Comparison of Nitrile Hydratases in Rhodococcus Rhodochrous DAP 96253 and DAP 96622 Growing on Inducing and Non- Inducing Media
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Genomes of Polyextremophilic Cyanidiales Contain 1% 2 Horizontally Transferred Genes with Diverse Adaptive Functions 3 4 Alessandro W
bioRxiv preprint doi: https://doi.org/10.1101/526111; this version posted January 23, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 1 The genomes of polyextremophilic Cyanidiales contain 1% 2 horizontally transferred genes with diverse adaptive functions 3 4 Alessandro W. Rossoni1#, Dana C. Price2, Mark Seger3, Dagmar Lyska1, Peter Lammers3, 5 Debashish Bhattacharya4 & Andreas P.M. Weber1* 6 7 1Institute of Plant Biochemistry, Cluster of Excellence on Plant Sciences (CEPLAS), Heinrich 8 Heine University, Universitätsstraße 1, 40225 Düsseldorf, Germany 9 2Department of Plant Biology, Rutgers University, New Brunswick, NJ 08901, USA 10 3Arizona Center for Algae Technology and Innovation, Arizona State University, Mesa, AZ 11 85212, USA 12 4Department of Biochemistry and Microbiology, Rutgers University, New Brunswick, NJ 13 08901, USA 14 15 *Corresponding author: Prof. Dr. Andreas P.M. Weber, 16 e-mail: [email protected] 17 bioRxiv preprint doi: https://doi.org/10.1101/526111; this version posted January 23, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 18 Abstract 19 The role and extent of horizontal gene transfer (HGT) in eukaryotes are hotly disputed topics 20 that impact our understanding regarding the origin of metabolic processes and the role of 21 organelles in cellular evolution. -
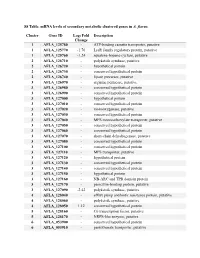
S8 Table. Mrna Levels of Secondary Metabolic Clustered Genes in A
S8 Table. mRNA levels of secondary metabolic clustered genes in A. flavus. Cluster Gene ID Log2 Fold Description Change 1 AFLA_125780 - ATP-binding cassette transporter, putative 1 AFLA_125770 -1.76 LysR family regulatory protein, putative 1 AFLA_125760 -1.24 squalene-hopene-cyclase, putative 2 AFLA_126710 - polyketide synthase, putative 2 AFLA_126720 - hypothetical protein 2 AFLA_126730 - conserved hypothetical protein 2 AFLA_126740 - lipase precursor, putative 3 AFLA_126970 - arginine permease, putative 3 AFLA_126980 - conserved hypothetical protein 3 AFLA_126990 - conserved hypothetical protein 3 AFLA_127000 - hypothetical protein 3 AFLA_127010 - conserved hypothetical protein 3 AFLA_127020 - monooxygenase, putative 3 AFLA_127030 - conserved hypothetical protein 3 AFLA_127040 - MFS monocarboxylate transporter, putative 3 AFLA_127050 - conserved hypothetical protein 3 AFLA_127060 - conserved hypothetical protein 3 AFLA_127070 - short-chain dehydrogenase, putative 3 AFLA_127080 - conserved hypothetical protein 3 AFLA_127100 - conserved hypothetical protein 3 AFLA_127110 - MFS transporter, putative 3 AFLA_127120 - hypothetical protein 3 AFLA_127130 - conserved hypothetical protein 3 AFLA_127140 - conserved hypothetical protein 3 AFLA_127150 - hypothetical protein 3 AFLA_127160 - NB-ARC and TPR domain protein 3 AFLA_127170 - penicillin-binding protein, putative 3 AFLA_127090 -2.42 polyketide synthase, putative 4 AFLA_128040 - efflux pump antibiotic resistance protein, putative 4 AFLA_128060 - polyketide synthase, putative 4 AFLA_128050 -

Comparative Genomics of Bradyrhizobium Japonicum CPAC
Siqueira et al. BMC Genomics 2014, 15:420 http://www.biomedcentral.com/1471-2164/15/420 RESEARCH ARTICLE Open Access Comparative genomics of Bradyrhizobium japonicum CPAC 15 and Bradyrhizobium diazoefficiens CPAC 7: elite model strains for understanding symbiotic performance with soybean Arthur Fernandes Siqueira1,2†, Ernesto Ormeño-Orrillo3†,RangelCelsoSouza4,ElisetePainsRodrigues5, Luiz Gonzaga Paula Almeida4, Fernando Gomes Barcellos5, Jesiane Stefânia Silva Batista6, Andre Shigueyoshi Nakatani2, Esperanza Martínez-Romero3, Ana Tereza Ribeiro Vasconcelos4 and Mariangela Hungria1,2* Abstract Background: The soybean-Bradyrhizobium symbiosis can be highly efficient in fixing nitrogen, but few genomic sequences of elite inoculant strains are available. Here we contribute with information on the genomes of two commercial strains that are broadly applied to soybean crops in the tropics. B. japonicum CPAC 15 (=SEMIA 5079) is outstanding in its saprophytic capacity and competitiveness, whereas B. diazoefficiens CPAC 7 (=SEMIA 5080) is known for its high efficiency in fixing nitrogen. Both are well adapted to tropical soils. The genomes of CPAC 15 and CPAC 7 were compared to each other and also to those of B. japonicum USDA 6T and B. diazoefficiens USDA 110T. Results: Differences in genome size were found between species, with B. japonicum having larger genomes than B. diazoefficiens. Although most of the four genomes were syntenic, genome rearrangements within and between species were observed, including events in the symbiosis island. In addition to the symbiotic region, several genomic islands were identified. Altogether, these features must confer high genomic plasticity that might explain adaptation and differences in symbiotic performance. It was not possible to attribute known functions to half of the predicted genes. -

Discovery of an Alternate Metabolic Pathway for Urea Synthesis in Adult Aedes Aegypti Mosquitoes
Discovery of an alternate metabolic pathway for urea synthesis in adult Aedes aegypti mosquitoes Patricia Y. Scaraffia*†‡, Guanhong Tan§, Jun Isoe*†, Vicki H. Wysocki*§, Michael A. Wells*†, and Roger L. Miesfeld*† Departments of §Chemistry and *Biochemistry and Molecular Biophysics and †Center for Insect Science, University of Arizona, Tucson, AZ 85721-0088 Edited by Anthony A. James, University of California, Irvine, CA, and approved December 4, 2007 (received for review August 27, 2007) We demonstrate the presence of an alternate metabolic pathway We previously reported that mosquitoes dispose of toxic for urea synthesis in Aedes aegypti mosquitoes that converts uric ammonia through glutamine (Gln) and proline (Pro) synthesis, acid to urea via an amphibian-like uricolytic pathway. For these along with excretion of ammonia, uric acid, and urea (20). By studies, female mosquitoes were fed a sucrose solution containing using labeled isotopes and mass spectrometry techniques (21), 15 15 15 15 15 NH4Cl, [5- N]-glutamine, [ N]-proline, allantoin, or allantoic we have recently determined how the N from NH4Cl is acid. At 24 h after feeding, the feces were collected and analyzed incorporated into the amide side chain of Gln, and then into Pro, in a mass spectrometer. Specific enzyme inhibitors confirmed that in Ae. aegypti (22). In the present article we demonstrate that the 15 15 15 mosquitoes incorporate N from NH4Cl into [5- N]-glutamine nitrogen of the amide group of Gln contributes to uric acid and use the 15N of the amide group of glutamine to produce synthesis in mosquitoes and, surprisingly, that uric acid can be 15 labeled uric acid. -

Francisella Tularensis 6/06 Tularemia Is a Commonly Acquired Laboratory Colony Morphology Infection; All Work on Suspect F
Francisella tularensis 6/06 Tularemia is a commonly acquired laboratory Colony Morphology infection; all work on suspect F. tularensis cultures .Aerobic, fastidious, requires cysteine for growth should be performed at minimum under BSL2 .Grows poorly on Blood Agar (BA) conditions with BSL3 practices. .Chocolate Agar (CA): tiny, grey-white, opaque A colonies, 1-2 mm ≥48hr B .Cysteine Heart Agar (CHA): greenish-blue colonies, 2-4 mm ≥48h .Colonies are butyrous and smooth Gram Stain .Tiny, 0.2–0.7 μm pleomorphic, poorly stained gram-negative coccobacilli .Mostly single cells Growth on BA (A) 48 h, (B) 72 h Biochemical/Test Reactions .Oxidase: Negative A B .Catalase: Weak positive .Urease: Negative Additional Information .Can be misidentified as: Haemophilus influenzae, Actinobacillus spp. by automated ID systems .Infective Dose: 10 colony forming units Biosafety Level 3 agent (once Francisella tularensis is . Growth on CA (A) 48 h, (B) 72 h suspected, work should only be done in a certified Class II Biosafety Cabinet) .Transmission: Inhalation, insect bite, contact with tissues or bodily fluids of infected animals .Contagious: No Acceptable Specimen Types .Tissue biopsy .Whole blood: 5-10 ml blood in EDTA, and/or Inoculated blood culture bottle Swab of lesion in transport media . Gram stain Sentinel Laboratory Rule-Out of Francisella tularensis Oxidase Little to no growth on BA >48 h Small, grey-white opaque colonies on CA after ≥48 h at 35/37ºC Positive Weak Negative Positive Catalase Tiny, pleomorphic, faintly stained, gram-negative coccobacilli (red, round, and random) Perform all additional work in a certified Class II Positive Biosafety Cabinet Weak Negative Positive *Oxidase: Negative Urease *Catalase: Weak positive *Urease: Negative *Oxidase, Catalase, and Urease: Appearances of test results are not agent-specific. -

Purification and Characterization of a Novel Nitrilase of Rhodococcus
JOURNAL OF BACTERIOLOGY, Sept. 1990, p. 4807-4815 Vol. 172, No. 9 0021-9193/90/094807-09$02.00/0 Copyright X3 1990, American Society for Microbiology Purification and Characterization of a Novel Nitrilase of Rhodococcus rhodochrous K22 That Acts on Aliphatic Nitriles MICHIHIKO KOBAYASHI,* NORIYUKI YANAKA, TORU NAGASAWA, AND HIDEAKI YAMADA Department ofAgricultural Chemistry, Faculty ofAgriculture, Kyoto University, Sakyo-ku, Kyoto 606, Japan Received 18 April 1990/Accepted 8 June 1990 A novel nitrilase that preferentially catalyzes the hydrolysis of aliphatic nitriles to the corresponding carboxylic acids and ammonia was found in the cells of a facultative crotononitrile-utilizing actinomycete isolated from soil. The strain was taxonomically studied and identified as Rhodococcus rhodochrous. The nitrilase was purified, with 9.08% overall recovery, through five steps from a cell extract of the stain. After the last step, the purified enzyme appeared to be homogeneous, as judged by polyacrylamide gel electrophoresis, analytical centrifugation, and double immunodiffusion in agarose. The relative molecular weight values for the native enzyme, estimated from the ultracentrifugal equilibrium and by high-performance liquid chromatog- raphy, were approximately 604,000 + 30,000 and 650,000, respectively, and the enzyme consisted of 15 to 16 subunits identical in molecular weight (41,000). The enzyme acted on aliphatic olefinic nitriles such as crotononitrile and acrylonitrile as the most suitable substrates. The apparent Km values for crotononitrile and acrylonitrile were 18.9 and 1.14 mM, respectively. The nitrilase also catalyzed the direct hydrolysis of saturated aliphatic nitriles, such as valeronitrile, 4-chlorobutyronitrile, and glutaronitrile, to the correspond- ing acids without the formation of amide intermediates. -

Extended Spectrum Β-Lactamases and Class C Β
F1000Research 2020, 9:774 Last updated: 01 APR 2021 RESEARCH ARTICLE Extended spectrum β-lactamases and class C β-lactamases gene frequency in Pseudomonas aeruginosa isolated from various clinical specimens in Khartoum State, Sudan: a cross sectional study [version 1; peer review: 1 not approved] Dina N. Abdelrahman 1,2, Aya A. Taha2, Mazar M. Dafaallah2, Alaa Abdelgafoor Mohammed3, Abdel Rahim M. El Hussein 1, Ahmed I. Hashim 2, Yousif F. Hamedelnil2, Hisham N. Altayb 4 1Department of Virology, Central Laboratory, Khartoum, Sudan 2Department of Microbiology, College of Medical Laboratory Sciences, Sudan University of Science and Technology, Khartoum, Sudan 3Department of Pharmaceutical Biotechnology, College of Pharmacy, Ahfad University for Women, Omdurman, Khartoum, Sudan 4Biochemistry Department, Faculty of Sciences, King Abdulaziz University, Jeddah, Saudi Arabia v1 First published: 27 Jul 2020, 9:774 Open Peer Review https://doi.org/10.12688/f1000research.24818.1 Second version: 15 Sep 2020, 9:774 https://doi.org/10.12688/f1000research.24818.2 Reviewer Status Latest published: 01 Apr 2021, 9:774 https://doi.org/10.12688/f1000research.24818.3 Invited Reviewers 1 2 Abstract Background: Pseudomonas aeruginosa is a pathogenic bacterium, version 3 causing nosocomial infections with intrinsic and acquired resistance (revision) mechanisms to a large group of antibiotics, including β-lactams. This 01 Apr 2021 study aimed to determine the susceptibility pattern to selected antibiotics and to index the first reported β-lactamases gene version 2 (extended spectrum β-lactamases (ESBLs) genes and class C β- (revision) report report lactamases genes) frequency in Ps. aeruginosa in Khartoum State, 15 Sep 2020 Sudan. Methods: 121 Ps. -

Direct Link to Fulltext
ISOLATION, IDENTIFICATION AND SUBSTRATE SPECIFICITY OF A NITRILASE PRODUCING BACTERIA, Acidovorax sp. SK1 Soumya Koippully, Subha Swaraj Pattnaik, Siddhardha Busi* Address(es): Department of Microbiology, School of Life Sciences, Pondicherry University, Puducherry-605 014, India. *Corresponding author: [email protected] doi: 10.15414/jmbfs.2018.8.2.788-793 ARTICLE INFO ABSTRACT Received 13. 5. 2018 The process of biocatalysis or biotransformation remains the core of industrial biotechnology owing to its importance in the synthesis of Revised 3. 9. 2018 high-value products in a cost effective manner. Nitrile biotransformation has also achieved considerable attention in last few decades Accepted 5. 9. 2018 due to its widespread biotechnological and industrial applications. In the present study, a versatile, potent nitrile-degrading bacterium, Published 1. 10. 2018 Acidovorax sp. SK1 was isolated from the cracker waste dumping site of Sivakasi, Tamilnadu and characterized for its biocatalytic potential, nitrilase production and enzyme activity. A pH sensitive indicator-based assay was performed to identify the nitrile degrading ability of the isolated samples. Semi quantitative High performance thin layer chromatographic (HPTLC) method was performed for Regular article quantitative measurement of mandelic acid produced from degradation of nitrile compound, mandelonitrile. The optimization of medium and nutritional parameters were studied for the improvement of nitrilase activity, which indicated that maximum nitrilase activity was observed at an optimum pH of 7.0, agitation at 100 rpm, glucose as best carbon source (10 g/L) and yeast extract (0.1 g/L) as principal nitrogen source. Biomass is also a critical parameter in the biocatalysis of mandelonitrile to mandelic acid and at a biomass of 100 mg/L, maximum nitrilase activity of 0.026 I.U was observed. -

(12) United States Patent (10) Patent No.: US 9.422,609 B2 Teichberg (45) Date of Patent: Aug
USOO9422609B2 (12) United States Patent (10) Patent No.: US 9.422,609 B2 Teichberg (45) Date of Patent: Aug. 23, 2016 (54) METHODS, COMPOSITIONS AND DEVICES (58) Field of Classification Search FOR MANTAINING CHEMICAL BALANCE CPC ........................ C02F 1/725; C12Y 305/01005 OF CHLORINATED WATER USPC ........................... 210/754; 435/195, 227 231 See application file for complete search history. (75) Inventor: Vivian I. Teichberg, Savyon (IL) (56) References Cited (73) Assignees: Mia Levite, Savyon (IL); Yaar Teichberg, Savyon (IL); Nof Lyle U.S. PATENT DOCUMENTS Teichberg, Savyon (IL) 4,793,935 A * 12/1988 Stillman ............... CO2F 1.5236 21Of727 (*) Notice: Subject to any disclaimer, the term of this 6,673,582 B2 * 1/2004 McTavish ..................... 435/122 patent is extended or adjusted under 35 U.S.C. 154(b) by 1044 days. (Continued) (21) Appl. No.: 12/225.18O FOREIGN PATENT DOCUMENTS y x- - - 9 AU 41971 5, 1979 (22) PCT Filed: Mar. 14, 2007 GB 2025919 1, 1980 (86). PCT No.: PCT/L2007/OOO336 (Continued) S 371 (c)(1) OTHER PUBLICATIONS (2), (4) Date: Sep. 16, 2008 Examiner's Report Dated Oct. 6, 2010 From the Australian Govern (87) PCT Pub. No.: WO2007/107.981 ment, IP Australia Re. Application No. 2007228391. (Continued) PCT Pub. Date: Sep. 27, 2007 (65) Prior Publication Data Primary Examiner — Peter Keyworth (74) Attorney, Agent, or Firm — Browdy and Neimark, US 201OfO270228A1 Oct. 28, 2010 PLLC Related U.S. Application Data (57) ABSTRACT (60) Provisional application No. 60/783,028, filed on Mar. A composition-of-matter for use in water treatment, com 17, 2006. -

Purification and Characterization of Nitrilase from Fusarium Solani
Process Biochemistry 45 (2010) 1115–1120 Contents lists available at ScienceDirect Process Biochemistry journal homepage: www.elsevier.com/locate/procbio Purification and characterization of nitrilase from Fusarium solani IMI196840 Vojtechˇ Vejvoda a, David Kubácˇ a,Alzbˇ etaˇ Davidová a, Ondrejˇ Kaplan a, Miroslav Sulcˇ a,b, Ondrejˇ Svedaˇ a, Radka Chaloupková c, Ludmila Martínková a,∗ a Institute of Microbiology, Centre of Biocatalysis and Biotransformation, Academy of Sciences of the Czech Republic, Vídenskᡠ1083, CZ-142 20 Prague, Czech Republic b Department of Biochemistry, Faculty of Science, Charles University Prague, Hlavova 8, CZ-128 40 Prague, Czech Republic c Loschmidt Laboratories, Department of Experimental Biology and Centre of Biocatalysis and Biotransformation, Faculty of Science, Masaryk University, Kamenice 5/A4, CZ-625 00 Brno, Czech Republic article info abstract Article history: Nitrilase activity in Fusarium solani IMI196840 (approx. 1500 U l−1 of culture broth) was induced by 2- Received 11 November 2009 cyanopyridine. The enzyme was purified by a factor of 20.3 at a yield of 26.9%. According to gel filtration, Received in revised form 26 March 2010 the holoenzyme was an approx. 550-kDa homooligomer consisting of subunits with a molecular weight Accepted 30 March 2010 of approximately 40 kDa, as determined by SDS-PAGE. Mass spectrometry analysis of the tryptic frag- ments suggested a high similarity of this enzyme to the hypothetical CN hydrolases from Aspergillus Keywords: oryzae, Gibberella zeae, Gibberella moniliformis and Nectria haematococca. Circular dichroism and fluo- Fusarium solani rescence spectra indicated that secondary structure content and overall tertiary structure, respectively, Nitrilase were almost identical in nitrilases from F. -

Structures, Functions, and Mechanisms of Filament Forming Enzymes: a Renaissance of Enzyme Filamentation
Structures, Functions, and Mechanisms of Filament Forming Enzymes: A Renaissance of Enzyme Filamentation A Review By Chad K. Park & Nancy C. Horton Department of Molecular and Cellular Biology University of Arizona Tucson, AZ 85721 N. C. Horton ([email protected], ORCID: 0000-0003-2710-8284) C. K. Park ([email protected], ORCID: 0000-0003-1089-9091) Keywords: Enzyme, Regulation, DNA binding, Nuclease, Run-On Oligomerization, self-association 1 Abstract Filament formation by non-cytoskeletal enzymes has been known for decades, yet only relatively recently has its wide-spread role in enzyme regulation and biology come to be appreciated. This comprehensive review summarizes what is known for each enzyme confirmed to form filamentous structures in vitro, and for the many that are known only to form large self-assemblies within cells. For some enzymes, studies describing both the in vitro filamentous structures and cellular self-assembly formation are also known and described. Special attention is paid to the detailed structures of each type of enzyme filament, as well as the roles the structures play in enzyme regulation and in biology. Where it is known or hypothesized, the advantages conferred by enzyme filamentation are reviewed. Finally, the similarities, differences, and comparison to the SgrAI system are also highlighted. 2 Contents INTRODUCTION…………………………………………………………..4 STRUCTURALLY CHARACTERIZED ENZYME FILAMENTS…….5 Acetyl CoA Carboxylase (ACC)……………………………………………………………………5 Phosphofructokinase (PFK)……………………………………………………………………….6 -

Letters to Nature
letters to nature Received 7 July; accepted 21 September 1998. 26. Tronrud, D. E. Conjugate-direction minimization: an improved method for the re®nement of macromolecules. Acta Crystallogr. A 48, 912±916 (1992). 1. Dalbey, R. E., Lively, M. O., Bron, S. & van Dijl, J. M. The chemistry and enzymology of the type 1 27. Wolfe, P. B., Wickner, W. & Goodman, J. M. Sequence of the leader peptidase gene of Escherichia coli signal peptidases. Protein Sci. 6, 1129±1138 (1997). and the orientation of leader peptidase in the bacterial envelope. J. Biol. Chem. 258, 12073±12080 2. Kuo, D. W. et al. Escherichia coli leader peptidase: production of an active form lacking a requirement (1983). for detergent and development of peptide substrates. Arch. Biochem. Biophys. 303, 274±280 (1993). 28. Kraulis, P.G. Molscript: a program to produce both detailed and schematic plots of protein structures. 3. Tschantz, W. R. et al. Characterization of a soluble, catalytically active form of Escherichia coli leader J. Appl. Crystallogr. 24, 946±950 (1991). peptidase: requirement of detergent or phospholipid for optimal activity. Biochemistry 34, 3935±3941 29. Nicholls, A., Sharp, K. A. & Honig, B. Protein folding and association: insights from the interfacial and (1995). the thermodynamic properties of hydrocarbons. Proteins Struct. Funct. Genet. 11, 281±296 (1991). 4. Allsop, A. E. et al.inAnti-Infectives, Recent Advances in Chemistry and Structure-Activity Relationships 30. Meritt, E. A. & Bacon, D. J. Raster3D: photorealistic molecular graphics. Methods Enzymol. 277, 505± (eds Bently, P. H. & O'Hanlon, P. J.) 61±72 (R. Soc. Chem., Cambridge, 1997).