Blueprint Genetics Hereditary Leukemia Panel
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Exostoses, Enchondromatosis and Metachondromatosis; Diagnosis and Management
Acta Orthop. Belg., 2016, 82, 102-105 ORIGINAL STUDY Exostoses, enchondromatosis and metachondromatosis; diagnosis and management John MCFARLANE, Tim KNIGHT, Anubha SINHA, Trevor COLE, Nigel KIELY, Rob FREEMAN From the Department of Orthopaedics, Robert Jones Agnes Hunt Hospital, Oswestry, UK We describe a 5 years old girl who presented to the region of long bones and are composed of a carti- multidisciplinary skeletal dysplasia clinic following lage lump outside the bone which may be peduncu- excision of two bony lumps from her fingers. Based on lated or sessile, the knee is the most common clinical examination, radiolographs and histological site (1,10). An isolated exostosis is a common inci- results an initial diagnosis of hereditary multiple dental finding rarely requiring treatment. Disorders exostosis (HME) was made. Four years later she developed further lumps which had the radiological associated with exostoses include HME, Langer- appearance of enchondromas. The appearance of Giedion syndrome, Gardner syndrome and meta- both exostoses and enchondromas suggested a possi- chondromatosis. ble diagnosis of metachondromatosis. Genetic testing Enchondroma are the second most common be- revealed a splice site mutation at the end of exon 11 on nign bone tumour characterised by the formation of the PTPN11 gene, confirming the diagnosis of meta- hyaline cartilage in the medulla of a bone. It occurs chondromatosis. While both single or multiple exosto- most frequently in the hand (60%) and then the feet. ses and enchondromas occur relatively commonly on The typical radiological features are of a well- their own, the appearance of multiple exostoses and defined lucent defect with endosteal scalloping and enchondromas together is rare and should raise the differential diagnosis of metachondromatosis. -

Cardiomyopathy Precision Panel Overview Indications
Cardiomyopathy Precision Panel Overview Cardiomyopathies are a group of conditions with a strong genetic background that structurally hinder the heart to pump out blood to the rest of the body due to weakness in the heart muscles. These diseases affect individuals of all ages and can lead to heart failure and sudden cardiac death. If there is a family history of cardiomyopathy it is strongly recommended to undergo genetic testing to be aware of the family risk, personal risk, and treatment options. Most types of cardiomyopathies are inherited in a dominant manner, which means that one altered copy of the gene is enough for the disease to present in an individual. The symptoms of cardiomyopathy are variable, and these diseases can present in different ways. There are 5 types of cardiomyopathies, the most common being hypertrophic cardiomyopathy: 1. Hypertrophic cardiomyopathy (HCM) 2. Dilated cardiomyopathy (DCM) 3. Restrictive cardiomyopathy (RCM) 4. Arrhythmogenic Right Ventricular Cardiomyopathy (ARVC) 5. Isolated Left Ventricular Non-Compaction Cardiomyopathy (LVNC). The Igenomix Cardiomyopathy Precision Panel serves as a diagnostic and tool ultimately leading to a better management and prognosis of the disease. It provides a comprehensive analysis of the genes involved in this disease using next-generation sequencing (NGS) to fully understand the spectrum of relevant genes. Indications The Igenomix Cardiomyopathy Precision Panel is indicated in those cases where there is a clinical suspicion of cardiomyopathy with or without the following manifestations: - Shortness of breath - Fatigue - Arrythmia (abnormal heart rhythm) - Family history of arrhythmia - Abnormal scans - Ventricular tachycardia - Ventricular fibrillation - Chest Pain - Dizziness - Sudden cardiac death in the family 1 Clinical Utility The clinical utility of this panel is: - The genetic and molecular diagnosis for an accurate clinical diagnosis of a patient with personal or family history of cardiomyopathy, channelopathy or sudden cardiac death. -

Targeted Disruption of Shp2 in Chondrocytes Leads to Metachondromatosis with Multiple Cartilaginous Protrusions
ORIGINAL ARTICLE JBMR Targeted Disruption of Shp2 in Chondrocytes Leads to Metachondromatosis With Multiple Cartilaginous Protrusions Harry KW Kim,1,2 Gen‐Sheng Feng,3 Di Chen,4 Philip D King,5 and Nobuhiro Kamiya1,2 1Texas Scottish Rite Hospital for Children, Dallas, TX, USA 2Orthopaedic Surgery, University of Texas Southwestern Medical Center, Dallas, TX, USA 3Department of Pathology and Division of Biological Sciences, University of California San Diego, La Jolla, CA, USA 4Department of Biochemistry, Rush University Medical Center, Chicago, IL, USA 5Department of Microbiology and Immunology, University of Michigan Medical School, Ann Arbor, MI, USA ABSTRACT Metachondromatosis is a benign bone disease predominantly observed in the hands and feet of children or young adults demonstrating two different manifestations: a cartilage‐capped bony outgrowth on the surface of the bone called exostosis and ectopic cartilaginous nodules inside the bone called enchondroma. Recently, it has been reported that loss‐of‐function mutations of the SHP2 gene, which encodes the SHP2 protein tyrosine phosphatase, are associated with metachondromatosis. The purpose of this study was to investigate the role of SHP2 in postnatal cartilage development, which is largely unknown. We disrupted Shp2 during the postnatal stage of mouse development in a chondrocyte‐specific manner using a tamoxifen‐inducible system. We found tumor‐like nodules on the hands and feet within a month after the initial induction. The SHP2‐deficient mice demonstrated an exostosis‐like and enchondroma‐like phenotype in multiple bones of the hands, feet, and ribs as assessed by X‐ray and micro‐computed tomography (CT). Histological assessment revealed the disorganization of the growth plate cartilage, a cartilaginous protrusion from the epiphyseal bone, and ectopic cartilage nodules within the bones, which is consistent with the pathological features of metachondromatosis in humans (ie, both exostosis and enchondroma). -

Prevalence and Incidence of Rare Diseases: Bibliographic Data
Number 1 | January 2019 Prevalence and incidence of rare diseases: Bibliographic data Prevalence, incidence or number of published cases listed by diseases (in alphabetical order) www.orpha.net www.orphadata.org If a range of national data is available, the average is Methodology calculated to estimate the worldwide or European prevalence or incidence. When a range of data sources is available, the most Orphanet carries out a systematic survey of literature in recent data source that meets a certain number of quality order to estimate the prevalence and incidence of rare criteria is favoured (registries, meta-analyses, diseases. This study aims to collect new data regarding population-based studies, large cohorts studies). point prevalence, birth prevalence and incidence, and to update already published data according to new For congenital diseases, the prevalence is estimated, so scientific studies or other available data. that: Prevalence = birth prevalence x (patient life This data is presented in the following reports published expectancy/general population life expectancy). biannually: When only incidence data is documented, the prevalence is estimated when possible, so that : • Prevalence, incidence or number of published cases listed by diseases (in alphabetical order); Prevalence = incidence x disease mean duration. • Diseases listed by decreasing prevalence, incidence When neither prevalence nor incidence data is available, or number of published cases; which is the case for very rare diseases, the number of cases or families documented in the medical literature is Data collection provided. A number of different sources are used : Limitations of the study • Registries (RARECARE, EUROCAT, etc) ; The prevalence and incidence data presented in this report are only estimations and cannot be considered to • National/international health institutes and agencies be absolutely correct. -

Neoreviews Quiz
Birthmarks of Potential Medical Significance Jacinto A. Hernández and Joseph G. Morelli NeoReviews 2003;4;263 DOI: 10.1542/neo.4-10-e263 The online version of this article, along with updated information and services, is located on the World Wide Web at: http://neoreviews.aappublications.org/cgi/content/full/neoreviews;4/10/e263 NeoReviews is the official journal of the American Academy of Pediatrics. A monthly publication, it has been published continuously since 2000. NeoReviews is owned, published, and trademarked by the American Academy of Pediatrics, 141 Northwest Point Boulevard, Elk Grove Village, Illinois, 60007. Copyright © 2003 by the American Academy of Pediatrics. All rights reserved. Online ISSN: 1526-9906. Downloaded from http://neoreviews.aappublications.org by J Michael Coleman on August 19, 2010 Article dermatology Birthmarks of Potential Medical Significance Jacinto A. Herna´ndez, Objectives After completing this article, readers should be able to: MD*, Joseph G. Morelli, MD† 1. Describe the clinical manifestations of neurofibromatosis type 1. 2. List conditions associated with cafe´au lait macules. 3. Explain the clinical significance of congenital melanocytic nevi. 4. Characterize tuberous sclerosis complex. 5. List the two syndromes associated with port-wine stains and extracutaneous abnormalities. 6. Categorize and describe the three types of hemangiomas. Introduction Birthmarks are common (ϳ8% to 10%) in newborns. Most birthmarks represent vascular and pigmentary lesions. The natural history of these lesions varies from being transient phenomena and essentially normal variants of no clinical significance to permanent cutaneous abnormalities that may be associated with significant systemic complications or diseases. Table 1 lists some neonatal skin lesions that should be recognized by the clinician as clues to more serious disorders. -

Whole Exome Sequencing Gene Package Intellectual Disability, Version 9.1, 31-1-2020
Whole Exome Sequencing Gene package Intellectual disability, version 9.1, 31-1-2020 Technical information DNA was enriched using Agilent SureSelect DNA + SureSelect OneSeq 300kb CNV Backbone + Human All Exon V7 capture and paired-end sequenced on the Illumina platform (outsourced). The aim is to obtain 10 Giga base pairs per exome with a mapped fraction of 0.99. The average coverage of the exome is ~50x. Duplicate and non-unique reads are excluded. Data are demultiplexed with bcl2fastq Conversion Software from Illumina. Reads are mapped to the genome using the BWA-MEM algorithm (reference: http://bio-bwa.sourceforge.net/). Variant detection is performed by the Genome Analysis Toolkit HaplotypeCaller (reference: http://www.broadinstitute.org/gatk/). The detected variants are filtered and annotated with Cartagenia software and classified with Alamut Visual. It is not excluded that pathogenic mutations are being missed using this technology. At this moment, there is not enough information about the sensitivity of this technique with respect to the detection of deletions and duplications of more than 5 nucleotides and of somatic mosaic mutations (all types of sequence changes). HGNC approved Phenotype description including OMIM phenotype ID(s) OMIM median depth % covered % covered % covered gene symbol gene ID >10x >20x >30x A2ML1 {Otitis media, susceptibility to}, 166760 610627 66 100 100 96 AARS1 Charcot-Marie-Tooth disease, axonal, type 2N, 613287 601065 63 100 97 90 Epileptic encephalopathy, early infantile, 29, 616339 AASS Hyperlysinemia, -

Psykisk Utviklingshemming Og Forsinket Utvikling
Psykisk utviklingshemming og forsinket utvikling Genpanel, versjon v03 Tabellen er sortert på gennavn (HGNC gensymbol) Navn på gen er iht. HGNC >x10 Andel av genet som har blitt lest med tilfredstillende kvalitet flere enn 10 ganger under sekvensering x10 er forventet dekning; faktisk dekning vil variere. Gen Gen (HGNC Transkript >10x Fenotype (symbol) ID) AAAS 13666 NM_015665.5 100% Achalasia-addisonianism-alacrimia syndrome OMIM AARS 20 NM_001605.2 100% Charcot-Marie-Tooth disease, axonal, type 2N OMIM Epileptic encephalopathy, early infantile, 29 OMIM AASS 17366 NM_005763.3 100% Hyperlysinemia OMIM Saccharopinuria OMIM ABCB11 42 NM_003742.2 100% Cholestasis, benign recurrent intrahepatic, 2 OMIM Cholestasis, progressive familial intrahepatic 2 OMIM ABCB7 48 NM_004299.5 100% Anemia, sideroblastic, with ataxia OMIM ABCC6 57 NM_001171.5 93% Arterial calcification, generalized, of infancy, 2 OMIM Pseudoxanthoma elasticum OMIM Pseudoxanthoma elasticum, forme fruste OMIM ABCC9 60 NM_005691.3 100% Hypertrichotic osteochondrodysplasia OMIM ABCD1 61 NM_000033.3 77% Adrenoleukodystrophy OMIM Adrenomyeloneuropathy, adult OMIM ABCD4 68 NM_005050.3 100% Methylmalonic aciduria and homocystinuria, cblJ type OMIM ABHD5 21396 NM_016006.4 100% Chanarin-Dorfman syndrome OMIM ACAD9 21497 NM_014049.4 99% Mitochondrial complex I deficiency due to ACAD9 deficiency OMIM ACADM 89 NM_000016.5 100% Acyl-CoA dehydrogenase, medium chain, deficiency of OMIM ACADS 90 NM_000017.3 100% Acyl-CoA dehydrogenase, short-chain, deficiency of OMIM ACADVL 92 NM_000018.3 100% VLCAD -
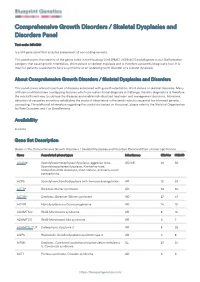
Blueprint Genetics Comprehensive Growth Disorders / Skeletal
Comprehensive Growth Disorders / Skeletal Dysplasias and Disorders Panel Test code: MA4301 Is a 374 gene panel that includes assessment of non-coding variants. This panel covers the majority of the genes listed in the Nosology 2015 (PMID: 26394607) and all genes in our Malformation category that cause growth retardation, short stature or skeletal dysplasia and is therefore a powerful diagnostic tool. It is ideal for patients suspected to have a syndromic or an isolated growth disorder or a skeletal dysplasia. About Comprehensive Growth Disorders / Skeletal Dysplasias and Disorders This panel covers a broad spectrum of diseases associated with growth retardation, short stature or skeletal dysplasia. Many of these conditions have overlapping features which can make clinical diagnosis a challenge. Genetic diagnostics is therefore the most efficient way to subtype the diseases and enable individualized treatment and management decisions. Moreover, detection of causative mutations establishes the mode of inheritance in the family which is essential for informed genetic counseling. For additional information regarding the conditions tested on this panel, please refer to the National Organization for Rare Disorders and / or GeneReviews. Availability 4 weeks Gene Set Description Genes in the Comprehensive Growth Disorders / Skeletal Dysplasias and Disorders Panel and their clinical significance Gene Associated phenotypes Inheritance ClinVar HGMD ACAN# Spondyloepimetaphyseal dysplasia, aggrecan type, AD/AR 20 56 Spondyloepiphyseal dysplasia, Kimberley -

Blueprint Genetics Comprehensive Skeletal Dysplasias and Disorders
Comprehensive Skeletal Dysplasias and Disorders Panel Test code: MA3301 Is a 251 gene panel that includes assessment of non-coding variants. Is ideal for patients with a clinical suspicion of disorders involving the skeletal system. About Comprehensive Skeletal Dysplasias and Disorders This panel covers a broad spectrum of skeletal disorders including common and rare skeletal dysplasias (eg. achondroplasia, COL2A1 related dysplasias, diastrophic dysplasia, various types of spondylo-metaphyseal dysplasias), various ciliopathies with skeletal involvement (eg. short rib-polydactylies, asphyxiating thoracic dysplasia dysplasias and Ellis-van Creveld syndrome), various subtypes of osteogenesis imperfecta, campomelic dysplasia, slender bone dysplasias, dysplasias with multiple joint dislocations, chondrodysplasia punctata group of disorders, neonatal osteosclerotic dysplasias, osteopetrosis and related disorders, abnormal mineralization group of disorders (eg hypopohosphatasia), osteolysis group of disorders, disorders with disorganized development of skeletal components, overgrowth syndromes with skeletal involvement, craniosynostosis syndromes, dysostoses with predominant craniofacial involvement, dysostoses with predominant vertebral involvement, patellar dysostoses, brachydactylies, some disorders with limb hypoplasia-reduction defects, ectrodactyly with and without other manifestations, polydactyly-syndactyly-triphalangism group of disorders, and disorders with defects in joint formation and synostoses. Availability 4 weeks Gene Set Description -

Blueprint Genetics Bone Marrow Failure Syndrome Panel
Bone Marrow Failure Syndrome Panel Test code: HE0801 Is a 135 gene panel that includes assessment of non-coding variants. Is ideal for patients with a clinical suspicion of inherited bone marrow failure syndromes. The genes on this panel are included in the Comprehensive Hematology Panel. About Bone Marrow Failure Syndrome Inherited bone marrow failure syndromes (IBMFS) are a diverse set of genetic disorders characterized by the inability of the bone marrow to produce sufficient circulating blood cells. Bone marrow failure can affect all blood cell lineages causing clinical symptoms similar to aplastic anemia, or be restricted to one or two blood cell lineages. The clinical presentation may include thrombocytopenia or neutropenia. Hematological manifestations may be accompanied by physical features such as short stature and abnormal skin pigmentation in Fanconi anemia and dystrophic nails, lacy reticular pigmentation and oral leukoplakia in dyskeratosis congenita. Patients with IBMFS have an increased risk of developing cancer—either hematological or solid tumors. Early and correct disease recognition is important for management and surveillance of the diseases. Currently, accurate genetic diagnosis is essential to confirm the clinical diagnosis. The most common phenotypes that are covered by the panel are Fanconi anemia, Diamond-Blackfan anemia, dyskeratosis congenita, Shwachman-Diamond syndrome and WAS-related disorders. Availability 4 weeks Gene Set Description Genes in the Bone Marrow Failure Syndrome Panel and their clinical significance -

Noonan Spectrum Disorders Panel, Sequencing ARUP Test Code 2010772 Noonan Disorders Sequencing Specimen Whole Blood
Patient Report |FINAL Client: Example Client ABC123 Patient: Patient, Example 123 Test Drive Salt Lake City, UT 84108 DOB 12/9/2011 UNITED STATES Gender: Female Patient Identifiers: 01234567890ABCD, 012345 Physician: Doctor, Example Visit Number (FIN): 01234567890ABCD Collection Date: 01/01/2017 12:34 Noonan Spectrum Disorders Panel, Sequencing ARUP test code 2010772 Noonan Disorders Sequencing Specimen Whole Blood Noonan Disorders Sequencing Interp Positive INDICATION FOR TESTING Short stature and facial features suggestive of Noonan syndrome. RESULT One pathogenic variant was detected in the PTPN11 gene. PATHOGENIC VARIANT Gene: PTPN11 (NM_002834.3) Nucleic Acid Change: c.188A>G; Heterozygous Amino Acid Alteration: p.Tyr63Cys Inheritance: Autosomal Dominant INTERPRETATION One pathogenic variant, c.188A>G; p.Tyr63Cys, was detected in the PTPN11 gene by massively parallel sequencing and confirmed by Sanger sequencing. Pathogenic PTPN11 variants are inherited in an autosomal dominant manner, and are associated with Noonan syndrome 1 (MIM: 163950), metachondromatosis (MIM: 156250), and LEOPARD syndrome 1 (MIM: 151100). No additional pathogenic variants were identified in the other targeted genes by massively parallel sequencing. Please refer to the background information included in this report for a list of the genes analyzed and limitations of this test. Evidence for variant classification: The PTPN11 c.188A>G, p.Tyr63Cys variant (rs121918459) has been reported in multiple patients diagnosed with Noonan syndrome (Tartaglia 2001, Jongmans 2011, Martinelli 2012, Hashida 2013, Lepri 2014, Okamoto 2015). This variant is located in a structurally important region of the catalytic N-terminal SH2 domain of PTPN11 (Hof 1998), and several additional variants in neighboring codons have also been identified in Noonan patients (Jongmans 2011, Tartaglia 2002, Tartaglia 2006). -

Cutaneous Markers of Systemic Disease and a Window to New Aspects of Tumorigenesis
JMG Online First, published on June 15, 2005 as 10.1136/jmg.2003.017806 J Med Genet: first published as 10.1136/jmg.2003.017806 on 15 June 2005. Downloaded from The lentiginoses: Cutaneous markers of systemic disease and a window to new aspects of tumorigenesis Andrew J. Bauer (1, 2) and Constantine A. Stratakis (1) @ From the 1. Section on Endocrinology & Genetics (SEGEN), Developmental Endocrinology Branch (DEB), National Institute of Child Health and Human Development (NICHD), National Institutes of Health, Bethesda, MD 20892-1103, and 2. Walter Reed Army Medical Center, Department of Pediatrics, Washington, DC, 20307 http://jmg.bmj.com/ Short title: Familial lentiginoses Key Words: Carney complex, Peutz-Jeghers syndrome, L.E.O.P.A.R.D. syndrome, Noonan syndrome, on September 30, 2021 by guest. Protected copyright. Cowden disease @ Address all correspondence and reprint requests to: Dr. Constantine A. Stratakis, Section on Endocrinology & Genetics, DEB, NICHD, NIH, Building 10, CRC, Room I-3330, 10 Center Dr., MSC 1103, Bethesda, Maryland 20892, Tel: 301-4964686, Fax: 301-4020574, E-mail: [email protected]; the corresponding author has the right to grant on behalf of all authors and does grant on behalf of all authors, a non-exclusive licence (because both Drs Stratakis and Bauer are U.S. government employees) on a worldwide basis to the BMJ Publishing Group Ltd to permit this article to be published in JMG/ptMJPGL products and sublicences such use and exploit all subsidiary rights, as set out in the Journal’s site http://JMG.bmjjournals.com/misc/ifora/licenceform.shtml. Copyright Article author (or their employer) 2005.