Binding of Androgen- and Estrogen-Like Flavonoids to Their Cognate (Non)Nuclear Receptors: a Comparison by Computational Prediction
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Dehydroepiandrosterone – Is the Fountain of Youth Drying Out?
Physiol. Res. 52: 397-407, 2003 MINIREVIEW Dehydroepiandrosterone – Is the Fountain of Youth Drying Out? P. CELEC 1,2, L. STÁRKA3 1Faculty of Medicine, 2Faculty of Natural Sciences, Comenius University, Bratislava, Slovakia and 3Institute of Endocrinology, Prague, Czech Republic Received September 15, 2002 Accepted October 7, 2002 Summary Dehydroepiandrosterone (DHEA) and its sulphate-bound form (DHEAS) are important steroids mainly of adrenal origin. Their physiological and pathophysiological functions are not yet fully identified, although a number of various possible features have been hypothesized. Most popular is the description of the “hormone of youth” as the long-term dynamics of DHEA levels are characterized by a sharp age-related decline in the late adulthood and later. Low levels of DHEA are, however, associated not only with the ageing process but also with diabetes mellitus, cardiovascular diseases and some neurological or immunological entities. In the past decade, a number of brief studies have concentrated on these relationships and also on the role of exogenous DHEA in health, disease and human well-being. This article tries to summarize some of the most important facts achieved recently. Key words Dehydroepiandrosterone • Intracrinology • Hormone replacement therapy • Steroids Introduction functions: 1) DHEA is an endogenous metabolite that cannot be patented so that pharmaceutical companies are In 1934 Butenandt and Dannenbaum isolated not interested in supporting research in this field. dehydroepiandrosterone (DHEA) from urine and in 1944 2) DHEA can be described as a “human molecule” Munson and colleagues identified its 3β-sulphate because other investigated species have much lower (DHEAS). Even now, nearly 70 years later, we still do concentrations. -

The Orphan Receptor GPR17 Is Unresponsive to Uracil Nucleotides and Cysteinyl Leukotrienes S
Supplemental material to this article can be found at: http://molpharm.aspetjournals.org/content/suppl/2017/03/02/mol.116.107904.DC1 1521-0111/91/5/518–532$25.00 https://doi.org/10.1124/mol.116.107904 MOLECULAR PHARMACOLOGY Mol Pharmacol 91:518–532, May 2017 Copyright ª 2017 by The American Society for Pharmacology and Experimental Therapeutics The Orphan Receptor GPR17 Is Unresponsive to Uracil Nucleotides and Cysteinyl Leukotrienes s Katharina Simon, Nicole Merten, Ralf Schröder, Stephanie Hennen, Philip Preis, Nina-Katharina Schmitt, Lucas Peters, Ramona Schrage,1 Celine Vermeiren, Michel Gillard, Klaus Mohr, Jesus Gomeza, and Evi Kostenis Molecular, Cellular and Pharmacobiology Section, Institute of Pharmaceutical Biology (K.S., N.M., Ral.S., S.H., P.P., N.-K.S, L.P., J.G., E.K.), Research Training Group 1873 (K.S., E.K.), Pharmacology and Toxicology Section, Institute of Pharmacy (Ram.S., K.M.), University of Bonn, Bonn, Germany; UCB Pharma, CNS Research, Braine l’Alleud, Belgium (C.V., M.G.). Downloaded from Received December 16, 2016; accepted March 1, 2017 ABSTRACT Pairing orphan G protein–coupled receptors (GPCRs) with their using eight distinct functional assay platforms based on label- cognate endogenous ligands is expected to have a major im- free pathway-unbiased biosensor technologies, as well as molpharm.aspetjournals.org pact on our understanding of GPCR biology. It follows that the canonical second-messenger or biochemical assays. Appraisal reproducibility of orphan receptor ligand pairs should be of of GPR17 activity can be accomplished with neither the coapplica- fundamental importance to guide meaningful investigations into tion of both ligand classes nor the exogenous transfection of partner the pharmacology and function of individual receptors. -

Edinburgh Research Explorer
Edinburgh Research Explorer International Union of Basic and Clinical Pharmacology. LXXXVIII. G protein-coupled receptor list Citation for published version: Davenport, AP, Alexander, SPH, Sharman, JL, Pawson, AJ, Benson, HE, Monaghan, AE, Liew, WC, Mpamhanga, CP, Bonner, TI, Neubig, RR, Pin, JP, Spedding, M & Harmar, AJ 2013, 'International Union of Basic and Clinical Pharmacology. LXXXVIII. G protein-coupled receptor list: recommendations for new pairings with cognate ligands', Pharmacological reviews, vol. 65, no. 3, pp. 967-86. https://doi.org/10.1124/pr.112.007179 Digital Object Identifier (DOI): 10.1124/pr.112.007179 Link: Link to publication record in Edinburgh Research Explorer Document Version: Publisher's PDF, also known as Version of record Published In: Pharmacological reviews Publisher Rights Statement: U.S. Government work not protected by U.S. copyright General rights Copyright for the publications made accessible via the Edinburgh Research Explorer is retained by the author(s) and / or other copyright owners and it is a condition of accessing these publications that users recognise and abide by the legal requirements associated with these rights. Take down policy The University of Edinburgh has made every reasonable effort to ensure that Edinburgh Research Explorer content complies with UK legislation. If you believe that the public display of this file breaches copyright please contact [email protected] providing details, and we will remove access to the work immediately and investigate your claim. Download date: 02. Oct. 2021 1521-0081/65/3/967–986$25.00 http://dx.doi.org/10.1124/pr.112.007179 PHARMACOLOGICAL REVIEWS Pharmacol Rev 65:967–986, July 2013 U.S. -

Targeting Lysophosphatidic Acid in Cancer: the Issues in Moving from Bench to Bedside
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by IUPUIScholarWorks cancers Review Targeting Lysophosphatidic Acid in Cancer: The Issues in Moving from Bench to Bedside Yan Xu Department of Obstetrics and Gynecology, Indiana University School of Medicine, 950 W. Walnut Street R2-E380, Indianapolis, IN 46202, USA; [email protected]; Tel.: +1-317-274-3972 Received: 28 August 2019; Accepted: 8 October 2019; Published: 10 October 2019 Abstract: Since the clear demonstration of lysophosphatidic acid (LPA)’s pathological roles in cancer in the mid-1990s, more than 1000 papers relating LPA to various types of cancer were published. Through these studies, LPA was established as a target for cancer. Although LPA-related inhibitors entered clinical trials for fibrosis, the concept of targeting LPA is yet to be moved to clinical cancer treatment. The major challenges that we are facing in moving LPA application from bench to bedside include the intrinsic and complicated metabolic, functional, and signaling properties of LPA, as well as technical issues, which are discussed in this review. Potential strategies and perspectives to improve the translational progress are suggested. Despite these challenges, we are optimistic that LPA blockage, particularly in combination with other agents, is on the horizon to be incorporated into clinical applications. Keywords: Autotaxin (ATX); ovarian cancer (OC); cancer stem cell (CSC); electrospray ionization tandem mass spectrometry (ESI-MS/MS); G-protein coupled receptor (GPCR); lipid phosphate phosphatase enzymes (LPPs); lysophosphatidic acid (LPA); phospholipase A2 enzymes (PLA2s); nuclear receptor peroxisome proliferator-activated receptor (PPAR); sphingosine-1 phosphate (S1P) 1. -

Estroquench™ Hormone Specific Formulation™
PRODUCT DATA DOUGLAS LABORATORIES® 08/2014 1 EstroQuench™ Hormone Specific Formulation™ DESCRIPTION EstroQuench™ is a Hormone Specific Formulation™ of ingredients that have documented anti-aromatase activity as well as androgenic adaptogens which support the function of endogenous aromatase inhibitors. Collectively these herbs promote minimal production and function of estrogens, while promoting testosterone function, including optimal sexual function in both genders. This formulation is designed to quench excessive production of estrogens and aberrant functions of them while supporting optimal function of androgens by maintaining the health of androgen producing glands.† Hormone Specific Formulation™ provided by Douglas Laboratories® and formulated by Dr. Joseph J Collins is created to support the optimal function of specific hormones through the use of hormone specific adaptogens, hormone specific agonists and hormone specific functional mimetics. This formulation may be used as part of a hormone health program with dietary and nutrient support. In addition, this formulation may be used by clinicians as an adjuvant to support optimal hormone health in patients who have been prescribed bioidentical hormone therapies. FUNCTIONS Aromatase (a cytochrome P450 enzyme {CYP19}) is the enzyme that controls the conversion of androgens to estrogens. More specifically, aromatase is the enzyme responsible for catalyzing the biosynthesis of androstenedione into estrone, and the biosynthesis of testosterone to estradiol. Estrogens include the broad range of aromatized hormones created form androgens. The specific attribute of estrogens that separate them from progestogens, androgens and corticoids is that estrogens are the only aromatized steroid hormones. Estradiol, the most potent endogenous estrogen, is biosynthesized by aromatization from androgens by aromatase (which is also called estrogen synthase). -

Test Report Comprehensive Hormone Insights™
698814 COMPREHENSIVE HORMONE INSIGHTS™ TEST REPORT Dr. Maximus, N.D. E: [email protected] Date of Collection: P: 403-241-4500 Time of Collection: F: 403-241-4501 Date of Receipt: www.rmalab.com Reported On: CHI Accession: 698814 Healthcare Professional Patient Age: Dr. Maximus, N.D. Date of Birth: Gender: Male F: Relevant Medications Biometrics Curcumin Height (in) : 73 Weight (lb) : 180 BMI : 24 Waist (in) : 35 Hip (in) : 41 CHI Accession: 698814 SUMMARY HMUS01 How to read the graphs LEGEND: 50 66 Sex Steroid Hormones 50 66 Middle third of 33 33 84 reference population Hormone Start of 83 100 80 100 highest 16 Percentile Precursors 16 Percentile third of Sum of Androgens Sum of Estrogens 50 66 reference 0 0 population (T, DHT, α+β androstanediol) Listed in Interp Guide 33 84 End of 100 16 lowest Percentile00 50 66 50 66 third of 33 33 84 reference 0 population Patient’s percentile rank 81 100 95 100 compared to reference 16 Percentile 16 Percentile population (see summary) DHEA + Metabolites Sum of Progesterone Metabolites 0 (DHEA + A + E) 0 α+β Pregnanediol Cortisol Melatonin Oxidative Stress Free Cortisol Profile (ng/mg) 100 50 66 50 66 33 84 33 84 80 64 100 0 100 16 Percentile 16 Percentile 60 6-sulfatoxy 8-Hydroxy-2- 0 Melatonin 0 deoxyguanosine 40 (Overnight) (Overnight) 20 6-sulfatoxymelatonin provides 8-hydroxy-2-deoxyguanosine is Cortisol/Creatinine (ng/mg) insight into melatonin levels. a marker of oxidative stress 0 Morning Dinner Bedtime A B C 50 66 Free cortisol Cortisol Metabolites 33 84 profile is used to provides a general Testosterone Cortisol assess diurnal assessment of 16 100 cortisol rhythm adrenal cortisol 16 Percentile Cortisol production Cortisol Metabolites 0 (α+β THF + THE) Testosterone Cortisol/Testosterone provides insight into relative catabolic (cortisol) and anabolic (testosterone) states. -
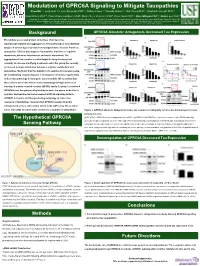
Background the Hypothetical GPRC6A Sensing Pathway
Modulation of GPRC6A Signaling to Mitigate Tauopathies Chao Ma1,2, Jerry Hunt1,3, Leslie Sandusky PhD1,3, William Fraser1,3, Ronald Alvarez1,3, Dali Zheng PhD1,4, Siddharth Kamath PhD1,2, Chad Dickey PhD1,4, Hans Bräuner-Osborne PhD5, Daniel Sejer Pedersen PhD5, Kevin Nash PhD1,2, Dave Morgan PhD1,2, Daniel Lee PhD1,3 1:Byrd Alzheimer‘s Institute, University of South Florida, Tampa, FL 33613, USA. 2:Morsani College of Medicine, Department of Molecular Pharmacology and Physiology, University of South Florida, Tampa, FL 33612, USA. 3:College of Pharmacy, Department of Pharmaceutical Sciences, University of South Florida, Tampa, FL 33612, USA. 4:Morsani College of Medicine, Department of Molecular Medicine, University of South Florida, Tampa, FL 33612, USA. 5:Department of Drug Design and Pharmacology, Faculty of Health and Medical Sciences, University of Copenhagen, Jagtvej 162, 2100 Copenhagen, Denmark Background GPRC6A Allosteric Antagonists Decreased Tau Expression Microtubule associated protein named tau, often becomes hyperphosphorylated and aggregates to form pathological neurofibrillary tangles in several age-associated neurodegenerative diseases known as tauopathies. Clinical phenotypes of tauopathies manifest as cognitive impairment, behavior disturbances and motor impairment. The aggregation of tau remains a central target for drug discovery, but currently no disease-modifying treatments exist. Our group has recently uncovered a unique interaction between L-arginine metabolism and tauopathies. We found that the depletion of L-arginine by overexpressing the metabolizing enzyme arginase 1 and arginine deiminase significantly reduced tau pathology in transgenic mouse models. We speculate that these effects associate with increased autophagy through amino acid sensing. G protein-coupled receptor (GPCR), family C, group 6, member A (GPRC6A) was deorphanized by binding to basic L-α amino acids, like L- arginine. -

2018 Apigenin Male Infertility (2).Pdf
Original Article Thai Journal of Pharmaceutical Sciences Apigenin and baicalin, each alone or in low-dose combination, attenuated chloroquine induced male infertility in adult rats Amira Akilah, Mohamed Balaha*, Mohamed-Nabeih Abd-El Rahman, Sabiha Hedya Department of Pharmacology, Faculty of Medicine, Tanta University, Postal No. 31527, El-Gish Street, Tanta, Egypt Corresponding Author: Department of Pharmacology, ABSTRACT Faculty of Medicine, Tanta University, Postal No. 31527, Introduction: Male infertility is a worldwide health problem, which accounts for about 50% of all El-Gish Street, Tanta, Egypt. cases of infertility and considered as the most common single defined cause of infertility. Recently, Tel.: +201284451952. E-mail: apigenin and baicalin exhibited a powerful antioxidant and antiapoptotic activities. Consequently, in [email protected]. the present study, we evaluated the possible protective effect of apigenin and baicalin, either alone or Edu.Eg (M. Balaha). in low-dose combination, on a rat model of male infertility, regarding for its effects on the hormonal assay, testicular weight, sperm parameters, oxidative-stress state, apoptosis, and histopathological Received: Mar 06, 2018 changes. Material and methods: 12-week-old adult male Wister rats received 10 mg/kg/d Accepted: May 08, 2018 chloroquine orally for 30 days to induce male infertility. Either apigenin (30 or 15 mg/kg/d), baicalin Published: July 10, 2018 (100 or 50 mg/kg/d) or a combination of 15 mg/kg/d apigenin and 50 mg/kg/d baicalin received daily -

Regulation of Immune Cells by Eicosanoid Receptors
Regulation of Immune Cells by Eicosanoid Receptors The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation Kim, Nancy D., and Andrew D. Luster. 2007. “Regulation of Immune Cells by Eicosanoid Receptors.” The Scientific World Journal 7 (1): 1307-1328. doi:10.1100/tsw.2007.181. http://dx.doi.org/10.1100/ tsw.2007.181. Published Version doi:10.1100/tsw.2007.181 Citable link http://nrs.harvard.edu/urn-3:HUL.InstRepos:37298366 Terms of Use This article was downloaded from Harvard University’s DASH repository, and is made available under the terms and conditions applicable to Other Posted Material, as set forth at http:// nrs.harvard.edu/urn-3:HUL.InstRepos:dash.current.terms-of- use#LAA Review Article Special Issue: Eicosanoid Receptors and Inflammation TheScientificWorldJOURNAL (2007) 7, 1307–1328 ISSN 1537-744X; DOI 10.1100/tsw.2007.181 Regulation of Immune Cells by Eicosanoid Receptors Nancy D. Kim and Andrew D. Luster* Center for Immunology and Inflammatory Diseases, Division of Rheumatology, Allergy, and Immunology, Massachusetts General Hospital, Harvard Medical School, Boston E-mail: [email protected] Received March 13, 2007; Revised June 14, 2007; Accepted July 2, 2007; Published September 1, 2007 Eicosanoids are potent, bioactive, lipid mediators that regulate important components of the immune response, including defense against infection, ischemia, and injury, as well as instigating and perpetuating autoimmune and inflammatory conditions. Although these lipids have numerous effects on diverse cell types and organs, a greater understanding of their specific effects on key players of the immune system has been gained in recent years through the characterization of individual eicosanoid receptors, the identification and development of specific receptor agonists and inhibitors, and the generation of mice genetically deficient in various eicosanoid receptors. -

(Apigenin, Luteolin, Chrysin) and Their Importance for Health
Mellifera 2020, 20(1):16-27 MELLIFERA REVIEW ARTICLE Flavones (Apigenin, Luteolin, Chrysin) and Their Importance for Health Fatma Özsel ÖZCAN1*, Ozan ALDEMİR2, Bülent KARABULUT3 1Beykoz Üniversity, Faculty of Art and Design, Department of Gastronomy and Culinary Arts, İstanbul, Turkey 2Ege University Faculty of Medicine, Department of Biochemistry, İzmir, TURKEY 3Ege University Faculty of Medicine, Department of Oncology, İzmir, TURKEY * Corresponding author e-mail: [email protected] Received: 14th March, 2020; accepted: 4th May, 2020; published: 29th June, 2020 A B S T R A C T It has been shown in recent years that foods called functional foods may protect against certain types of cancer, cardiovascular diseases and cognitive dysfunctions. In the studies performed, the flavonoids (apigenin, chrysin, luteolin) which are subclass of flavonoids have been shown to have antioxidant, antiinflammatory, antiallergic, neuroprotective and cardioprotective effects and it is presented as the current treatment method in the treatment of some diseases. The structure function, nutritional resources and potential therapeutic properties of the flavones, which are also used as supplement in the compost, have been studied. The purpose of this review is to evaluate the therapeutic effects of flavones in certain diseases. The positive effect of 16 flavones on health can be proven in many experimental studies and can be proven in the long run. Keywords: Functional nutrients, flavones, apigenin, luteolin, chrysin Introduction Polyphenolic flavonoids are among the affected by factors such as plant species, wide variety of phytochemicals found in the environment, genetics, light, maturity, human diet. Current studies reveal that harvest [3]. Flavones, a subset of dietary flavonoids are inversely related to flavonoids, form glycosylation and contain many cancers and age-related diseases [1]. -

Glucocorticoid Regulation of the G-Protein Coupled Estrogen Receptor (GPER) in Mouse Hippocampal Neurons
Glucocorticoid Regulation of the G-Protein Coupled Estrogen Receptor (GPER) in Mouse Hippocampal Neurons by Kate Colleen Eliza Nicholson A Thesis presented to The University of Guelph In partial fulfilment of requirements for the degree of Master of Science in Biomedical Sciences Guelph, Ontario, Canada © Kate Colleen Eliza Nicholson, August, 2019 ABSTRACT GLUCOCORTICOID REGULATION OF THE G-PROTEIN COUPLED ESTROGEN RECEPTOR (GPER) IN MOUSE HIPPOCAMPAL NEURONS Kate Colleen Eliza Nicholson Advisor: University of Guelph, 2019 Dr. Neil J. MacLusky The most prevalent estrogen, 17β-estradiol, binds the non-classical G-protein coupled estrogen receptor (GPER) with high affinity resulting in rapid activation of the c- jun N terminal kinase (JNK) pathway. GPER activation mediates some of the rapid neurotrophic and memory-enhancing effects of 17β-estradiol in the female hippocampus. However, exposure to stressful stimuli may impair these beneficial effects. This thesis characterizes neurosteroid receptor expression in murine-derived mHippoE cell lines that are subsequently used to investigate the glucocorticoid regulation of GPER protein expression and functional activation. This thesis demonstrates that 24-hour treatment with a glucocorticoid receptor agonist reduces GPER protein expression and activation of JNK in female-derived mHippoE-14s. Using an in vivo model, treatment with glucocorticoids significantly reduces hippocampal activation of JNK in female ovariectomized CD1 mice. Collectively, this thesis uses in vitro and in vivo models to characterize glucocorticoid regulation of GPER expression and signalling in female murine hippocampal neurons. ACKNOWLEDGEMENTS To Dr. MacLusky: I would like to thank you for inspiring my passion for science and pursuit of knowledge. Over the past 2 years, you have provided me with countless opportunities to grow as a young researcher and I am tremendously grateful for this. -

G Protein-Coupled Estrogen Receptor Inhibits the P2Y Receptor-Mediated Ca2+ Signaling Pathway in Human Airway Epithelia
Pflugers Arch - Eur J Physiol DOI 10.1007/s00424-016-1840-7 SIGNALING AND CELL PHYSIOLOGY G protein-coupled estrogen receptor inhibits the P2Y receptor-mediated Ca2+ signaling pathway in human airway epithelia Yuan Hao1 & Alison W. Chow1 & Wallace C. Yip 1 & Chi H. Li1 & Tai F. Wan 1 & Benjamin C. Tong2 & King H. Cheung2 & Wood Y. Chan 1 & Yangchao Chen 1 & Christopher H. Cheng1 & Wing H. Ko1 Received: 21 January 2016 /Revised: 11 May 2016 /Accepted: 22 May 2016 # The Author(s) 2016. This article is published with open access at Springerlink.com Abstract P2Y receptor activation causes the release of in- inhibitory effects of G1 or E2 on P2Y receptor-mediated flammatory cytokines in the bronchial epithelium, whereas Ca2+ mobilization and cytokine secretion were due to G protein-coupled estrogen receptor (GPER), a novel estrogen GPER-mediated activation of a cAMP-dependent PKA path- (E2) receptor, may play an anti-inflammatory role in this pro- way. This study has reported, for the first time, the expression cess. We investigated the cellular mechanisms underlying the and function of GPER as an anti-inflammatory component in inhibitory effect of GPER activation on the P2Y receptor- human bronchial epithelia, which may mediate through its mediated Ca2+ signaling pathway and cytokine production in opposing effects on the pro‐inflammatory pathway activated airway epithelia. Expression of GPER in primary human by the P2Y receptors in inflamed airway epithelia. bronchial epithelial (HBE) or 16HBE14o- cells was con- firmed on both the mRNA and protein levels. Stimulation of Keywords GPER . P2Y receptor signaling pathway . Human HBE or 16HBE14o- cells with E2 or G1, a specific agonist of .