Global Identification of MLL2-Targeted Loci Reveals Mll2ts Role in Diverse
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
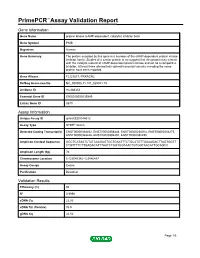
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name protein kinase (cAMP-dependent, catalytic) inhibitor beta Gene Symbol PKIB Organism Human Gene Summary The protein encoded by this gene is a member of the cAMP-dependent protein kinase inhibitor family. Studies of a similar protein in rat suggest that this protein may interact with the catalytic subunit of cAMP-dependent protein kinase and act as a competitive inhibitor. At least three alternatively spliced transcript variants encoding the same protein have been reported. Gene Aliases FLJ23817, PRKACN2 RefSeq Accession No. NC_000006.11, NT_025741.15 UniGene ID Hs.486354 Ensembl Gene ID ENSG00000135549 Entrez Gene ID 5570 Assay Information Unique Assay ID qHsaCED0044612 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000368452, ENST00000368448, ENST00000258014, ENST00000354275, ENST00000368446, ENST00000392491, ENST00000392490 Amplicon Context Sequence GGCTCATAATCTATCAAGAGTGCTGAATTTCTGCATGTTGAAAGACTTAGTGGTT CTGTTTTCTTGAGACATTTAATCTGGTGGTAACTGTGGTAACATTGCAGCC Amplicon Length (bp) 76 Chromosome Location 6:123046342-123046447 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 95 R2 0.9996 cDNA Cq 22.83 cDNA Tm (Celsius) 76.5 gDNA Cq 24.52 Page 1/5 PrimePCR™Assay Validation Report Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report PKIB, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve -

Location Analysis of Estrogen Receptor Target Promoters Reveals That
Location analysis of estrogen receptor ␣ target promoters reveals that FOXA1 defines a domain of the estrogen response Jose´ e Laganie` re*†, Genevie` ve Deblois*, Ce´ line Lefebvre*, Alain R. Bataille‡, Franc¸ois Robert‡, and Vincent Gigue` re*†§ *Molecular Oncology Group, Departments of Medicine and Oncology, McGill University Health Centre, Montreal, QC, Canada H3A 1A1; †Department of Biochemistry, McGill University, Montreal, QC, Canada H3G 1Y6; and ‡Laboratory of Chromatin and Genomic Expression, Institut de Recherches Cliniques de Montre´al, Montreal, QC, Canada H2W 1R7 Communicated by Ronald M. Evans, The Salk Institute for Biological Studies, La Jolla, CA, July 1, 2005 (received for review June 3, 2005) Nuclear receptors can activate diverse biological pathways within general absence of large scale functional data linking these putative a target cell in response to their cognate ligands, but how this binding sites with gene expression in specific cell types. compartmentalization is achieved at the level of gene regulation is Recently, chromatin immunoprecipitation (ChIP) has been used poorly understood. We used a genome-wide analysis of promoter in combination with promoter or genomic DNA microarrays to occupancy by the estrogen receptor ␣ (ER␣) in MCF-7 cells to identify loci recognized by transcription factors in a genome-wide investigate the molecular mechanisms underlying the action of manner in mammalian cells (20–24). This technology, termed 17-estradiol (E2) in controlling the growth of breast cancer cells. ChIP-on-chip or location analysis, can therefore be used to deter- We identified 153 promoters bound by ER␣ in the presence of E2. mine the global gene expression program that characterize the Motif-finding algorithms demonstrated that the estrogen re- action of a nuclear receptor in response to its natural ligand. -

Rage (Receptor for Advanced Glycation End Products) in Melanoma
RAGE (RECEPTOR FOR ADVANCED GLYCATION END PRODUCTS) IN MELANOMA PROGRESSION A Dissertation Submitted to the Graduate Faculty of the North Dakota State University of Agriculture and Applied Science By Varsha Meghnani In Partial Fulfillment for the Degree of DOCTOR OF PHILOSOPHY Major Department: Pharmaceutical Sciences May 2014 Fargo, North Dakota North Dakota State University Graduate School Title RAGE (RECEPTOR FOR ADVANCED GLYCATION END PRODUCTS) IN MELANOMA PROGRESSION By VARSHA MEGHNANI The Supervisory Committee certifies that this disquisition complies with North Dakota State University’s regulations and meets the accepted standards for the degree of DOCTOR OF PHILOSOPHY SUPERVISORY COMMITTEE: ESTELLE LECLERC Chair BIN GUO STEPHEN O’ROURKE JANE SCHUH Approved: 5/22/2014 JAGDISH SINGH Date Department Chair ABSTRACT The Receptor for Advanced Glycation End Products (RAGE) and its ligands are expressed in multiple cancer types and are implicated in cancer progression. Our lab has previously reported higher transcript levels of RAGE and S100B in advanced staged melanoma patients. The contribution of RAGE in the pathophysiology of melanoma has not been well studied. Based on previous reports, we hypothesized that RAGE, when over-expressed in melanoma cells, promotes melanoma progression. To study the pathogenic role of RAGE in melanoma, a primary melanoma cell line, WM115, was selected and stably transfected with full length RAGE (FL_RAGE) to generate a model cell line over-expressing RAGE (WM115_RAGE). WM266, a sister cell line of WM115, originated from a metastatic tumor of the same patient was used as a metastatic control cell line in the study. After assessing the expression levels of RAGE in the transfected cells, the influence of RAGE on their cellular properties was examined. -

The UVB-Induced Gene Expression Profile of Human Epidermis in Vivo Is Different from That of Cultured Keratinocytes
Oncogene (2006) 25, 2601–2614 & 2006 Nature Publishing Group All rights reserved 0950-9232/06 $30.00 www.nature.com/onc ORIGINAL ARTICLE The UVB-induced gene expression profile of human epidermis in vivo is different from that of cultured keratinocytes CD Enk1, J Jacob-Hirsch2, H Gal3, I Verbovetski4, N Amariglio2, D Mevorach4, A Ingber1, D Givol3, G Rechavi2 and M Hochberg1 1Department of Dermatology, The Hadassah-Hebrew University Medical Center, Jerusalem, Israel; 2Department of Pediatric Hemato-Oncology and Functional Genomics, Safra Children’s Hospital, Sheba Medical Center and Sackler School of Medicine, Tel-Aviv University,Tel Aviv, Israel; 3Department of Molecular Cell Biology, Weizmann Institute of Science, Rehovot, Israel and 4The Laboratory for Cellular and Molecular Immunology, Department of Medicine, The Hadassah-Hebrew University Medical Center, Jerusalem, Israel In order to obtain a comprehensive picture of the radiation. UVB, with a wavelength range between 290 molecular events regulating cutaneous photodamage of and 320 nm, represents one of the most important intact human epidermis, suction blister roofs obtained environmental hazards affectinghuman skin (Hahn after a single dose of in vivo ultraviolet (UV)B exposure and Weinberg, 2002). To protect itself against the were used for microarray profiling. We found a changed DNA-damaging effects of sunlight, the skin disposes expression of 619 genes. Half of the UVB-regulated genes over highly complicated cellular programs, including had returned to pre-exposure baseline levels at 72 h, cell-cycle arrest, DNA repair and apoptosis (Brash et al., underscoring the transient character of the molecular 1996). Failure in selected elements of these defensive cutaneous UVB response. -

8296.Full.Pdf
Inflammation-Induced Chondrocyte Hypertrophy Is Driven by Receptor for Advanced Glycation End Products This information is current as Denise L. Cecil, Kristen Johnson, John Rediske, Martin of September 28, 2021. Lotz, Ann Marie Schmidt and Robert Terkeltaub J Immunol 2005; 175:8296-8302; ; doi: 10.4049/jimmunol.175.12.8296 http://www.jimmunol.org/content/175/12/8296 Downloaded from References This article cites 43 articles, 13 of which you can access for free at: http://www.jimmunol.org/content/175/12/8296.full#ref-list-1 http://www.jimmunol.org/ Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication by guest on September 28, 2021 *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2005 by The American Association of Immunologists All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Inflammation-Induced Chondrocyte Hypertrophy Is Driven by Receptor for Advanced Glycation End Products1 Denise L. Cecil,* Kristen Johnson,* John Rediske,‡ Martin Lotz,§ Ann Marie Schmidt,† and Robert Terkeltaub2* The multiligand receptor for advanced glycation end products (RAGE) mediates certain chronic vascular and neurologic degen- erative diseases accompanied by low-grade inflammation. -

(Rage) in Progression of Pancreatic Cancer
The Texas Medical Center Library DigitalCommons@TMC The University of Texas MD Anderson Cancer Center UTHealth Graduate School of The University of Texas MD Anderson Cancer Biomedical Sciences Dissertations and Theses Center UTHealth Graduate School of (Open Access) Biomedical Sciences 8-2017 INVOLVEMENT OF THE RECEPTOR FOR ADVANCED GLYCATION END PRODUCTS (RAGE) IN PROGRESSION OF PANCREATIC CANCER Nancy Azizian MS Follow this and additional works at: https://digitalcommons.library.tmc.edu/utgsbs_dissertations Part of the Biology Commons, and the Medicine and Health Sciences Commons Recommended Citation Azizian, Nancy MS, "INVOLVEMENT OF THE RECEPTOR FOR ADVANCED GLYCATION END PRODUCTS (RAGE) IN PROGRESSION OF PANCREATIC CANCER" (2017). The University of Texas MD Anderson Cancer Center UTHealth Graduate School of Biomedical Sciences Dissertations and Theses (Open Access). 748. https://digitalcommons.library.tmc.edu/utgsbs_dissertations/748 This Dissertation (PhD) is brought to you for free and open access by the The University of Texas MD Anderson Cancer Center UTHealth Graduate School of Biomedical Sciences at DigitalCommons@TMC. It has been accepted for inclusion in The University of Texas MD Anderson Cancer Center UTHealth Graduate School of Biomedical Sciences Dissertations and Theses (Open Access) by an authorized administrator of DigitalCommons@TMC. For more information, please contact [email protected]. INVOLVEMENT OF THE RECEPTOR FOR ADVANCED GLYCATION END PRODUCTS (RAGE) IN PROGRESSION OF PANCREATIC CANCER by Nancy -

Investigation of the Underlying Hub Genes and Molexular Pathogensis in Gastric Cancer by Integrated Bioinformatic Analyses
bioRxiv preprint doi: https://doi.org/10.1101/2020.12.20.423656; this version posted December 22, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Investigation of the underlying hub genes and molexular pathogensis in gastric cancer by integrated bioinformatic analyses Basavaraj Vastrad1, Chanabasayya Vastrad*2 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karanataka, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India bioRxiv preprint doi: https://doi.org/10.1101/2020.12.20.423656; this version posted December 22, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract The high mortality rate of gastric cancer (GC) is in part due to the absence of initial disclosure of its biomarkers. The recognition of important genes associated in GC is therefore recommended to advance clinical prognosis, diagnosis and and treatment outcomes. The current investigation used the microarray dataset GSE113255 RNA seq data from the Gene Expression Omnibus database to diagnose differentially expressed genes (DEGs). Pathway and gene ontology enrichment analyses were performed, and a proteinprotein interaction network, modules, target genes - miRNA regulatory network and target genes - TF regulatory network were constructed and analyzed. Finally, validation of hub genes was performed. The 1008 DEGs identified consisted of 505 up regulated genes and 503 down regulated genes. -

Molecular Effects of Isoflavone Supplementation Human Intervention Studies and Quantitative Models for Risk Assessment
Molecular effects of isoflavone supplementation Human intervention studies and quantitative models for risk assessment Vera van der Velpen Thesis committee Promotors Prof. Dr Pieter van ‘t Veer Professor of Nutritional Epidemiology Wageningen University Prof. Dr Evert G. Schouten Emeritus Professor of Epidemiology and Prevention Wageningen University Co-promotors Dr Anouk Geelen Assistant professor, Division of Human Nutrition Wageningen University Dr Lydia A. Afman Assistant professor, Division of Human Nutrition Wageningen University Other members Prof. Dr Jaap Keijer, Wageningen University Dr Hubert P.J.M. Noteborn, Netherlands Food en Consumer Product Safety Authority Prof. Dr Yvonne T. van der Schouw, UMC Utrecht Dr Wendy L. Hall, King’s College London This research was conducted under the auspices of the Graduate School VLAG (Advanced studies in Food Technology, Agrobiotechnology, Nutrition and Health Sciences). Molecular effects of isoflavone supplementation Human intervention studies and quantitative models for risk assessment Vera van der Velpen Thesis submitted in fulfilment of the requirements for the degree of doctor at Wageningen University by the authority of the Rector Magnificus Prof. Dr M.J. Kropff, in the presence of the Thesis Committee appointed by the Academic Board to be defended in public on Friday 20 June 2014 at 13.30 p.m. in the Aula. Vera van der Velpen Molecular effects of isoflavone supplementation: Human intervention studies and quantitative models for risk assessment 154 pages PhD thesis, Wageningen University, Wageningen, NL (2014) With references, with summaries in Dutch and English ISBN: 978-94-6173-952-0 ABSTRact Background: Risk assessment can potentially be improved by closely linked experiments in the disciplines of epidemiology and toxicology. -

75 2. INTRODUCTION Triple-Negative Breast Cancer (TNBC)
[Frontiers in Bioscience, Scholar, 11, 75-88, March 1, 2019] The persisting puzzle of racial disparity in triple negative breast cancer: looking through a new lens Chakravarthy Garlapati1, Shriya Joshi1, Bikram Sahoo1, Shobhna Kapoor2, Ritu Aneja1 1Department of Biology, Georgia State University, Atlanta, GA, USA, 2Department of Chemistry, Indian Institute of Technology Bombay, Powai, India TABLE OF CONTENTS 1. Abstract 2. Introduction 3. Dissecting the TNBC racially disparate burden 3.1. Does race influence TNBC onset and progression? 3.2. Tumor microenvironment in TNBC and racial disparity 3.3. Differential gene signatures and pathways in racially distinct TNBC 3.4. Our Perspective: Looking racial disparity through a new lens 4. Conclusion 5. Acknowledgement 6. References 1. ABSTRACT 2. INTRODUCTION Triple-negative breast cancer (TNBC) Triple-negative breast cancer (TNBC), is characterized by the absence of estrogen a subtype of breast cancer (BC), accounts for and progesterone receptors and absence 15-20% of all BC diagnoses in the US. It has of amplification of human epidermal growth been recognized that women of African descent factor receptor (HER2). This disease has no are twice as likely to develop TNBC than approved treatment with a poor prognosis women of European descent (1). As the name particularly in African-American (AA) as foretells, TNBCs lack estrogen, progesterone, compared to European-American (EA) and human epidermal growth factor receptors. patients. Gene ontology analysis showed Unfortunately, TNBCs are defined by what they specific gene pathways that are differentially “lack” rather than what they “have” and thus this regulated and gene signatures that are negative nomenclature provides no actionable differentially expressed in AA as compared to information on “druggable” targets. -

Transcriptional Activation of the Human S100A2 Promoter by Wild-Type P53
FEBS 21606 FEBS Letters 445 (1999) 265^268 Transcriptional activation of the human S100A2 promoter by wild-type p53 Mingjia Tana, Claus W. Heizmannb, Kunliang Guanc, Beat W. Schaferb, Yi Suna;* aDepartment of Molecular Biology, Parke-Davis Pharmaceutical Research, Division of Warner-Lambert Company, Ann Arbor, MI 48105, USA bDepartment of Pediatrics, Division of Clinical Chemistry and Biochemistry, University of Zurich, Zurich, Switzerland cDepartment of Biological Chemistry, University of Michigan, Ann Arbor, MI 48109, USA Received 18 January 1999 2. Materials and methods Abstract S100A2, a calcium binding protein of the EF-hand family, was recently identified to be inducible by etoposide, a p53 2.1. Cell culture and drug treatment activator. A potential p53 binding site was identified in the Two human osteogenic sarcoma cell lines, U2-OS and Saos-2 promoter of the S100A2 gene, which binds to purified p53 as well (ATCC), were grown in McCory or DMEM supplemented with as p53 in nuclear extract activated by etoposide. Transactivation 10% FCS, respectively. U2-OS cells harbor a wild-type p53 [13], assays using the promoter driven luciferase reporters revealed whereas Saos-2 cells have the p53 gene deleted [14]. For drug treat- that the S100A2 promoter was transcriptionally activated by ment, U2-OS and Saos-2 cells were exposed to etoposide (25 WM, wild-type p53, but not by p53 mutants, in a dose-dependent as Sigma) for various periods of time up to 48 h. well as a p53 binding site-dependent manner. The p53-induced 2.2. Gel shift assay transactivation of the S100A2 promoter was enhanced by The assay was performed as described previously [15], using as the etoposide and blocked by a dominant negative p53 mutant. -

S100 Alpha 2 (S100A2) (NM 005978) Human Tagged ORF Clone Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for RC201203 S100 alpha 2 (S100A2) (NM_005978) Human Tagged ORF Clone Product data: Product Type: Expression Plasmids Product Name: S100 alpha 2 (S100A2) (NM_005978) Human Tagged ORF Clone Tag: Myc-DDK Symbol: S100A2 Synonyms: CAN19; S100L Vector: pCMV6-Entry (PS100001) E. coli Selection: Kanamycin (25 ug/mL) Cell Selection: Neomycin ORF Nucleotide >RC201203 ORF sequence Sequence: Red=Cloning site Blue=ORF Green=Tags(s) TTTTGTAATACGACTCACTATAGGGCGGCCGGGAATTCGTCGACTGGATCCGGTACCGAGGAGATCTGCC GCCGCGATCGCC ATGATGTGCAGTTCTCTGGAGCAGGCGCTGGCTGTGCTGGTCACTACCTTCCACAAGTACTCCTGCCAAG AGGGCGACAAGTTCAAGCTGAGTAAGGGGGAAATGAAGGAACTTCTGCACAAGGAGCTGCCCAGCTTTGT GGGGGAGAAAGTGGATGAGGAGGGGCTGAAGAAGCTGATGGGCAGCCTGGATGAGAACAGTGACCAGCAG GTGGACTTCCAGGAGTATGCTGTTTTCCTGGCACTCATCACTGTCATGTGCAATGACTTCTTCCAGGGCT GCCCAGACCGACCC ACGCGTACGCGGCCGCTCGAGCAGAAACTCATCTCAGAAGAGGATCTGGCAGCAAATGATATCCTGGATT ACAAGGATGACGACGATAAGGTTTAA Protein Sequence: >RC201203 protein sequence Red=Cloning site Green=Tags(s) MMCSSLEQALAVLVTTFHKYSCQEGDKFKLSKGEMKELLHKELPSFVGEKVDEEGLKKLMGSLDENSDQQ VDFQEYAVFLALITVMCNDFFQGCPDRP TRTRPLEQKLISEEDLAANDILDYKDDDDKV Chromatograms: https://cdn.origene.com/chromatograms/mk6143_g11.zip Restriction Sites: SgfI-MluI This product is to be used for laboratory only. Not for diagnostic or therapeutic use. View online » ©2021 OriGene Technologies, Inc., 9620 Medical -

Integrated Analysis of Gene Expression Changes Associated
Miao et al. Lipids in Health and Disease (2019) 18:92 https://doi.org/10.1186/s12944-019-1032-5 RESEARCH Open Access Integrated analysis of gene expression changes associated with coronary artery disease Liu Miao1 , Rui-Xing Yin1,2,3* , Feng Huang1,2,3, Shuo Yang1, Wu-Xian Chen1 and Jin-Zhen Wu1 Abstract Background: This study investigated the pathways and genes involved in coronary artery disease (CAD) and the associated mechanisms. Methods: Two array data sets of GSE19339 and GSE56885 were downloaded. The limma package was used to analyze the differentially expressed genes (DEGs) in normal and CAD specimens. Examination of DEGs through Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment and Gene Ontology annotation was achieved by Database for Annotation, Visualization and Integrated Discovery (DAVID). The Cytoscape software facilitated the establishment of the protein-protein interaction (PPI) network and Molecular Complex Detection (MCODE) was performed for the significant modules. Results: We identified 413 DEGs (291 up-regulated and 122 down-regulated). Approximately 256 biological processes, only 1 cellular component, and 21 molecular functions were identified by GO analysis and 10 pathways were enriched by KEGG. Moreover, 264 protein pairs and 64 nodes were visualized by the PPI network. After the MCODE analysis, the top 4 high degree genes, including interleukin 1 beta (IL1B, degree = 29), intercellular adhesion molecule 1 (ICAM1, degree = 25), Jun proto-oncogene (JUN, degree = 23) and C-C motif chemokine ligand 2 (CCL2, degree = 20) had been identified to validate in RT-PCR and Cox proportional hazards regression between CAD and normals. Conclusions: The relative expression of IL1B, ICAM1 and CCL2 was higher in CAD than in normal controls (P <0.05–0.