Integrated Analysis of Gene Expression Changes Associated
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Characterization of TRIB2-Mediated Resistance to Pharmacological Inhibition of MEK
VANESSA MENDES HENRIQUES Characterization of TRIB2-mediated resistance to pharmacological inhibition of MEK Oncobiology Master Thesis Faro, 2017 VANESSA MENDES HENRIQUES Characterization of TRIB2-mediated resistance to pharmacological inhibition of MEK Supervisors: Dr. Wolfgang Link Dr. Bibiana Ferreira Oncobiology Master Thesis Faro, 2017 Título: “Characterization of TRIB2-mediated resistance to pharmacological inhibition of MEK” Declaração de autoria do trabalho Declaro ser a autora deste trabalho, que é original e inédito. Autores e trabalhos consultados estão devidamente citados no texto e constam da listagem de referências incluída. Copyright Vanessa Mendes Henriques _____________________________ A Universidade do Algarve tem o direito, perpétuo e sem limites geográficos, de arquivar e publicitar este trabalho através de exemplares impressos reproduzidos em papel ou de forma digital, ou por qualquer outro meio conhecido ou que venha a ser inventado, de o divulgar através de repositórios científicos e de admitir a sua cópia e distribuição com objetivos educacionais ou de investigação, não comerciais, desde que seja dado crédito ao autor e editor. i Acknowledgements First, I would like to thank the greatest opportunity given by professor doctor Wolfgang Link to accepting me into his team, contributing to my scientific and personal progress. Thank You for all Your knowledge and help across the year. A special thanks to Bibiana Ferreira who stayed by me all year and trained me. Thank You for all you taught me, thank you for all your patience and time and all the support. Thank you for all the great times that You provided me. It was an amazing experience to learn and work with You, which made me grow as a scientist and also as a person. -
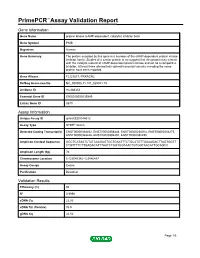
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name protein kinase (cAMP-dependent, catalytic) inhibitor beta Gene Symbol PKIB Organism Human Gene Summary The protein encoded by this gene is a member of the cAMP-dependent protein kinase inhibitor family. Studies of a similar protein in rat suggest that this protein may interact with the catalytic subunit of cAMP-dependent protein kinase and act as a competitive inhibitor. At least three alternatively spliced transcript variants encoding the same protein have been reported. Gene Aliases FLJ23817, PRKACN2 RefSeq Accession No. NC_000006.11, NT_025741.15 UniGene ID Hs.486354 Ensembl Gene ID ENSG00000135549 Entrez Gene ID 5570 Assay Information Unique Assay ID qHsaCED0044612 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000368452, ENST00000368448, ENST00000258014, ENST00000354275, ENST00000368446, ENST00000392491, ENST00000392490 Amplicon Context Sequence GGCTCATAATCTATCAAGAGTGCTGAATTTCTGCATGTTGAAAGACTTAGTGGTT CTGTTTTCTTGAGACATTTAATCTGGTGGTAACTGTGGTAACATTGCAGCC Amplicon Length (bp) 76 Chromosome Location 6:123046342-123046447 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 95 R2 0.9996 cDNA Cq 22.83 cDNA Tm (Celsius) 76.5 gDNA Cq 24.52 Page 1/5 PrimePCR™Assay Validation Report Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report PKIB, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve -

Location Analysis of Estrogen Receptor Target Promoters Reveals That
Location analysis of estrogen receptor ␣ target promoters reveals that FOXA1 defines a domain of the estrogen response Jose´ e Laganie` re*†, Genevie` ve Deblois*, Ce´ line Lefebvre*, Alain R. Bataille‡, Franc¸ois Robert‡, and Vincent Gigue` re*†§ *Molecular Oncology Group, Departments of Medicine and Oncology, McGill University Health Centre, Montreal, QC, Canada H3A 1A1; †Department of Biochemistry, McGill University, Montreal, QC, Canada H3G 1Y6; and ‡Laboratory of Chromatin and Genomic Expression, Institut de Recherches Cliniques de Montre´al, Montreal, QC, Canada H2W 1R7 Communicated by Ronald M. Evans, The Salk Institute for Biological Studies, La Jolla, CA, July 1, 2005 (received for review June 3, 2005) Nuclear receptors can activate diverse biological pathways within general absence of large scale functional data linking these putative a target cell in response to their cognate ligands, but how this binding sites with gene expression in specific cell types. compartmentalization is achieved at the level of gene regulation is Recently, chromatin immunoprecipitation (ChIP) has been used poorly understood. We used a genome-wide analysis of promoter in combination with promoter or genomic DNA microarrays to occupancy by the estrogen receptor ␣ (ER␣) in MCF-7 cells to identify loci recognized by transcription factors in a genome-wide investigate the molecular mechanisms underlying the action of manner in mammalian cells (20–24). This technology, termed 17-estradiol (E2) in controlling the growth of breast cancer cells. ChIP-on-chip or location analysis, can therefore be used to deter- We identified 153 promoters bound by ER␣ in the presence of E2. mine the global gene expression program that characterize the Motif-finding algorithms demonstrated that the estrogen re- action of a nuclear receptor in response to its natural ligand. -

Investigation of the Underlying Hub Genes and Molexular Pathogensis in Gastric Cancer by Integrated Bioinformatic Analyses
bioRxiv preprint doi: https://doi.org/10.1101/2020.12.20.423656; this version posted December 22, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Investigation of the underlying hub genes and molexular pathogensis in gastric cancer by integrated bioinformatic analyses Basavaraj Vastrad1, Chanabasayya Vastrad*2 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karanataka, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India bioRxiv preprint doi: https://doi.org/10.1101/2020.12.20.423656; this version posted December 22, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract The high mortality rate of gastric cancer (GC) is in part due to the absence of initial disclosure of its biomarkers. The recognition of important genes associated in GC is therefore recommended to advance clinical prognosis, diagnosis and and treatment outcomes. The current investigation used the microarray dataset GSE113255 RNA seq data from the Gene Expression Omnibus database to diagnose differentially expressed genes (DEGs). Pathway and gene ontology enrichment analyses were performed, and a proteinprotein interaction network, modules, target genes - miRNA regulatory network and target genes - TF regulatory network were constructed and analyzed. Finally, validation of hub genes was performed. The 1008 DEGs identified consisted of 505 up regulated genes and 503 down regulated genes. -

Molecular Effects of Isoflavone Supplementation Human Intervention Studies and Quantitative Models for Risk Assessment
Molecular effects of isoflavone supplementation Human intervention studies and quantitative models for risk assessment Vera van der Velpen Thesis committee Promotors Prof. Dr Pieter van ‘t Veer Professor of Nutritional Epidemiology Wageningen University Prof. Dr Evert G. Schouten Emeritus Professor of Epidemiology and Prevention Wageningen University Co-promotors Dr Anouk Geelen Assistant professor, Division of Human Nutrition Wageningen University Dr Lydia A. Afman Assistant professor, Division of Human Nutrition Wageningen University Other members Prof. Dr Jaap Keijer, Wageningen University Dr Hubert P.J.M. Noteborn, Netherlands Food en Consumer Product Safety Authority Prof. Dr Yvonne T. van der Schouw, UMC Utrecht Dr Wendy L. Hall, King’s College London This research was conducted under the auspices of the Graduate School VLAG (Advanced studies in Food Technology, Agrobiotechnology, Nutrition and Health Sciences). Molecular effects of isoflavone supplementation Human intervention studies and quantitative models for risk assessment Vera van der Velpen Thesis submitted in fulfilment of the requirements for the degree of doctor at Wageningen University by the authority of the Rector Magnificus Prof. Dr M.J. Kropff, in the presence of the Thesis Committee appointed by the Academic Board to be defended in public on Friday 20 June 2014 at 13.30 p.m. in the Aula. Vera van der Velpen Molecular effects of isoflavone supplementation: Human intervention studies and quantitative models for risk assessment 154 pages PhD thesis, Wageningen University, Wageningen, NL (2014) With references, with summaries in Dutch and English ISBN: 978-94-6173-952-0 ABSTRact Background: Risk assessment can potentially be improved by closely linked experiments in the disciplines of epidemiology and toxicology. -

Genome-Wide Association Study of Diabetic Kidney Disease Highlights Biology Involved in Glomerular Basement Membrane Collagen
CLINICAL RESEARCH www.jasn.org Genome-Wide Association Study of Diabetic Kidney Disease Highlights Biology Involved in Glomerular Basement Membrane Collagen Rany M. Salem ,1 Jennifer N. Todd,2,3,4 Niina Sandholm ,5,6,7 Joanne B. Cole ,2,3,4 Wei-Min Chen,8 Darrell Andrews,9 Marcus G. Pezzolesi,10 Paul M. McKeigue,11 Linda T. Hiraki,12 Chengxiang Qiu,13 Viji Nair,14 Chen Di Liao,12 Jing Jing Cao,12 Erkka Valo ,5,6,7 Suna Onengut-Gumuscu,8 Adam M. Smiles,15 Stuart J. McGurnaghan,16 Jani K. Haukka,5,6,7 Valma Harjutsalo,5,6,7,17 Eoin P. Brennan,9 Natalie van Zuydam,18,19 Emma Ahlqvist,20 Ross Doyle,9 Tarunveer S. Ahluwalia ,21 Maria Lajer,21 Maria F. Hughes,9 Jihwan Park,13 Jan Skupien,15 Athina Spiliopoulou,11 Andrew Liu,22 Rajasree Menon,14,23 Carine M. Boustany-Kari,24 Hyun M. Kang,23,25 Robert G. Nelson,26 Ronald Klein,27 Barbara E. Klein,27 Kristine E. Lee ,27 Xiaoyu Gao,28 Michael Mauer,29 Silvia Maestroni,30 Maria Luiza Caramori,29 Ian H. de Boer ,31 Rachel G. Miller,32 Jingchuan Guo ,32 Andrew P. Boright,12 David Tregouet,33,34 Beata Gyorgy,33,34 Janet K. Snell-Bergeon,35 David M. Maahs,36 Shelley B. Bull ,37 Angelo J. Canty,38 Colin N.A. Palmer,39 Lars Stechemesser,40 Bernhard Paulweber,40 Raimund Weitgasser,40,41 Jelizaveta Sokolovska,42 Vita Rovıte,43 Valdis Pırags, 42,44 Edita Prakapiene,45 Lina Radzeviciene,46 Rasa Verkauskiene,46 Nicolae Mircea Panduru,6,47 Leif C. -

140503 IPF Signatures Supplement Withfigs Thorax
Supplementary material for Heterogeneous gene expression signatures correspond to distinct lung pathologies and biomarkers of disease severity in idiopathic pulmonary fibrosis Daryle J. DePianto1*, Sanjay Chandriani1⌘*, Alexander R. Abbas1, Guiquan Jia1, Elsa N. N’Diaye1, Patrick Caplazi1, Steven E. Kauder1, Sabyasachi Biswas1, Satyajit K. Karnik1#, Connie Ha1, Zora Modrusan1, Michael A. Matthay2, Jasleen Kukreja3, Harold R. Collard2, Jackson G. Egen1, Paul J. Wolters2§, and Joseph R. Arron1§ 1Genentech Research and Early Development, South San Francisco, CA 2Department of Medicine, University of California, San Francisco, CA 3Department of Surgery, University of California, San Francisco, CA ⌘Current address: Novartis Institutes for Biomedical Research, Emeryville, CA. #Current address: Gilead Sciences, Foster City, CA. *DJD and SC contributed equally to this manuscript §PJW and JRA co-directed this project Address correspondence to Paul J. Wolters, MD University of California, San Francisco Department of Medicine Box 0111 San Francisco, CA 94143-0111 [email protected] or Joseph R. Arron, MD, PhD Genentech, Inc. MS 231C 1 DNA Way South San Francisco, CA 94080 [email protected] 1 METHODS Human lung tissue samples Tissues were obtained at UCSF from clinical samples from IPF patients at the time of biopsy or lung transplantation. All patients were seen at UCSF and the diagnosis of IPF was established through multidisciplinary review of clinical, radiological, and pathological data according to criteria established by the consensus classification of the American Thoracic Society (ATS) and European Respiratory Society (ERS), Japanese Respiratory Society (JRS), and the Latin American Thoracic Association (ALAT) (ref. 5 in main text). Non-diseased normal lung tissues were procured from lungs not used by the Northern California Transplant Donor Network. -

75 2. INTRODUCTION Triple-Negative Breast Cancer (TNBC)
[Frontiers in Bioscience, Scholar, 11, 75-88, March 1, 2019] The persisting puzzle of racial disparity in triple negative breast cancer: looking through a new lens Chakravarthy Garlapati1, Shriya Joshi1, Bikram Sahoo1, Shobhna Kapoor2, Ritu Aneja1 1Department of Biology, Georgia State University, Atlanta, GA, USA, 2Department of Chemistry, Indian Institute of Technology Bombay, Powai, India TABLE OF CONTENTS 1. Abstract 2. Introduction 3. Dissecting the TNBC racially disparate burden 3.1. Does race influence TNBC onset and progression? 3.2. Tumor microenvironment in TNBC and racial disparity 3.3. Differential gene signatures and pathways in racially distinct TNBC 3.4. Our Perspective: Looking racial disparity through a new lens 4. Conclusion 5. Acknowledgement 6. References 1. ABSTRACT 2. INTRODUCTION Triple-negative breast cancer (TNBC) Triple-negative breast cancer (TNBC), is characterized by the absence of estrogen a subtype of breast cancer (BC), accounts for and progesterone receptors and absence 15-20% of all BC diagnoses in the US. It has of amplification of human epidermal growth been recognized that women of African descent factor receptor (HER2). This disease has no are twice as likely to develop TNBC than approved treatment with a poor prognosis women of European descent (1). As the name particularly in African-American (AA) as foretells, TNBCs lack estrogen, progesterone, compared to European-American (EA) and human epidermal growth factor receptors. patients. Gene ontology analysis showed Unfortunately, TNBCs are defined by what they specific gene pathways that are differentially “lack” rather than what they “have” and thus this regulated and gene signatures that are negative nomenclature provides no actionable differentially expressed in AA as compared to information on “druggable” targets. -

Inhibition of Mitochondrial Complex II in Neuronal Cells Triggers Unique
www.nature.com/scientificreports OPEN Inhibition of mitochondrial complex II in neuronal cells triggers unique pathways culminating in autophagy with implications for neurodegeneration Sathyanarayanan Ranganayaki1, Neema Jamshidi2, Mohamad Aiyaz3, Santhosh‑Kumar Rashmi4, Narayanappa Gayathri4, Pulleri Kandi Harsha5, Balasundaram Padmanabhan6 & Muchukunte Mukunda Srinivas Bharath7* Mitochondrial dysfunction and neurodegeneration underlie movement disorders such as Parkinson’s disease, Huntington’s disease and Manganism among others. As a corollary, inhibition of mitochondrial complex I (CI) and complex II (CII) by toxins 1‑methyl‑4‑phenylpyridinium (MPP+) and 3‑nitropropionic acid (3‑NPA) respectively, induced degenerative changes noted in such neurodegenerative diseases. We aimed to unravel the down‑stream pathways associated with CII inhibition and compared with CI inhibition and the Manganese (Mn) neurotoxicity. Genome‑wide transcriptomics of N27 neuronal cells exposed to 3‑NPA, compared with MPP+ and Mn revealed varied transcriptomic profle. Along with mitochondrial and synaptic pathways, Autophagy was the predominant pathway diferentially regulated in the 3‑NPA model with implications for neuronal survival. This pathway was unique to 3‑NPA, as substantiated by in silico modelling of the three toxins. Morphological and biochemical validation of autophagy markers in the cell model of 3‑NPA revealed incomplete autophagy mediated by mechanistic Target of Rapamycin Complex 2 (mTORC2) pathway. Interestingly, Brain Derived Neurotrophic Factor -

The New Antitumor Drug ABTL0812 Inhibits the Akt/Mtorc1 Axis By
Published OnlineFirst December 15, 2015; DOI: 10.1158/1078-0432.CCR-15-1808 Cancer Therapy: Preclinical Clinical Cancer Research The New Antitumor Drug ABTL0812 Inhibits the Akt/mTORC1 Axis by Upregulating Tribbles-3 Pseudokinase Tatiana Erazo1, Mar Lorente2,3, Anna Lopez-Plana 4, Pau Munoz-Guardiola~ 1,5, Patricia Fernandez-Nogueira 4, Jose A. García-Martínez5, Paloma Bragado4, Gemma Fuster4, María Salazar2, Jordi Espadaler5, Javier Hernandez-Losa 6, Jose Ramon Bayascas7, Marc Cortal5, Laura Vidal8, Pedro Gascon 4,8, Mariana Gomez-Ferreria 5, Jose Alfon 5, Guillermo Velasco2,3, Carles Domenech 5, and Jose M. Lizcano1 Abstract Purpose: ABTL0812 is a novel first-in-class, small molecule tion of Tribbles-3 pseudokinase (TRIB3) gene expression. Upre- which showed antiproliferative effect on tumor cells in pheno- gulated TRIB3 binds cellular Akt, preventing its activation by typic assays. Here we describe the mechanism of action of this upstream kinases, resulting in Akt inhibition and suppression of antitumor drug, which is currently in clinical development. the Akt/mTORC1 axis. Pharmacologic inhibition of PPARa/g or Experimental Design: We investigated the effect of ABTL0812 TRIB3 silencing prevented ABTL0812-induced cell death. on cancer cell death, proliferation, and modulation of intracel- ABTL0812 treatment induced Akt inhibition in cancer cells, tumor lular signaling pathways, using human lung (A549) and pancre- xenografts, and peripheral blood mononuclear cells from patients atic (MiaPaCa-2) cancer cells and tumor xenografts. To identify enrolled in phase I/Ib first-in-human clinical trial. cellular targets, we performed in silico high-throughput screening Conclusions: ABTL0812 has a unique and novel mechanism of comparing ABTL0812 chemical structure against ChEMBL15 action, that defines a new and drugable cellular route that links database. -

Genome-Wide Association and Trans-Ethnic Meta-Analysis for Advanced Diabetic Kidney Disease: Family Investigation of Nephropathy and Diabetes (FIND)
RESEARCH ARTICLE Genome-Wide Association and Trans-ethnic Meta-Analysis for Advanced Diabetic Kidney Disease: Family Investigation of Nephropathy and Diabetes (FIND) Sudha K. Iyengar1☯*, John R. Sedor2,3☯*, Barry I. Freedman4☯*, W. H. Linda Kao5†, Matthias Kretzler6, Benjamin J. Keller6, Hanna E. Abboud7†, Sharon G. Adler8, Lyle G. Best9, Donald W. Bowden10, Allison Burlock6, Yii-Der Ida Chen11, Shelley A. Cole12, Mary E. Comeau13, Jeffrey M. Curtis14, Jasmin Divers13, Christiane Drechsler15, Ravi Duggirala12, Robert C. Elston1, Xiuqing Guo11, Huateng Huang16, Michael Marcus Hoffmann17, Barbara V. Howard18, Eli Ipp19, Paul L. Kimmel20, Michael J. Klag21, William C. Knowler14, Orly F. Kohn22, Tennille S. Leak6, David J. Leehey23, Man Li24, Alka Malhotra14, Winfried März25, Viji Nair6, Robert G. Nelson14, Susanne B. Nicholas26, Stephen J. O’Brien27, Madeleine V. Pahl28, Rulan S. Parekh29, Marcus G. Pezzolesi30, Rebekah S. Rasooly31, Charles N. Rotimi32, Jerome I. Rotter11, Jeffrey R. Schelling2, Michael F. Seldin33, Vallabh O. Shah34, Adam M. Smiles35, Michael W. Smith36, Kent D. Taylor11, Farook Thameem37¤, Denyse P. Thornley-Brown38, Barbara J. Truitt1, OPEN ACCESS Christoph Wanner39, E. Jennifer Weil14, Cheryl A. Winkler40, Philip G. Zager41, Robert P. Igo, Jr1‡, Robert L. Hanson14‡, Carl D. Langefeld11‡, Family Investigation of Citation: Iyengar SK, Sedor JR, Freedman BI, Kao Nephropathy and Diabetes (FIND)¶ WHL, Kretzler M, Keller BJ, et al. (2015) Genome- Wide Association and Trans-ethnic Meta-Analysis for 1 Department of Epidemiology & Biostatistics, Case Western Reserve University, Cleveland, Ohio, Advanced Diabetic Kidney Disease: Family United States of America, 2 Departments of Medicine, Case Western Reserve University, Cleveland, Ohio, Investigation of Nephropathy and Diabetes (FIND). -

Chemical Agent and Antibodies B-Raf Inhibitor RAF265
Supplemental Materials and Methods: Chemical agent and antibodies B-Raf inhibitor RAF265 [5-(2-(5-(trifluromethyl)-1H-imidazol-2-yl)pyridin-4-yloxy)-N-(4-trifluoromethyl)phenyl-1-methyl-1H-benzp{D, }imidazol-2- amine] was kindly provided by Novartis Pharma AG and dissolved in solvent ethanol:propylene glycol:2.5% tween-80 (percentage 6:23:71) for oral delivery to mice by gavage. Antibodies to phospho-ERK1/2 Thr202/Tyr204(4370), phosphoMEK1/2(2338 and 9121)), phospho-cyclin D1(3300), cyclin D1 (2978), PLK1 (4513) BIM (2933), BAX (2772), BCL2 (2876) were from Cell Signaling Technology. Additional antibodies for phospho-ERK1,2 detection for western blot were from Promega (V803A), and Santa Cruz (E-Y, SC7383). Total ERK antibody for western blot analysis was K-23 from Santa Cruz (SC-94). Ki67 antibody (ab833) was from ABCAM, Mcl1 antibody (559027) was from BD Biosciences, Factor VIII antibody was from Dako (A082), CD31 antibody was from Dianova, (DIA310), and Cot antibody was from Santa Cruz Biotechnology (sc-373677). For the cyclin D1 second antibody staining was with an Alexa Fluor 568 donkey anti-rabbit IgG (Invitrogen, A10042) (1:200 dilution). The pMEK1 fluorescence was developed using the Alexa Fluor 488 chicken anti-rabbit IgG second antibody (1:200 dilution).TUNEL staining kits were from Promega (G2350). Mouse Implant Studies: Biopsy tissues were delivered to research laboratory in ice-cold Dulbecco's Modified Eagle Medium (DMEM) buffer solution. As the tissue mass available from each biopsy was limited, we first passaged the biopsy tissue in Balb/c nu/Foxn1 athymic nude mice (6-8 weeks of age and weighing 22-25g, purchased from Harlan Sprague Dawley, USA) to increase the volume of tumor for further implantation.