Characterization of Foxp2 Functions in the Mouse Cortex Vera Medvedeva
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Cellular and Synaptic Network Defects in Autism
Cellular and synaptic network defects in autism The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation Peca, Joao, and Guoping Feng. “Cellular and Synaptic Network Defects in Autism.” Current Opinion in Neurobiology 22, no. 5 (October 2012): 866–872. As Published http://dx.doi.org/10.1016/j.conb.2012.02.015 Publisher Elsevier Version Author's final manuscript Citable link http://hdl.handle.net/1721.1/102179 Terms of Use Creative Commons Attribution-Noncommercial-NoDerivatives Detailed Terms http://creativecommons.org/licenses/by-nc-nd/4.0/ NIH Public Access Author Manuscript Curr Opin Neurobiol. Author manuscript; available in PMC 2013 October 01. Published in final edited form as: Curr Opin Neurobiol. 2012 October ; 22(5): 866–872. doi:10.1016/j.conb.2012.02.015. Cellular and synaptic network defects in autism João Peça1 and Guoping Feng1,2 $watermark-text1McGovern $watermark-text Institute $watermark-text for Brain Research, Department of Brain and Cognitive Sciences, Massachusetts Institute of Technology, Cambridge, MA 02139, USA 2Stanley Center for Psychiatric Research, Broad Institute, Cambridge, MA 02142, USA Abstract Many candidate genes are now thought to confer susceptibility to autism spectrum disorder (ASD). Here we review four interrelated complexes, each composed of multiple families of genes that functionally coalesce on common cellular pathways. We illustrate a common thread in the organization of glutamatergic synapses and suggest a link between genes involved in Tuberous Sclerosis Complex, Fragile X syndrome, Angelman syndrome and several synaptic ASD candidate genes. When viewed in this context, progress in deciphering the molecular architecture of cellular protein-protein interactions together with the unraveling of synaptic dysfunction in neural networks may prove pivotal to advancing our understanding of ASDs. -
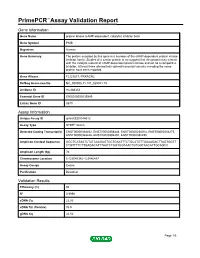
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name protein kinase (cAMP-dependent, catalytic) inhibitor beta Gene Symbol PKIB Organism Human Gene Summary The protein encoded by this gene is a member of the cAMP-dependent protein kinase inhibitor family. Studies of a similar protein in rat suggest that this protein may interact with the catalytic subunit of cAMP-dependent protein kinase and act as a competitive inhibitor. At least three alternatively spliced transcript variants encoding the same protein have been reported. Gene Aliases FLJ23817, PRKACN2 RefSeq Accession No. NC_000006.11, NT_025741.15 UniGene ID Hs.486354 Ensembl Gene ID ENSG00000135549 Entrez Gene ID 5570 Assay Information Unique Assay ID qHsaCED0044612 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000368452, ENST00000368448, ENST00000258014, ENST00000354275, ENST00000368446, ENST00000392491, ENST00000392490 Amplicon Context Sequence GGCTCATAATCTATCAAGAGTGCTGAATTTCTGCATGTTGAAAGACTTAGTGGTT CTGTTTTCTTGAGACATTTAATCTGGTGGTAACTGTGGTAACATTGCAGCC Amplicon Length (bp) 76 Chromosome Location 6:123046342-123046447 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 95 R2 0.9996 cDNA Cq 22.83 cDNA Tm (Celsius) 76.5 gDNA Cq 24.52 Page 1/5 PrimePCR™Assay Validation Report Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report PKIB, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve -

Location Analysis of Estrogen Receptor Target Promoters Reveals That
Location analysis of estrogen receptor ␣ target promoters reveals that FOXA1 defines a domain of the estrogen response Jose´ e Laganie` re*†, Genevie` ve Deblois*, Ce´ line Lefebvre*, Alain R. Bataille‡, Franc¸ois Robert‡, and Vincent Gigue` re*†§ *Molecular Oncology Group, Departments of Medicine and Oncology, McGill University Health Centre, Montreal, QC, Canada H3A 1A1; †Department of Biochemistry, McGill University, Montreal, QC, Canada H3G 1Y6; and ‡Laboratory of Chromatin and Genomic Expression, Institut de Recherches Cliniques de Montre´al, Montreal, QC, Canada H2W 1R7 Communicated by Ronald M. Evans, The Salk Institute for Biological Studies, La Jolla, CA, July 1, 2005 (received for review June 3, 2005) Nuclear receptors can activate diverse biological pathways within general absence of large scale functional data linking these putative a target cell in response to their cognate ligands, but how this binding sites with gene expression in specific cell types. compartmentalization is achieved at the level of gene regulation is Recently, chromatin immunoprecipitation (ChIP) has been used poorly understood. We used a genome-wide analysis of promoter in combination with promoter or genomic DNA microarrays to occupancy by the estrogen receptor ␣ (ER␣) in MCF-7 cells to identify loci recognized by transcription factors in a genome-wide investigate the molecular mechanisms underlying the action of manner in mammalian cells (20–24). This technology, termed 17-estradiol (E2) in controlling the growth of breast cancer cells. ChIP-on-chip or location analysis, can therefore be used to deter- We identified 153 promoters bound by ER␣ in the presence of E2. mine the global gene expression program that characterize the Motif-finding algorithms demonstrated that the estrogen re- action of a nuclear receptor in response to its natural ligand. -

Preclinical Evaluation of Protein Disulfide Isomerase Inhibitors for the Treatment of Glioblastoma by Andrea Shergalis
Preclinical Evaluation of Protein Disulfide Isomerase Inhibitors for the Treatment of Glioblastoma By Andrea Shergalis A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy (Medicinal Chemistry) in the University of Michigan 2020 Doctoral Committee: Professor Nouri Neamati, Chair Professor George A. Garcia Professor Peter J. H. Scott Professor Shaomeng Wang Andrea G. Shergalis [email protected] ORCID 0000-0002-1155-1583 © Andrea Shergalis 2020 All Rights Reserved ACKNOWLEDGEMENTS So many people have been involved in bringing this project to life and making this dissertation possible. First, I want to thank my advisor, Prof. Nouri Neamati, for his guidance, encouragement, and patience. Prof. Neamati instilled an enthusiasm in me for science and drug discovery, while allowing me the space to independently explore complex biochemical problems, and I am grateful for his kind and patient mentorship. I also thank my committee members, Profs. George Garcia, Peter Scott, and Shaomeng Wang, for their patience, guidance, and support throughout my graduate career. I am thankful to them for taking time to meet with me and have thoughtful conversations about medicinal chemistry and science in general. From the Neamati lab, I would like to thank so many. First and foremost, I have to thank Shuzo Tamara for being an incredible, kind, and patient teacher and mentor. Shuzo is one of the hardest workers I know. In addition to a strong work ethic, he taught me pretty much everything I know and laid the foundation for the article published as Chapter 3 of this dissertation. The work published in this dissertation really began with the initial identification of PDI as a target by Shili Xu, and I am grateful for his advice and guidance (from afar!). -

Modulating Hallmarks of Cholangiocarcinoma
University of Nebraska Medical Center DigitalCommons@UNMC Theses & Dissertations Graduate Studies Fall 12-14-2018 Modulating Hallmarks of Cholangiocarcinoma Cody Wehrkamp University of Nebraska Medical Center Follow this and additional works at: https://digitalcommons.unmc.edu/etd Part of the Molecular Biology Commons Recommended Citation Wehrkamp, Cody, "Modulating Hallmarks of Cholangiocarcinoma" (2018). Theses & Dissertations. 337. https://digitalcommons.unmc.edu/etd/337 This Dissertation is brought to you for free and open access by the Graduate Studies at DigitalCommons@UNMC. It has been accepted for inclusion in Theses & Dissertations by an authorized administrator of DigitalCommons@UNMC. For more information, please contact [email protected]. MODULATING HALLMARKS OF CHOLANGIOCARCINOMA by Cody J. Wehrkamp A DISSERTATION Presented to the Faculty of the University of Nebraska Graduate College in Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy Biochemistry and Molecular Biology Graduate Program Under the Supervision of Professor Justin L. Mott University of Nebraska Medical Center Omaha, Nebraska November 2018 Supervisory Committee: Kaustubh Datta, Ph.D. Melissa Teoh‐Fitzgerald, Ph.D. Richard G. MacDonald, Ph.D. Acknowledgements This endeavor has led to scientific as well as personal growth for me. I am indebted to many for their knowledge, influence, and support along the way. To my mentor, Dr. Justin L. Mott, you have been an incomparable teacher and invaluable guide. You upheld for me the concept that science is intrepid, even when the experience is trying. Through my training, and now here at the end, I can say that it has been an honor to be your protégé. When you have shaped your future graduates to be and do great, I will be privileged to say that I was your first one. -

Investigation of the Underlying Hub Genes and Molexular Pathogensis in Gastric Cancer by Integrated Bioinformatic Analyses
bioRxiv preprint doi: https://doi.org/10.1101/2020.12.20.423656; this version posted December 22, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Investigation of the underlying hub genes and molexular pathogensis in gastric cancer by integrated bioinformatic analyses Basavaraj Vastrad1, Chanabasayya Vastrad*2 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karanataka, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India bioRxiv preprint doi: https://doi.org/10.1101/2020.12.20.423656; this version posted December 22, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract The high mortality rate of gastric cancer (GC) is in part due to the absence of initial disclosure of its biomarkers. The recognition of important genes associated in GC is therefore recommended to advance clinical prognosis, diagnosis and and treatment outcomes. The current investigation used the microarray dataset GSE113255 RNA seq data from the Gene Expression Omnibus database to diagnose differentially expressed genes (DEGs). Pathway and gene ontology enrichment analyses were performed, and a proteinprotein interaction network, modules, target genes - miRNA regulatory network and target genes - TF regulatory network were constructed and analyzed. Finally, validation of hub genes was performed. The 1008 DEGs identified consisted of 505 up regulated genes and 503 down regulated genes. -

Lrrtms and Neuroligins Bind Neurexins with a Differential Code to Cooperate in Glutamate Synapse Development
The Journal of Neuroscience, June 2, 2010 • 30(22):7495–7506 • 7495 Cellular/Molecular LRRTMs and Neuroligins Bind Neurexins with a Differential Code to Cooperate in Glutamate Synapse Development Tabrez J. Siddiqui, Raika Pancaroglu, Yunhee Kang, Amanda Rooyakkers, and Ann Marie Craig Brain Research Centre and Department of Psychiatry, University of British Columbia, Vancouver, British Columbia V6T 2B5, Canada Leucine-rich repeat transmembrane neuronal proteins (LRRTMs) were recently found to instruct presynaptic and mediate postsynaptic glutamatergic differentiation. In a candidate screen, here we identify neurexin-1 lacking an insert at splice site 4 (ϪS4) as a ligand for LRRTM2. Neurexins bind LRRTM2 with a similar affinity but distinct code from the code for binding neuroligin-1 (the predominant form of neuroligin-1 at glutamate synapses, containing the B splice site insert). Whereas neuroligin-1 binds to neurexins 1, 2, and 3  but not ␣ variants, regardless of insert at splice site 4, LRRTM2 binds to neurexins 1, 2, and 3 ␣ and  variants specifically lacking an insert at splicesite4.WefurthershowthatthisbindingcodeisconservedinLRRTM1,thefamilymemberlinkedtoschizophreniaandhandedness, and that the code is functional in a coculture hemisynapse formation assay. Mutagenesis of LRRTM2 to prevent binding to neurexins abolishes presynaptic inducing activity of LRRTM2. Remarkably, mutagenesis of neurexins shows that the binding face on neurexin-1 (ϪS4) is highly overlapping for the structurally distinct LRRTM2 and neuroligin-1 partners. Finally, we explore here the interplay of neuroligin-1 and LRRTM2 in synapse regulation. In neuron cultures, LRRTM2 is more potent than neuroligin-1 in promoting synaptic differentiation, and, most importantly, these two families of neurexin-binding partners cooperate in an additive or synergistic manner. -

WO 2019/079361 Al 25 April 2019 (25.04.2019) W 1P O PCT
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization I International Bureau (10) International Publication Number (43) International Publication Date WO 2019/079361 Al 25 April 2019 (25.04.2019) W 1P O PCT (51) International Patent Classification: CA, CH, CL, CN, CO, CR, CU, CZ, DE, DJ, DK, DM, DO, C12Q 1/68 (2018.01) A61P 31/18 (2006.01) DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, HN, C12Q 1/70 (2006.01) HR, HU, ID, IL, IN, IR, IS, JO, JP, KE, KG, KH, KN, KP, KR, KW, KZ, LA, LC, LK, LR, LS, LU, LY, MA, MD, ME, (21) International Application Number: MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, PCT/US2018/056167 OM, PA, PE, PG, PH, PL, PT, QA, RO, RS, RU, RW, SA, (22) International Filing Date: SC, SD, SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, 16 October 2018 (16. 10.2018) TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. (25) Filing Language: English (84) Designated States (unless otherwise indicated, for every kind of regional protection available): ARIPO (BW, GH, (26) Publication Language: English GM, KE, LR, LS, MW, MZ, NA, RW, SD, SL, ST, SZ, TZ, (30) Priority Data: UG, ZM, ZW), Eurasian (AM, AZ, BY, KG, KZ, RU, TJ, 62/573,025 16 October 2017 (16. 10.2017) US TM), European (AL, AT, BE, BG, CH, CY, CZ, DE, DK, EE, ES, FI, FR, GB, GR, HR, HU, ΓΕ , IS, IT, LT, LU, LV, (71) Applicant: MASSACHUSETTS INSTITUTE OF MC, MK, MT, NL, NO, PL, PT, RO, RS, SE, SI, SK, SM, TECHNOLOGY [US/US]; 77 Massachusetts Avenue, TR), OAPI (BF, BJ, CF, CG, CI, CM, GA, GN, GQ, GW, Cambridge, Massachusetts 02139 (US). -

Molecular Effects of Isoflavone Supplementation Human Intervention Studies and Quantitative Models for Risk Assessment
Molecular effects of isoflavone supplementation Human intervention studies and quantitative models for risk assessment Vera van der Velpen Thesis committee Promotors Prof. Dr Pieter van ‘t Veer Professor of Nutritional Epidemiology Wageningen University Prof. Dr Evert G. Schouten Emeritus Professor of Epidemiology and Prevention Wageningen University Co-promotors Dr Anouk Geelen Assistant professor, Division of Human Nutrition Wageningen University Dr Lydia A. Afman Assistant professor, Division of Human Nutrition Wageningen University Other members Prof. Dr Jaap Keijer, Wageningen University Dr Hubert P.J.M. Noteborn, Netherlands Food en Consumer Product Safety Authority Prof. Dr Yvonne T. van der Schouw, UMC Utrecht Dr Wendy L. Hall, King’s College London This research was conducted under the auspices of the Graduate School VLAG (Advanced studies in Food Technology, Agrobiotechnology, Nutrition and Health Sciences). Molecular effects of isoflavone supplementation Human intervention studies and quantitative models for risk assessment Vera van der Velpen Thesis submitted in fulfilment of the requirements for the degree of doctor at Wageningen University by the authority of the Rector Magnificus Prof. Dr M.J. Kropff, in the presence of the Thesis Committee appointed by the Academic Board to be defended in public on Friday 20 June 2014 at 13.30 p.m. in the Aula. Vera van der Velpen Molecular effects of isoflavone supplementation: Human intervention studies and quantitative models for risk assessment 154 pages PhD thesis, Wageningen University, Wageningen, NL (2014) With references, with summaries in Dutch and English ISBN: 978-94-6173-952-0 ABSTRact Background: Risk assessment can potentially be improved by closely linked experiments in the disciplines of epidemiology and toxicology. -

Environmental and Genetic Factors in Autism Spectrum Disorders: Special Emphasis on Data from Arabian Studies
International Journal of Environmental Research and Public Health Review Environmental and Genetic Factors in Autism Spectrum Disorders: Special Emphasis on Data from Arabian Studies Noor B. Almandil 1,† , Deem N. Alkuroud 2,†, Sayed AbdulAzeez 2, Abdulla AlSulaiman 3, Abdelhamid Elaissari 4 and J. Francis Borgio 2,* 1 Department of Clinical Pharmacy Research, Institute for Research and Medical Consultation (IRMC), Imam Abdulrahman Bin Faisal University, Dammam 31441, Saudi Arabia; [email protected] 2 Department of Genetic Research, Institute for Research and Medical Consultation (IRMC), Imam Abdulrahman Bin Faisal University, Dammam 31441, Saudi Arabia; [email protected] (D.N.A.); [email protected] (S.A.) 3 Department of Neurology, College of Medicine, Imam Abdulrahman Bin Faisal University, Dammam 31441, Saudi Arabia; [email protected] or [email protected] 4 Univ Lyon, University Claude Bernard Lyon-1, CNRS, LAGEP-UMR 5007, F-69622 Lyon, France; [email protected] * Correspondence: [email protected] or [email protected]; Tel.: +966-13-333-0864 † These authors contributed equally to this work. Received: 26 January 2019; Accepted: 19 February 2019; Published: 23 February 2019 Abstract: One of the most common neurodevelopmental disorders worldwide is autism spectrum disorder (ASD), which is characterized by language delay, impaired communication interactions, and repetitive patterns of behavior caused by environmental and genetic factors. This review aims to provide a comprehensive survey of recently published literature on ASD and especially novel insights into excitatory synaptic transmission. Even though numerous genes have been discovered that play roles in ASD, a good understanding of the pathophysiologic process of ASD is still lacking. -

Slitrks Control Excitatory and Inhibitory Synapse Formation with LAR
Slitrks control excitatory and inhibitory synapse SEE COMMENTARY formation with LAR receptor protein tyrosine phosphatases Yeong Shin Yima,1, Younghee Kwonb,1, Jungyong Namc, Hong In Yoona, Kangduk Leeb, Dong Goo Kima, Eunjoon Kimc, Chul Hoon Kima,2, and Jaewon Kob,2 aDepartment of Pharmacology, Brain Research Institute, Brain Korea 21 Project for Medical Science, Severance Biomedical Science Institute, Yonsei University College of Medicine, Seoul 120-752, Korea; bDepartment of Biochemistry, College of Life Science and Biotechnology, Yonsei University, Seoul 120-749, Korea; and cCenter for Synaptic Brain Dysfunctions, Institute for Basic Science, Department of Biological Sciences, Korea Advanced Institute of Science and Technology, Daejeon 305-701, Korea Edited by Thomas C. Südhof, Stanford University School of Medicine, Stanford, CA, and approved December 26, 2012 (received for review June 11, 2012) The balance between excitatory and inhibitory synaptic inputs, share a similar domain organization comprising three Ig domains which is governed by multiple synapse organizers, controls neural and four to eight fibronectin type III repeats. LAR-RPTP family circuit functions and behaviors. Slit- and Trk-like proteins (Slitrks) are members are evolutionarily conserved and are functionally required a family of synapse organizers, whose emerging synaptic roles are for axon guidance and synapse formation (15). Recent studies have incompletely understood. Here, we report that Slitrks are enriched shown that netrin-G ligand-3 (NGL-3), neurotrophin receptor ty- in postsynaptic densities in rat brains. Overexpression of Slitrks rosine kinase C (TrkC), and IL-1 receptor accessory protein-like 1 promoted synapse formation, whereas RNAi-mediated knock- (IL1RAPL1) bind to all three LAR-RPTP family members or dis- down of Slitrks decreased synapse density. -

Deletion of Α-Neurexin II Results in Autism-Related
OPEN Citation: Transl Psychiatry (2014) 4, e484; doi:10.1038/tp.2014.123 www.nature.com/tp ORIGINAL ARTICLE Deletion of α-neurexin II results in autism-related behaviors in mice J Dachtler1, J Glasper2, RN Cohen1, JL Ivorra1, DJ Swiffen1, AJ Jackson1, MK Harte2, RJ Rodgers3 and SJ Clapcote1 Autism is a common and frequently disabling neurodevelopmental disorder with a strong genetic basis. Human genetic studies have discovered mutations disrupting exons of the NRXN2 gene, which encodes the synaptic adhesion protein α-neurexin II (Nrxn2α), in two unrelated individuals with autism, but a causal link between NRXN2 and the disorder remains unclear. To begin to test the hypothesis that Nrxn2α deficiency contributes to the symptoms of autism, we employed Nrxn2α knockout (KO) mice that genetically model Nrxn2α deficiency in vivo. We report that Nrxn2α KO mice displayed deficits in sociability and social memory when exposed to novel conspecifics. In tests of exploratory activity, Nrxn2α KO mice displayed an anxiety-like phenotype in comparison with wild-type littermates, with thigmotaxis in an open field, less time spent in the open arms of an elevated plus maze, more time spent in the enclosure of an emergence test and less time spent exploring novel objects. However, Nrxn2α KO mice did not exhibit any obvious changes in prepulse inhibition or in passive avoidance learning. Real-time PCR analysis of the frontal cortex and hippocampus revealed significant decreases in the mRNA levels of genes encoding proteins involved in both excitatory and inhibitory transmission. Quantification of protein expression revealed that Munc18-1, encoded by Stxbp1, was significantly decreased in the hippocampus of Nrxn2α KO mice, which is suggestive of deficiencies in presynaptic vesicular release.