Noonan Spectrum Disorders Panel, Sequencing ARUP Test Code 2010772 Noonan Disorders Sequencing Specimen Whole Blood
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
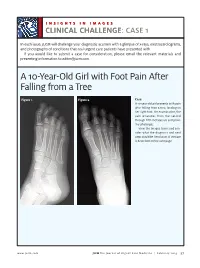
A 10-Year-Old Girl with Foot Pain After Falling from a Tree
INSIGHTS IN IMAGES CLINICAL CHALLENGECHALLENGE: CASE 1 In each issue, JUCM will challenge your diagnostic acumen with a glimpse of x-rays, electrocardiograms, and photographs of conditions that real urgent care patients have presented with. If you would like to submit a case for consideration, please email the relevant materials and presenting information to [email protected]. A 10-Year-Old Girl with Foot Pain After Falling from a Tree Figure 1. Figure 2. Case A 10-year-old girl presents with pain after falling from a tree, landing on her right foot. On examination, the pain emanates from the second through fifth metatarsals and proxi- mal phalanges. View the images taken and con- sider what the diagnosis and next steps would be. Resolution of the case is described on the next page. www.jucm.com JUCM The Journal of Urgent Care Medicine | February 2019 37 INSIGHTS IN IMAGES: CLINICAL CHALLENGE THE RESOLUTION Figure 1. Mediastinal air Figure 1. Differential Diagnosis Pearls for Urgent Care Management and Ⅲ Fracture of the distal fourth metatarsal Considerations for Transfer Ⅲ Plantar plate disruption Ⅲ Emergent transfer should be considered with associated neu- Ⅲ Sesamoiditis rologic deficit, compartment syndrome, open fracture, or vas- Ⅲ Turf toe cular compromise Ⅲ Referral to an orthopedist is warranted in the case of an in- Diagnosis tra-articular fracture, or with Lisfranc ligament injury or ten- Angulation of the distal fourth metatarsal metaphyseal cortex derness over the Lisfranc ligament and hairline lucency consistent with fracture. Acknowledgment: Images courtesy of Teleradiology Associates. Learnings/What to Look for Ⅲ Proximal metatarsal fractures are most often caused by crush- ing or direct blows Ⅲ In athletes, an axial load placed on a plantar-flexed foot should raise suspicion of a Lisfranc injury 38 JUCM The Journal of Urgent Care Medicine | February 2019 www.jucm.com INSIGHTS IN IMAGES CLINICAL CHALLENGE: CASE 2 A 55-Year-Old Man with 3 Hours of Epigastric Pain 55 years PR 249 QRSD 90 QT 471 QTc 425 AXES P 64 QRS -35 T 30 Figure 1. -

Genetic Mechanisms of Disease in Children: a New Look
Genetic Mechanisms of Disease in Children: A New Look Laurie Demmer, MD Tufts Medical Center and the Floating Hospital for Children Boston, MA Traditionally genetic disorders have been linked to the ‘one gene-one protein-one disease’ hypothesis. However recent advances in the field of molecular biology and biotechnology have afforded us the opportunity to greatly expand our knowledge of genetics, and we now know that the mechanisms of inherited disorders are often significantly more complex, and consequently, much more intriguing, than originally thought. Classical mendelian disorders with relatively simple genetic mechanisms do exist, but turn out to be far more rare than originally thought. All patients with sickle cell disease for example, carry the same A-to-T point mutation in the sixth codon of the beta globin gene. This results in a glutamate to valine substitution which changes the shape and the function of the globin molecule in a predictable way. Similarly, all patients with achondroplasia have a single base pair substitution at nucleotide #1138 of the FGFR3 gene. On the other hand, another common inherited disorder, cystic fibrosis, is known to result from changes in a specific transmembrane receptor (CFTR), but over 1000 different disease-causing mutations have been reported in this single gene. Since most commercial labs only test for between 23-100 different mutations, interpreting CFTR mutation testing is significantly complicated by the known risk of false negative results. Many examples of complex, or non-Mendelian, inheritance are now known to exist and include disorders of trinucleotide repeats, errors in imprinting, and gene dosage effects. -

Blueprint Genetics Hereditary Leukemia Panel
Hereditary Leukemia Panel Test code: ON0101 Is a 41 gene panel that includes assessment of non-coding variants. Is ideal for patients with a personal history of a syndrome that confers an increased risk of leukemia or patients with a family history of a syndrome that confers an increased risk of leukemia. About Hereditary Leukemia An inherited predisposition to hematological malignancies, namely acute lymphoblastic leukemia (ALL), acute myeloid leukemia (AML), and bone marrow myelodysplastic syndrome (MDS) may be associated with syndromic features or occur as the principal clinical feature. MDSs and AMLs can occur in the context of syndromic bone marrow failure (eg. dyskeratosis congenita, Fanconi anemia). Other hereditary syndromes with an increased risk of leukemia include Li-Fraumeni syndrome (TP53), ataxia telangiectasia (ATM), Bloom syndrome (BLM), neurofibromatosis type 1 (NF1) and less frequently Noonan syndrome (PTPN11, CBL). Some reports have also shown an association of biallelic germline mutations in constitutional mismatch repair-deficiency syndrome genes, MLH1, MSH2, MSH6, and PMS2 with the development of ALL. Isolated hematological malignancies are associated with germline mutations in RUNX1 (familial platelet syndrome with predisposition to acute myelogenous leukemia), CEBPA (familial AML), GATA2 (GATA2-associated syndromes) and DDX41(DDX41 -related myeloid neoplasms). There is a rapidly expanding list of germline mutations associated with increased risks for myeloid malignancies and inherited predisposition to hematologic malignancies may be more common than has been thought. Many different genetic defects associated with the development of leukemia have been described but the common underlying mechanism is a dysfunctional DNA damage response. Recognition of an inherited cause provides a specific molecular diagnosis and helps to guide treatment, understand unique disease features, prognosis and other organ systems that may be involved, and identify others in the family who may be at risk. -

Segmental Overvekst Og Vaskulærmalformasjoner V02
2/1/2021 Segmental overvekst og vaskulærmalformasjoner v02 Avdeling for medisinsk genetikk Segmental overvekst og vaskulærmalformasjoner Genpanel, versjon v02 * Enkelte genomiske regioner har lav eller ingen sekvensdekning ved eksomsekvensering. Dette skyldes at de har stor likhet med andre områder i genomet, slik at spesifikk gjenkjennelse av disse områdene og påvisning av varianter i disse områdene, blir vanskelig og upålitelig. Disse genetiske regionene har vi identifisert ved å benytte USCS segmental duplication hvor områder større enn 1 kb og ≥90% likhet med andre regioner i genomet, gjenkjennes (https://genome.ucsc.edu). Vi gjør oppmerksom på at ved identifiseringav ekson oppstrøms for startkodon kan eksonnummereringen endres uten at transkript ID endres. Avdelingens websider har en full oversikt over områder som er affisert av segmentale duplikasjoner. ** Transkriptets kodende ekson. Ekson Gen Gen affisert (HGNC (HGNC Transkript Ekson** Fenotype av symbol) ID) segdup* ACVRL1 175 NM_000020.3 2-10 Telangiectasia, hereditary hemorrhagic, type 2 OMIM ADAMTS3 219 NM_014243.3 1-22 Hennekam lymphangiectasia- lymphedema syndrome 3 OMIM AKT1 391 NM_005163.2 2-14 Cowden syndrome 6 OMIM Proteus syndrome, somatic OMIM AKT2 392 NM_001626.6 2-14 Diabetes mellitus, type II OMIM Hypoinsulinemic hypoglycemia with hemihypertrophy OMIM AKT3 393 NM_005465.7 2-14 Megalencephaly-polymicrogyria- polydactyly-hydrocephalus syndrome 2 OMIM file:///data/SegOv_v02-web.html 1/7 2/1/2021 Segmental overvekst og vaskulærmalformasjoner v02 Ekson Gen Gen affisert (HGNC (HGNC -

Exostoses, Enchondromatosis and Metachondromatosis; Diagnosis and Management
Acta Orthop. Belg., 2016, 82, 102-105 ORIGINAL STUDY Exostoses, enchondromatosis and metachondromatosis; diagnosis and management John MCFARLANE, Tim KNIGHT, Anubha SINHA, Trevor COLE, Nigel KIELY, Rob FREEMAN From the Department of Orthopaedics, Robert Jones Agnes Hunt Hospital, Oswestry, UK We describe a 5 years old girl who presented to the region of long bones and are composed of a carti- multidisciplinary skeletal dysplasia clinic following lage lump outside the bone which may be peduncu- excision of two bony lumps from her fingers. Based on lated or sessile, the knee is the most common clinical examination, radiolographs and histological site (1,10). An isolated exostosis is a common inci- results an initial diagnosis of hereditary multiple dental finding rarely requiring treatment. Disorders exostosis (HME) was made. Four years later she developed further lumps which had the radiological associated with exostoses include HME, Langer- appearance of enchondromas. The appearance of Giedion syndrome, Gardner syndrome and meta- both exostoses and enchondromas suggested a possi- chondromatosis. ble diagnosis of metachondromatosis. Genetic testing Enchondroma are the second most common be- revealed a splice site mutation at the end of exon 11 on nign bone tumour characterised by the formation of the PTPN11 gene, confirming the diagnosis of meta- hyaline cartilage in the medulla of a bone. It occurs chondromatosis. While both single or multiple exosto- most frequently in the hand (60%) and then the feet. ses and enchondromas occur relatively commonly on The typical radiological features are of a well- their own, the appearance of multiple exostoses and defined lucent defect with endosteal scalloping and enchondromas together is rare and should raise the differential diagnosis of metachondromatosis. -

Phenotypic and Genotypic Characterisation of Noonan-Like
1of5 ELECTRONIC LETTER J Med Genet: first published as 10.1136/jmg.2004.024091 on 2 February 2005. Downloaded from Phenotypic and genotypic characterisation of Noonan-like/ multiple giant cell lesion syndrome J S Lee, M Tartaglia, B D Gelb, K Fridrich, S Sachs, C A Stratakis, M Muenke, P G Robey, M T Collins, A Slavotinek ............................................................................................................................... J Med Genet 2005;42:e11 (http://www.jmedgenet.com/cgi/content/full/42/2/e11). doi: 10.1136/jmg.2004.024091 oonan-like/multiple giant cell lesion syndrome (NL/ MGCLS; OMIM 163955) is a rare condition1–3 with Key points Nphenotypic overlap with Noonan’s syndrome (OMIM 163950) and cherubism (OMIM 118400) (table 1). N Noonan-like/multiple giant cell lesion syndrome (NL/ Recently, missense mutations in the PTPN11 gene on MGCLS) has clinical similarities with Noonan’s syn- chromosome 12q24.1 have been identified as the cause of drome and cherubism. It is unclear whether it is a Noonan’s syndrome in 45% of familial and sporadic cases,45 distinct entity or a variant of Noonan’s syndrome or indicating genetic heterogeneity within the syndrome. In the cherubism. 5 study by Tartaglia et al, there was a family in which three N Three unrelated patients with NL/MGCLS were char- members had features of Noonan’s syndrome; two of these acterised, two of whom were found to have mutations had incidental mandibular giant cell lesions.3 All three in the PTPN11 gene, the mutation found in 45% of members were found to have a PTPN11 mutation known to patients with Noonan’s syndrome. -

Megalencephaly and Macrocephaly
277 Megalencephaly and Macrocephaly KellenD.Winden,MD,PhD1 Christopher J. Yuskaitis, MD, PhD1 Annapurna Poduri, MD, MPH2 1 Department of Neurology, Boston Children’s Hospital, Boston, Address for correspondence Annapurna Poduri, Epilepsy Genetics Massachusetts Program, Division of Epilepsy and Clinical Electrophysiology, 2 Epilepsy Genetics Program, Division of Epilepsy and Clinical Department of Neurology, Fegan 9, Boston Children’s Hospital, 300 Electrophysiology, Department of Neurology, Boston Children’s Longwood Avenue, Boston, MA 02115 Hospital, Boston, Massachusetts (e-mail: [email protected]). Semin Neurol 2015;35:277–287. Abstract Megalencephaly is a developmental disorder characterized by brain overgrowth secondary to increased size and/or numbers of neurons and glia. These disorders can be divided into metabolic and developmental categories based on their molecular etiologies. Metabolic megalencephalies are mostly caused by genetic defects in cellular metabolism, whereas developmental megalencephalies have recently been shown to be caused by alterations in signaling pathways that regulate neuronal replication, growth, and migration. These disorders often lead to epilepsy, developmental disabilities, and Keywords behavioral problems; specific disorders have associations with overgrowth or abnor- ► megalencephaly malities in other tissues. The molecular underpinnings of many of these disorders are ► hemimegalencephaly now understood, providing insight into how dysregulation of critical pathways leads to ► -

Cardiomyopathy Precision Panel Overview Indications
Cardiomyopathy Precision Panel Overview Cardiomyopathies are a group of conditions with a strong genetic background that structurally hinder the heart to pump out blood to the rest of the body due to weakness in the heart muscles. These diseases affect individuals of all ages and can lead to heart failure and sudden cardiac death. If there is a family history of cardiomyopathy it is strongly recommended to undergo genetic testing to be aware of the family risk, personal risk, and treatment options. Most types of cardiomyopathies are inherited in a dominant manner, which means that one altered copy of the gene is enough for the disease to present in an individual. The symptoms of cardiomyopathy are variable, and these diseases can present in different ways. There are 5 types of cardiomyopathies, the most common being hypertrophic cardiomyopathy: 1. Hypertrophic cardiomyopathy (HCM) 2. Dilated cardiomyopathy (DCM) 3. Restrictive cardiomyopathy (RCM) 4. Arrhythmogenic Right Ventricular Cardiomyopathy (ARVC) 5. Isolated Left Ventricular Non-Compaction Cardiomyopathy (LVNC). The Igenomix Cardiomyopathy Precision Panel serves as a diagnostic and tool ultimately leading to a better management and prognosis of the disease. It provides a comprehensive analysis of the genes involved in this disease using next-generation sequencing (NGS) to fully understand the spectrum of relevant genes. Indications The Igenomix Cardiomyopathy Precision Panel is indicated in those cases where there is a clinical suspicion of cardiomyopathy with or without the following manifestations: - Shortness of breath - Fatigue - Arrythmia (abnormal heart rhythm) - Family history of arrhythmia - Abnormal scans - Ventricular tachycardia - Ventricular fibrillation - Chest Pain - Dizziness - Sudden cardiac death in the family 1 Clinical Utility The clinical utility of this panel is: - The genetic and molecular diagnosis for an accurate clinical diagnosis of a patient with personal or family history of cardiomyopathy, channelopathy or sudden cardiac death. -

Targeted Disruption of Shp2 in Chondrocytes Leads to Metachondromatosis with Multiple Cartilaginous Protrusions
ORIGINAL ARTICLE JBMR Targeted Disruption of Shp2 in Chondrocytes Leads to Metachondromatosis With Multiple Cartilaginous Protrusions Harry KW Kim,1,2 Gen‐Sheng Feng,3 Di Chen,4 Philip D King,5 and Nobuhiro Kamiya1,2 1Texas Scottish Rite Hospital for Children, Dallas, TX, USA 2Orthopaedic Surgery, University of Texas Southwestern Medical Center, Dallas, TX, USA 3Department of Pathology and Division of Biological Sciences, University of California San Diego, La Jolla, CA, USA 4Department of Biochemistry, Rush University Medical Center, Chicago, IL, USA 5Department of Microbiology and Immunology, University of Michigan Medical School, Ann Arbor, MI, USA ABSTRACT Metachondromatosis is a benign bone disease predominantly observed in the hands and feet of children or young adults demonstrating two different manifestations: a cartilage‐capped bony outgrowth on the surface of the bone called exostosis and ectopic cartilaginous nodules inside the bone called enchondroma. Recently, it has been reported that loss‐of‐function mutations of the SHP2 gene, which encodes the SHP2 protein tyrosine phosphatase, are associated with metachondromatosis. The purpose of this study was to investigate the role of SHP2 in postnatal cartilage development, which is largely unknown. We disrupted Shp2 during the postnatal stage of mouse development in a chondrocyte‐specific manner using a tamoxifen‐inducible system. We found tumor‐like nodules on the hands and feet within a month after the initial induction. The SHP2‐deficient mice demonstrated an exostosis‐like and enchondroma‐like phenotype in multiple bones of the hands, feet, and ribs as assessed by X‐ray and micro‐computed tomography (CT). Histological assessment revealed the disorganization of the growth plate cartilage, a cartilaginous protrusion from the epiphyseal bone, and ectopic cartilage nodules within the bones, which is consistent with the pathological features of metachondromatosis in humans (ie, both exostosis and enchondroma). -

Prevalence and Incidence of Rare Diseases: Bibliographic Data
Number 1 | January 2019 Prevalence and incidence of rare diseases: Bibliographic data Prevalence, incidence or number of published cases listed by diseases (in alphabetical order) www.orpha.net www.orphadata.org If a range of national data is available, the average is Methodology calculated to estimate the worldwide or European prevalence or incidence. When a range of data sources is available, the most Orphanet carries out a systematic survey of literature in recent data source that meets a certain number of quality order to estimate the prevalence and incidence of rare criteria is favoured (registries, meta-analyses, diseases. This study aims to collect new data regarding population-based studies, large cohorts studies). point prevalence, birth prevalence and incidence, and to update already published data according to new For congenital diseases, the prevalence is estimated, so scientific studies or other available data. that: Prevalence = birth prevalence x (patient life This data is presented in the following reports published expectancy/general population life expectancy). biannually: When only incidence data is documented, the prevalence is estimated when possible, so that : • Prevalence, incidence or number of published cases listed by diseases (in alphabetical order); Prevalence = incidence x disease mean duration. • Diseases listed by decreasing prevalence, incidence When neither prevalence nor incidence data is available, or number of published cases; which is the case for very rare diseases, the number of cases or families documented in the medical literature is Data collection provided. A number of different sources are used : Limitations of the study • Registries (RARECARE, EUROCAT, etc) ; The prevalence and incidence data presented in this report are only estimations and cannot be considered to • National/international health institutes and agencies be absolutely correct. -

MECHANISMS in ENDOCRINOLOGY: Novel Genetic Causes of Short Stature
J M Wit and others Genetics of short stature 174:4 R145–R173 Review MECHANISMS IN ENDOCRINOLOGY Novel genetic causes of short stature 1 1 2 2 Jan M Wit , Wilma Oostdijk , Monique Losekoot , Hermine A van Duyvenvoorde , Correspondence Claudia A L Ruivenkamp2 and Sarina G Kant2 should be addressed to J M Wit Departments of 1Paediatrics and 2Clinical Genetics, Leiden University Medical Center, PO Box 9600, 2300 RC Leiden, Email The Netherlands [email protected] Abstract The fast technological development, particularly single nucleotide polymorphism array, array-comparative genomic hybridization, and whole exome sequencing, has led to the discovery of many novel genetic causes of growth failure. In this review we discuss a selection of these, according to a diagnostic classification centred on the epiphyseal growth plate. We successively discuss disorders in hormone signalling, paracrine factors, matrix molecules, intracellular pathways, and fundamental cellular processes, followed by chromosomal aberrations including copy number variants (CNVs) and imprinting disorders associated with short stature. Many novel causes of GH deficiency (GHD) as part of combined pituitary hormone deficiency have been uncovered. The most frequent genetic causes of isolated GHD are GH1 and GHRHR defects, but several novel causes have recently been found, such as GHSR, RNPC3, and IFT172 mutations. Besides well-defined causes of GH insensitivity (GHR, STAT5B, IGFALS, IGF1 defects), disorders of NFkB signalling, STAT3 and IGF2 have recently been discovered. Heterozygous IGF1R defects are a relatively frequent cause of prenatal and postnatal growth retardation. TRHA mutations cause a syndromic form of short stature with elevated T3/T4 ratio. Disorders of signalling of various paracrine factors (FGFs, BMPs, WNTs, PTHrP/IHH, and CNP/NPR2) or genetic defects affecting cartilage extracellular matrix usually cause disproportionate short stature. -

Whole Exome Sequencing Gene Package Intellectual Disability, Version 9.1, 31-1-2020
Whole Exome Sequencing Gene package Intellectual disability, version 9.1, 31-1-2020 Technical information DNA was enriched using Agilent SureSelect DNA + SureSelect OneSeq 300kb CNV Backbone + Human All Exon V7 capture and paired-end sequenced on the Illumina platform (outsourced). The aim is to obtain 10 Giga base pairs per exome with a mapped fraction of 0.99. The average coverage of the exome is ~50x. Duplicate and non-unique reads are excluded. Data are demultiplexed with bcl2fastq Conversion Software from Illumina. Reads are mapped to the genome using the BWA-MEM algorithm (reference: http://bio-bwa.sourceforge.net/). Variant detection is performed by the Genome Analysis Toolkit HaplotypeCaller (reference: http://www.broadinstitute.org/gatk/). The detected variants are filtered and annotated with Cartagenia software and classified with Alamut Visual. It is not excluded that pathogenic mutations are being missed using this technology. At this moment, there is not enough information about the sensitivity of this technique with respect to the detection of deletions and duplications of more than 5 nucleotides and of somatic mosaic mutations (all types of sequence changes). HGNC approved Phenotype description including OMIM phenotype ID(s) OMIM median depth % covered % covered % covered gene symbol gene ID >10x >20x >30x A2ML1 {Otitis media, susceptibility to}, 166760 610627 66 100 100 96 AARS1 Charcot-Marie-Tooth disease, axonal, type 2N, 613287 601065 63 100 97 90 Epileptic encephalopathy, early infantile, 29, 616339 AASS Hyperlysinemia,