Apr. 3, 2002 Assay.” J. Clin. Microbiol. 3 1:2402-2409, American Society for Microbiology (1993). (65) Prior Publication Data Giuliani, C.D., et al., “Expression of an active recombinant lysine US 2003 FOOT3811 A1 Apr. 17, 2003 49 phospholipase A myotoxin as a fusion protein in bacteria.” Toxicon 39:1595-1600, Elsevier Science Ltd. -

Combined Transcriptome and Metabolome Analyses to Understand the Dynamic Responses of Rice Plants to Attack by the Rice Stem
Liu et al. BMC Plant Biology (2016) 16:259 DOI 10.1186/s12870-016-0946-6 RESEARCHARTICLE Open Access Combined transcriptome and metabolome analyses to understand the dynamic responses of rice plants to attack by the rice stem borer Chilo suppressalis (Lepidoptera: Crambidae) Qingsong Liu1†, Xingyun Wang1†, Vered Tzin2, Jörg Romeis1,3, Yufa Peng1 and Yunhe Li1* Abstract Background: Rice (Oryza sativa L.), which is a staple food for more than half of the world’s population, is frequently attacked by herbivorous insects, including the rice stem borer, Chilo suppressalis. C. suppressalis substantially reduces rice yields in temperate regions of Asia, but little is known about how rice plants defend themselves against this herbivore at molecular and biochemical level. Results: In the current study, we combined next-generation RNA sequencing and metabolomics techniques to investigate the changes in gene expression and in metabolic processes in rice plants that had been continuously fed by C. suppressalis larvae for different durations (0, 24, 48, 72, and 96 h). Furthermore, the data were validated using quantitative real-time PCR. There were 4,729 genes and 151 metabolites differently regulated when rice plants were damaged by C. suppressalis larvae. Further analyses showed that defense-related phytohormones, transcript factors, shikimate-mediated and terpenoid-related secondary metabolism were activated, whereas the growth-related counterparts were suppressed by C. suppressalis feeding. The activated defense was fueled by catabolism of energy storage compounds such as monosaccharides, which meanwhile resulted in the increased levels of metabolites that were involved in rice plant defense response. Comparable analyses showed a correspondence between transcript patterns and metabolite profiles. -

Structure and Conformational Dynamics of Fatty Acid Synthases
Structure and Conformational Dynamics of Fatty Acid Synthases Inauguraldissertation zur Erlangung der Würde eines Doktors der Philosophie vorgelegt der Philosophisch-Naturwissenschaftlichen Fakultät der Universität Basel von Friederike Maria Carola Benning aus Deutschland Basel, 2017 Originaldokument gespeichert auf dem Dokumentenserver der Universität Basel edoc.unibas.ch Genehmigt von der Philosophisch-Naturwissenschaftlichen Fakultät auf Antrag von Prof. Dr. Timm Maier Prof. Dr. Sebastian Hiller Basel, den 20.06.2017 Prof. Dr. Martin Spiess Dekan 2 3 4 I Abstract Multistep reactions rely on substrate channeling between active sites. Carrier protein- based enzyme systems constitute the most versatile class of shuttling systems due to their capability of linking multiple catalytic centers. In eukaryotes and some bacteria, these systems have evolved to multifunctional enzymes, which integrate all functional domains involved into one or more giant polypeptide chains. The metazoan fatty acid synthase (FAS) is a key paradigm for carrier protein-based multienzymes. It catalyzes the de novo biosynthesis of fatty acids from carbohydrate-derived precursors in more than 40 individual reactions steps. Its seven functional domains are encoded on one polypeptide chain, which assembles into an X-shaped dimer for activity. The dimer features two lateral reaction clefts, each equipped with a full set of active sites and a flexibly tethered carrier protein. Substrate loading and condensation in the condensing region are structurally and functionally separated from the b-carbon processing domains in the modifying region. At the beginning of this thesis, only a single crystal structure of an intact metazoan FAS was known. FAS, in particular its modifying region, displays extensive conformational variability, according to electron microscopy (EM) studies. -

Product Sheet Info
Master Clone List for NR-19279 ® Vibrio cholerae Gateway Clone Set, Recombinant in Escherichia coli, Plates 1-46 Catalog No. NR-19279 Table 1: Vibrio cholerae Gateway® Clones, Plate 1 (NR-19679) Clone ID Well ORF Locus ID Symbol Product Accession Position Length Number 174071 A02 367 VC2271 ribD riboflavin-specific deaminase NP_231902.1 174346 A03 336 VC1877 lpxK tetraacyldisaccharide 4`-kinase NP_231511.1 174354 A04 342 VC0953 holA DNA polymerase III, delta subunit NP_230600.1 174115 A05 388 VC2085 sucC succinyl-CoA synthase, beta subunit NP_231717.1 174310 A06 506 VC2400 murC UDP-N-acetylmuramate--alanine ligase NP_232030.1 174523 A07 132 VC0644 rbfA ribosome-binding factor A NP_230293.2 174632 A08 322 VC0681 ribF riboflavin kinase-FMN adenylyltransferase NP_230330.1 174930 A09 433 VC0720 phoR histidine protein kinase PhoR NP_230369.1 174953 A10 206 VC1178 conserved hypothetical protein NP_230823.1 174976 A11 213 VC2358 hypothetical protein NP_231988.1 174898 A12 369 VC0154 trmA tRNA (uracil-5-)-methyltransferase NP_229811.1 174059 B01 73 VC2098 hypothetical protein NP_231730.1 174075 B02 82 VC0561 rpsP ribosomal protein S16 NP_230212.1 174087 B03 378 VC1843 cydB-1 cytochrome d ubiquinol oxidase, subunit II NP_231477.1 174099 B04 383 VC1798 eha eha protein NP_231433.1 174294 B05 494 VC0763 GTP-binding protein NP_230412.1 174311 B06 314 VC2183 prsA ribose-phosphate pyrophosphokinase NP_231814.1 174603 B07 108 VC0675 thyA thymidylate synthase NP_230324.1 174474 B08 466 VC1297 asnS asparaginyl-tRNA synthetase NP_230942.2 174933 B09 198 -

Development of Computational Resources for Predicting Disease Resistance Genes and Mapping Ngs- Transcripts to Secondary Metabolism in Plants
Ph.D DEVELOPMENT OF COMPUTATIONAL . RESOURCES FOR PREDICTING DISEASE RESISTANCE GENES AND MAPPING NGS- TRANSCRIPTS TO SECONDARY METABOLISM IN PLANTS TARUN PAL PAL TARUN Thesis submitted in fulfillment of the requirements for the Degree of DOCTOR OF PHILOSOPHY IN BIOINFORMATICS BY TARUN PAL JUIT,Waknaghat Department of Biotechnology & Bioinformatics 2017 JAYPEE UNIVERSITY OF INFORMATION TECHNOLOGY WAKNAGHAT, DISTRICT SOLAN, H.P., INDIA Month November Year 2017 DEVELOPMENT OF COMPUTATIONAL RESOURCES FOR PREDICTING DISEASE RESISTANCE GENES AND MAPPING NGS- TRANSCRIPTS TO SECONDARY METABOLISM IN PLANTS Thesis submitted in fulfillment of the requirements for the Degree of DOCTOR OF PHILOSOPHY IN BIOINFORMATICS BY TARUN PAL Department of Biotechnology & Bioinformatics JAYPEE UNIVERSITY OF INFORMATION TECHNOLOGY WAKNAGHAT, DISTRICT SOLAN, H.P., INDIA Month November Year 2017 Copyright @ JAYPEE UNIVERSITY OF INFORMATION TECHNOLOGY WAKNAGHAT DECEMBER 2017 ALL RIGHTS RESERVED DECLARATION BY THE SCHOLAR I hereby declare that the work reported in the Ph.D. thesis entitled “Development of computational resources for predicting disease resistance genes and mapping NGS-transcripts to secondary metabolism in plants” submitted at Jaypee University of Information Technology, Waknaghat, India, is an authentic record of my work carried out under the supervision of Prof. (Dr.) Rajinder Singh Chauhan. I have not submitted this work elsewhere for any other degree or diploma. I am fully responsible for the contents of my Ph.D. Theses. Tarun Pal Department of Biotechnology & Bioinformatics Jaypee University of Information Technology, Waknaghat, India Date: SUPERVISOR’S CERTIFICATE This is to certify that the work reported in the Ph.D. thesis entitled “Development of computational resources for predicting disease resistance genes and mapping NGS-transcripts to secondary metabolism in plants”, submitted by Tarun Pal at Jaypee University of Information Technology, Waknaghat, India, is a bonafide record of his / her original work carried out under my supervision. -

A Genomic Survey of Two Dinotoms
A GENOMIC SURVEY OF TWO DINOTOMS by Behzad Imanian MSc., The University of British Columbia, 2006 BSc., The University of British Columbia, 2002 A THESIS SUBMITTED IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE OF DOCTOR OF PHILOSOPHY in THE FACULTY OF GRADUATE STUDIES (Botany) THE UNIVERSITY OF BRITISH COLUMBIA (Vancouver) April 2013 © Behzad Imanian, 2013 Abstract Endosymbiosis has played a major role in shaping eukaryotic cells, their success and diversity. At the base of the eukaryotic tree, an α-proteobacterium endosymbiont in a protoeukaryotic cell was converted into the mitochondrion through its reductive evolution, endosymbiotic gene transfer (EGT) and the development of a protein targeting system to direct the products of the transferred genes to this organelle. Similar events mark the plastid evolution from a cyanobacterium. However, the primary endosymbiosis of plastid, unlike the mitochondrion, was followed by the secondary and tertiary movement of this organelle between eukaryotes through analogous endosymbiotic reduction, EGT and evolution of a protein targeting system and many subsequent independent losses from different eukaryotic lineages. The obligate tertiary diatom endosymbiont in a small group of dinoflagellates called ‘dinotoms’ is exceptional in that it retains most of its ancestral characters including a large nucleus, its own mitochondria, plastids and many other eukaryotic organelles and structures in a large cytoplasm all enclosed in and separated from its dinoflagellate host by a single membrane. This level of conservation of ancestral features in the endosymbiont suggests an early stage of integration. In order to investigate the impacts of endosymbiosis on the organelle genomes and to determine the extent of EGT and the contribution of the host nuclear genome to the proteomes of the organelles, I conducted mass pyrosequencing of the A+T-rich portion of the DNA extracted from two dinotoms, Durinskia baltica and Kryptoperidinium foliaceum, and the SL cDNA library constructed for D. -

When an Enzyme Isn't Just an Enzyme Anymore Extra Botany
eXtra Botany Insight When an enzyme isn’t just an enzyme anymore Brenda S.J. Winkel Department of Biological Sciences, Virginia Tech, Blacksburg, VA 24060, USA [email protected] In this issue (pages 1425–1440) Bross et al. provide evi- There are numerous other examples of glycolytic enzymes dence of surprising alternative functions for two iso- with alternative non-catalytic functions, though none as inten- forms of the plastid enzyme arogenate dehydratase. sively studied as GAPDH (Jeffery, 2014). The functions of This study points to a previously unsuspected con- these proteins range from serving as transcriptional regulators nection between central metabolism and chloroplast that are sensitive to sucrose concentrations to binding of plas- division and a potential new mechanism for retrograde minogen during microbial pathogenesis (Rolland et al., 2006; signaling. These findings add to a growing awareness He et al., 2013; Henderson and Martin, 2013; Vega et al., of the complexities of protein function that has sub- 2016). Dual functionality is also associated with enzymes of stantial implications for both basic and applied plant many other pathways of central metabolism, such as homoci- science. trate synthase, which is essential for lysine biosynthesis in mitochondria but can also mediate DNA repair in the nucleus We are in the midst of a paradigm shift in our understanding (Torres-Machorro et al., 2015). Aconitase is another well- of cellular metabolism, as growing numbers of proteins with established example, functioning not only as a key enzyme well-established catalytic roles are being found to have addi- in the TCA cycle, but also as an iron-responsive protein that tional, alternative functions. -
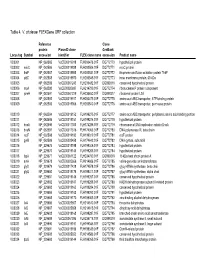
Table 4. V. Cholerae Flexgene ORF Collection
Table 4. V. cholerae FLEXGene ORF collection Reference Clone protein PlasmID clone GenBank Locus tag Symbol accession identifier FLEX clone name accession Product name VC0001 NP_062585 VcCD00019918 FLH200476.01F DQ772770 hypothetical protein VC0002 mioC NP_062586 VcCD00019938 FLH200506.01F DQ772771 mioC protein VC0003 thdF NP_062587 VcCD00019958 FLH200531.01F DQ772772 thiophene and furan oxidation protein ThdF VC0004 yidC NP_062588 VcCD00019970 FLH200545.01F DQ772773 inner membrane protein, 60 kDa VC0005 NP_062589 VcCD00061243 FLH236482.01F DQ899316 conserved hypothetical protein VC0006 rnpA NP_062590 VcCD00025697 FLH214799.01F DQ772774 ribonuclease P protein component VC0007 rpmH NP_062591 VcCD00061229 FLH236450.01F DQ899317 ribosomal protein L34 VC0008 NP_062592 VcCD00019917 FLH200475.01F DQ772775 amino acid ABC transporter, ATP-binding protein VC0009 NP_062593 VcCD00019966 FLH200540.01F DQ772776 amino acid ABC transproter, permease protein VC0010 NP_062594 VcCD00019152 FLH199275.01F DQ772777 amino acid ABC transporter, periplasmic amino acid-binding portion VC0011 NP_062595 VcCD00019151 FLH199274.01F DQ772778 hypothetical protein VC0012 dnaA NP_062596 VcCD00017363 FLH174286.01F DQ772779 chromosomal DNA replication initiator DnaA VC0013 dnaN NP_062597 VcCD00017316 FLH174063.01F DQ772780 DNA polymerase III, beta chain VC0014 recF NP_062598 VcCD00019182 FLH199319.01F DQ772781 recF protein VC0015 gyrB NP_062599 VcCD00025458 FLH174642.01F DQ772782 DNA gyrase, subunit B VC0016 NP_229675 VcCD00019198 FLH199346.01F DQ772783 hypothetical protein -

Chandran Et Al. Supporting Info.Pdf
Supporting Information (SI) Appendix Part 1. Impact of tissue preparation, LMD, and RNA amplification on array output. p. 2 Text S1: Detailed Experimental Design and Methods Figure S1A: Correlation analysis indicates tissue preparation has minimal impact on ATH1 array output. Figure S1B: Correlation analysis indicates RNA degradation does not significantly impact array output. Figure S1C: Correlation analysis indicates two-round amplification does not significantly impact array output. Figure S1D: Independent biological replicates of LMD samples are highly correlated. Figure S1E: Validation of LMD array expression by qPCR. Table S1: Tissue preparation-associated genes Part 2. Analysis of LMD and parallel whole leaf array data. p. 13 Table S2A: Known PM-impacted genes enriched in LMD dataset Table S2B: Dataset of LMD and whole leaf genes with PM-altered expression Table S2C: LMD PM MapMan Results and Bins Table S2D: ics1 vs. WT LMD PM MapMan Results and Bins Table S2E: Infection site-specific changes for redox and calcium categories Table S2F: cis-acting regulatory element motif analysis Part 3. Process network construction p. 149 3A. Photosynthesis 3B. Cold/dehydration response Part 4. Powdery mildew infection of WT and myb3r4 mutants p. 173 Text S4: Detailed Experimental Design and Methods (supplement to manuscript) Figure S4A. Uninfected 4 week old WT and myb3r4 plants Figure S4B. myb3r4 mutants exhibit reduced visible PM growth and reproduction Figure S4C. PM-infected WT and myb3r4 mutants do not exhibit cell death Figure S4D. Endoreduplication occurs at site of PM infection not distal to infection Figure S4E. Ploidy correlates with nuclear size. Part 1. Impact of tissue preparation, LMD, and RNA amplification on array output. -

Plastid-Localized Amino Acid Biosynthetic Pathways of Plantae Are Predominantly Composed of Non-Cyanobacterial Enzymes
Plastid-localized amino acid biosynthetic pathways of Plantae are predominantly SUBJECT AREAS: MOLECULAR EVOLUTION composed of non-cyanobacterial PHYLOGENETICS PLANT EVOLUTION enzymes PHYLOGENY Adrian Reyes-Prieto1* & Ahmed Moustafa2* Received 1 26 September 2012 Canadian Institute for Advanced Research and Department of Biology, University of New Brunswick, Fredericton, Canada, 2Department of Biology and Biotechnology Graduate Program, American University in Cairo, Egypt. Accepted 27 November 2012 Studies of photosynthetic eukaryotes have revealed that the evolution of plastids from cyanobacteria Published involved the recruitment of non-cyanobacterial proteins. Our phylogenetic survey of .100 Arabidopsis 11 December 2012 nuclear-encoded plastid enzymes involved in amino acid biosynthesis identified only 21 unambiguous cyanobacterial-derived proteins. Some of the several non-cyanobacterial plastid enzymes have a shared phylogenetic origin in the three Plantae lineages. We hypothesize that during the evolution of plastids some enzymes encoded in the host nuclear genome were mistargeted into the plastid. Then, the activity of those Correspondence and foreign enzymes was sustained by both the plastid metabolites and interactions with the native requests for materials cyanobacterial enzymes. Some of the novel enzymatic activities were favored by selective compartmentation should be addressed to of additional complementary enzymes. The mosaic phylogenetic composition of the plastid amino acid A.R.-P. ([email protected]) biosynthetic pathways and the reduced number of plastid-encoded proteins of non-cyanobacterial origin suggest that enzyme recruitment underlies the recompartmentation of metabolic routes during the evolution of plastids. * Equal contribution made by these authors. rimary plastids of plants and algae are the evolutionary outcome of an endosymbiotic association between eukaryotes and cyanobacteria1.