MECHANISMS in ENDOCRINOLOGY: Novel Genetic Causes of Short Stature
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Genetic Screening Identifies a Component of the SWI/SNF Complex, Arid1b As a Senescence Regulator
A genetic screening identifies a component of the SWI/SNF complex, Arid1b as a senescence regulator Sadaf Khan A thesis submitted to Imperial College London for the degree of Doctor in Philosophy MRC Clinical Sciences Centre Imperial College London, School of Medicine July 2013 Statement of originality All experiments included in this thesis were performed by myself unless otherwise stated. Copyright Declaration The copyright of this thesis rests with the author and is made available under a Creative Commons Attribution Non-Commercial No Derivatives license. Researchers are free to copy, distribute or transmit the thesis on the condition that they attribute it, that they do not use it for commercial purposes and that they do not alter, transform or build upon it. For any reuse or redistribution, researchers must make clear to others the license terms of this work. 2 Abstract Senescence is an important tumour suppressor mechanism, which prevents the proliferation of stressed or damaged cells. The use of RNA interference to identify genes with a role in senescence is an important tool in the discovery of novel cancer genes. In this work, a protocol was established for conducting bypass of senescence screenings, using shRNA libraries together with next-generation sequencing. Using this approach, the SWI/SNF subunit Arid1b was identified as a regulator of cellular lifespan in MEFs. SWI/SNF is a large multi-subunit complex that remodels chromatin. Mutations in SWI/SNF proteins are frequently associated with cancer, suggesting that SWI/SNF components are tumour suppressors. Here the role of ARID1B during senescence was investigated. Depletion of ARID1B extends the proliferative capacity of primary mouse and human fibroblasts. -
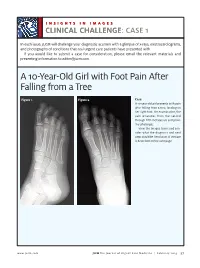
A 10-Year-Old Girl with Foot Pain After Falling from a Tree
INSIGHTS IN IMAGES CLINICAL CHALLENGECHALLENGE: CASE 1 In each issue, JUCM will challenge your diagnostic acumen with a glimpse of x-rays, electrocardiograms, and photographs of conditions that real urgent care patients have presented with. If you would like to submit a case for consideration, please email the relevant materials and presenting information to [email protected]. A 10-Year-Old Girl with Foot Pain After Falling from a Tree Figure 1. Figure 2. Case A 10-year-old girl presents with pain after falling from a tree, landing on her right foot. On examination, the pain emanates from the second through fifth metatarsals and proxi- mal phalanges. View the images taken and con- sider what the diagnosis and next steps would be. Resolution of the case is described on the next page. www.jucm.com JUCM The Journal of Urgent Care Medicine | February 2019 37 INSIGHTS IN IMAGES: CLINICAL CHALLENGE THE RESOLUTION Figure 1. Mediastinal air Figure 1. Differential Diagnosis Pearls for Urgent Care Management and Ⅲ Fracture of the distal fourth metatarsal Considerations for Transfer Ⅲ Plantar plate disruption Ⅲ Emergent transfer should be considered with associated neu- Ⅲ Sesamoiditis rologic deficit, compartment syndrome, open fracture, or vas- Ⅲ Turf toe cular compromise Ⅲ Referral to an orthopedist is warranted in the case of an in- Diagnosis tra-articular fracture, or with Lisfranc ligament injury or ten- Angulation of the distal fourth metatarsal metaphyseal cortex derness over the Lisfranc ligament and hairline lucency consistent with fracture. Acknowledgment: Images courtesy of Teleradiology Associates. Learnings/What to Look for Ⅲ Proximal metatarsal fractures are most often caused by crush- ing or direct blows Ⅲ In athletes, an axial load placed on a plantar-flexed foot should raise suspicion of a Lisfranc injury 38 JUCM The Journal of Urgent Care Medicine | February 2019 www.jucm.com INSIGHTS IN IMAGES CLINICAL CHALLENGE: CASE 2 A 55-Year-Old Man with 3 Hours of Epigastric Pain 55 years PR 249 QRSD 90 QT 471 QTc 425 AXES P 64 QRS -35 T 30 Figure 1. -

The Genetical Society of Great Britain
Heredity 59 (1987) 151—160 The Genetical Society of Great Britain THEGENETICAL SOCIETY (Abstracts of papers presented at the TVVO HUNDRED AND FIFTH MEETING of the Society held on Friday, 14th and Saturday, 15th November 1986 at UNIVERSITY COLLEGE, LONDON) 1. Selection of somatic cell D. J. Porteous, P. A. Boyd, N. D. Hastie and hybrids with specific chromosome V. van Heyningen content for mapping the WAGR MAC Clinical and Population Cytogenetics Unit, Western General Hospital, Crewe Road, syndrome Edinburgh EH4 2XU. J. M. Fletcher, H. Morrison, J. A. Fantes, Clonedprobes for a number of available chromo- A. Seawright, S. Christie, D. J. Porteous, some ii assigned genes were used to define the N. D. Hastie and V. van Heyningen extent of deletions associated with the Wilms' MAC Clinical and Population Cytogenetics Unit, tumour, aniridia, genitourinary abnormalities and Western General Hospital, Crewe Road, mental retardation (WAGR) syndrome. Establish- Edinburgh EH4 2XU. ing reliable dosage studies for a number of different probes has proved difficult. We have therefore WAGR(Wilms tumour, aniridia, genitourinary abnormalities and mental retardation) syndrome concentrated on segregating the deleted chromo- is frequently associated with deletions on the short some 11 from a number of patients in somatic cell arm of chromosome 11. The deletions vary in size hybrids and analysing DNA from these to produce but always include part of band lipl3. To home a consistent map of chromosome lip. At the same in on the Wilms tumour and aniridia loci the end time we have determined the deletion breakpoints points of the different deletion breakpoints need at a molecular level and shown that the results are to be defined at the DNA level. -

Next Generation Sequencing Panels for Disorders of Sex Development
Next Generation Sequencing Panels for Disorders of Sex Development Disorders of Sex Development – Overview Disorders of sex development (DSDs) occur when sex development does not follow the course of typical male or female patterning. Types of DSDs include congenital development of ambiguous genitalia, disjunction between the internal and external sex anatomy, incomplete development of the sex anatomy, and abnormalities of the development of gonads (such as ovotestes or streak ovaries) (1). Sex chromosome anomalies including Turner syndrome and Klinefelter syndrome as well as sex chromosome mosaicism are also considered to be DSDs. DSDs can be caused by a wide range of genetic abnormalities (2). Determining the etiology of a patient’s DSD can assist in deciding gender assignment, provide recurrence risk information for future pregnancies, and can identify potential health problems such as adrenal crisis or gonadoblastoma (1, 3). Sex chromosome aneuploidy and copy number variation are common genetic causes of DSDs. For this reason, chromosome analysis and/or microarray analysis typically should be the first genetic analysis in the case of a patient with ambiguous genitalia or other suspected disorder of sex development. Identifying whether a patient has a 46,XY or 46,XX karyotype can also be helpful in determining appropriate additional genetic testing. Abnormal/Ambiguous Genitalia Panel Our Abnormal/Ambiguous Genitalia Panel includes mutation analysis of 72 genes associated with both syndromic and non-syndromic DSDs. This comprehensive panel evaluates a broad range of genetic causes of ambiguous or abnormal genitalia, including conditions in which abnormal genitalia are the primary physical finding as well as syndromic conditions that involve abnormal genitalia in addition to other congenital anomalies. -

Nucleotide Excision Repair Gene ERCC2 and ERCC5 Variants Increase Risk of Uterine Cervical Cancer
pISSN 1598-2998, eISSN 2005-9256 Cancer Res Treat. 2016;48(2):708-714 http://dx.doi.org/10.4143/crt.2015.098 Original Article Open Access Nucleotide Excision Repair Gene ERCC2 and ERCC5 Variants Increase Risk of Uterine Cervical Cancer Jungnam Joo, PhD 1, Kyong-Ah Yoon, PhD 2, Tomonori Hayashi, PhD 3, Sun-Young Kong, MD, PhD 4, Hye-Jin Shin, MS 5, Boram Park, MS 1, Young Min Kim, PhD 6, Sang-Hyun Hwang, MD, PhD 7, Jeongseon Kim, PhD 8, Aesun Shin, MD, PhD 8,9 , Joo-Young Kim, MD, PhD 5,10 1Biometric Research Branch, 2Lung Cancer Branch, National Cancer Center, Goyang, Korea, 3Department of Radiobiology and Molecular Epidemiology, Radiation Effects Research Foundation, Hiroshima, Japan, 4Translational Epidemiology Research Branch and Department of Laboratory Medicine, 5Radiation Medicine Branch, National Cancer Center, Goyang, Korea, 6Department of Statistics, Radiation Effects Research Foundation, Hiroshima, Japan, 7Hematologic Malignancy Branch and Department of Laboratory Medicine, 8Molecular Epidemiology Branch, National Cancer Center, Goyang, 9Department of Preventive Medicine, Seoul National University College of Medicine, Seoul, 10 Center for Proton Therapy, Research Institute and Hospital, National Cancer Center, Goyang, Korea Supplementary Data Table of Contents Supplementary Fig. S1 ........................................................................................................................................................................... 2 Supplementary Table 1 ......................................................................................................................................................................... -

Genetic Mechanisms of Disease in Children: a New Look
Genetic Mechanisms of Disease in Children: A New Look Laurie Demmer, MD Tufts Medical Center and the Floating Hospital for Children Boston, MA Traditionally genetic disorders have been linked to the ‘one gene-one protein-one disease’ hypothesis. However recent advances in the field of molecular biology and biotechnology have afforded us the opportunity to greatly expand our knowledge of genetics, and we now know that the mechanisms of inherited disorders are often significantly more complex, and consequently, much more intriguing, than originally thought. Classical mendelian disorders with relatively simple genetic mechanisms do exist, but turn out to be far more rare than originally thought. All patients with sickle cell disease for example, carry the same A-to-T point mutation in the sixth codon of the beta globin gene. This results in a glutamate to valine substitution which changes the shape and the function of the globin molecule in a predictable way. Similarly, all patients with achondroplasia have a single base pair substitution at nucleotide #1138 of the FGFR3 gene. On the other hand, another common inherited disorder, cystic fibrosis, is known to result from changes in a specific transmembrane receptor (CFTR), but over 1000 different disease-causing mutations have been reported in this single gene. Since most commercial labs only test for between 23-100 different mutations, interpreting CFTR mutation testing is significantly complicated by the known risk of false negative results. Many examples of complex, or non-Mendelian, inheritance are now known to exist and include disorders of trinucleotide repeats, errors in imprinting, and gene dosage effects. -

Open Full Page
CCR PEDIATRIC ONCOLOGY SERIES CCR Pediatric Oncology Series Recommendations for Childhood Cancer Screening and Surveillance in DNA Repair Disorders Michael F. Walsh1, Vivian Y. Chang2, Wendy K. Kohlmann3, Hamish S. Scott4, Christopher Cunniff5, Franck Bourdeaut6, Jan J. Molenaar7, Christopher C. Porter8, John T. Sandlund9, Sharon E. Plon10, Lisa L. Wang10, and Sharon A. Savage11 Abstract DNA repair syndromes are heterogeneous disorders caused by around the world to discuss and develop cancer surveillance pathogenic variants in genes encoding proteins key in DNA guidelines for children with cancer-prone disorders. Herein, replication and/or the cellular response to DNA damage. The we focus on the more common of the rare DNA repair dis- majority of these syndromes are inherited in an autosomal- orders: ataxia telangiectasia, Bloom syndrome, Fanconi ane- recessive manner, but autosomal-dominant and X-linked reces- mia, dyskeratosis congenita, Nijmegen breakage syndrome, sive disorders also exist. The clinical features of patients with DNA Rothmund–Thomson syndrome, and Xeroderma pigmento- repair syndromes are highly varied and dependent on the under- sum. Dedicated syndrome registries and a combination of lying genetic cause. Notably, all patients have elevated risks of basic science and clinical research have led to important in- syndrome-associated cancers, and many of these cancers present sights into the underlying biology of these disorders. Given the in childhood. Although it is clear that the risk of cancer is rarity of these disorders, it is recommended that centralized increased, there are limited data defining the true incidence of centers of excellence be involved directly or through consulta- cancer and almost no evidence-based approaches to cancer tion in caring for patients with heritable DNA repair syn- surveillance in patients with DNA repair disorders. -

Seq2pathway Vignette
seq2pathway Vignette Bin Wang, Xinan Holly Yang, Arjun Kinstlick May 19, 2021 Contents 1 Abstract 1 2 Package Installation 2 3 runseq2pathway 2 4 Two main functions 3 4.1 seq2gene . .3 4.1.1 seq2gene flowchart . .3 4.1.2 runseq2gene inputs/parameters . .5 4.1.3 runseq2gene outputs . .8 4.2 gene2pathway . 10 4.2.1 gene2pathway flowchart . 11 4.2.2 gene2pathway test inputs/parameters . 11 4.2.3 gene2pathway test outputs . 12 5 Examples 13 5.1 ChIP-seq data analysis . 13 5.1.1 Map ChIP-seq enriched peaks to genes using runseq2gene .................... 13 5.1.2 Discover enriched GO terms using gene2pathway_test with gene scores . 15 5.1.3 Discover enriched GO terms using Fisher's Exact test without gene scores . 17 5.1.4 Add description for genes . 20 5.2 RNA-seq data analysis . 20 6 R environment session 23 1 Abstract Seq2pathway is a novel computational tool to analyze functional gene-sets (including signaling pathways) using variable next-generation sequencing data[1]. Integral to this tool are the \seq2gene" and \gene2pathway" components in series that infer a quantitative pathway-level profile for each sample. The seq2gene function assigns phenotype-associated significance of genomic regions to gene-level scores, where the significance could be p-values of SNPs or point mutations, protein-binding affinity, or transcriptional expression level. The seq2gene function has the feasibility to assign non-exon regions to a range of neighboring genes besides the nearest one, thus facilitating the study of functional non-coding elements[2]. Then the gene2pathway summarizes gene-level measurements to pathway-level scores, comparing the quantity of significance for gene members within a pathway with those outside a pathway. -

Autism Multiplex Family with 16P11.2P12.2 Microduplication Syndrome in Monozygotic Twins and Distal 16P11.2 Deletion in Their Brother
European Journal of Human Genetics (2012) 20, 540–546 & 2012 Macmillan Publishers Limited All rights reserved 1018-4813/12 www.nature.com/ejhg ARTICLE Autism multiplex family with 16p11.2p12.2 microduplication syndrome in monozygotic twins and distal 16p11.2 deletion in their brother Anne-Claude Tabet1,2,3,4, Marion Pilorge2,3,4, Richard Delorme5,6,Fre´de´rique Amsellem5,6, Jean-Marc Pinard7, Marion Leboyer6,8,9, Alain Verloes10, Brigitte Benzacken1,11,12 and Catalina Betancur*,2,3,4 The pericentromeric region of chromosome 16p is rich in segmental duplications that predispose to rearrangements through non-allelic homologous recombination. Several recurrent copy number variations have been described recently in chromosome 16p. 16p11.2 rearrangements (29.5–30.1 Mb) are associated with autism, intellectual disability (ID) and other neurodevelopmental disorders. Another recognizable but less common microdeletion syndrome in 16p11.2p12.2 (21.4 to 28.5–30.1 Mb) has been described in six individuals with ID, whereas apparently reciprocal duplications, studied by standard cytogenetic and fluorescence in situ hybridization techniques, have been reported in three patients with autism spectrum disorders. Here, we report a multiplex family with three boys affected with autism, including two monozygotic twins carrying a de novo 16p11.2p12.2 duplication of 8.95 Mb (21.28–30.23 Mb) characterized by single-nucleotide polymorphism array, encompassing both the 16p11.2 and 16p11.2p12.2 regions. The twins exhibited autism, severe ID, and dysmorphic features, including a triangular face, deep-set eyes, large and prominent nasal bridge, and tall, slender build. The eldest brother presented with autism, mild ID, early-onset obesity and normal craniofacial features, and carried a smaller, overlapping 16p11.2 microdeletion of 847 kb (28.40–29.25 Mb), inherited from his apparently healthy father. -

ARID1B Is a Specific Vulnerability in ARID1A-Mutant Cancers The
ARID1B is a specific vulnerability in ARID1A-mutant cancers The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters. Citation Helming, K. C., X. Wang, B. G. Wilson, F. Vazquez, J. R. Haswell, H. E. Manchester, Y. Kim, et al. 2014. “ARID1B is a specific vulnerability in ARID1A-mutant cancers.” Nature medicine 20 (3): 251-254. doi:10.1038/nm.3480. http://dx.doi.org/10.1038/nm.3480. Published Version doi:10.1038/nm.3480 Accessed February 16, 2015 10:04:32 PM EST Citable Link http://nrs.harvard.edu/urn-3:HUL.InstRepos:12987227 Terms of Use This article was downloaded from Harvard University's DASH repository, and is made available under the terms and conditions applicable to Other Posted Material, as set forth at http://nrs.harvard.edu/urn-3:HUL.InstRepos:dash.current.terms-of- use#LAA (Article begins on next page) NIH Public Access Author Manuscript Nat Med. Author manuscript; available in PMC 2014 September 01. NIH-PA Author ManuscriptPublished NIH-PA Author Manuscript in final edited NIH-PA Author Manuscript form as: Nat Med. 2014 March ; 20(3): 251–254. doi:10.1038/nm.3480. ARID1B is a specific vulnerability in ARID1A-mutant cancers Katherine C. Helming1,2,3,4,*, Xiaofeng Wang1,2,3,*, Boris G. Wilson1,2,3, Francisca Vazquez5, Jeffrey R. Haswell1,2,3, Haley E. Manchester1,2,3, Youngha Kim1,2,3, Gregory V. Kryukov5, Mahmoud Ghandi5, Andrew J. Aguirre5,6,7, Zainab Jagani8, Zhong Wang9, Levi A. Garraway6, William C. Hahn6,7, and Charles W. -

The Genetic Basis for Skeletal Diseases
insight review articles The genetic basis for skeletal diseases Elazar Zelzer & Bjorn R. Olsen Harvard Medical School, Department of Cell Biology, 240 Longwood Avenue, Boston, Massachusetts 02115, USA (e-mail: [email protected]) We walk, run, work and play, paying little attention to our bones, their joints and their muscle connections, because the system works. Evolution has refined robust genetic mechanisms for skeletal development and growth that are able to direct the formation of a complex, yet wonderfully adaptable organ system. How is it done? Recent studies of rare genetic diseases have identified many of the critical transcription factors and signalling pathways specifying the normal development of bones, confirming the wisdom of William Harvey when he said: “nature is nowhere accustomed more openly to display her secret mysteries than in cases where she shows traces of her workings apart from the beaten path”. enetic studies of diseases that affect skeletal differentiation to cartilage cells (chondrocytes) or bone cells development and growth are providing (osteoblasts) within the condensations. Subsequent growth invaluable insights into the roles not only of during the organogenesis phase generates cartilage models individual genes, but also of entire (anlagen) of future bones (as in limb bones) or membranous developmental pathways. Different mutations bones (as in the cranial vault) (Fig. 1). The cartilage anlagen Gin the same gene may result in a range of abnormalities, are replaced by bone and marrow in a process called endo- and disease ‘families’ are frequently caused by mutations in chondral ossification. Finally, a process of growth and components of the same pathway. -
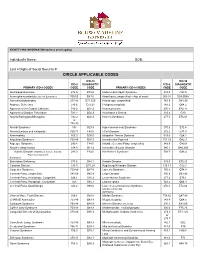
Circle Applicable Codes
IDENTIFYING INFORMATION (please print legibly) Individual’s Name: DOB: Last 4 Digits of Social Security #: CIRCLE APPLICABLE CODES ICD-10 ICD-10 ICD-9 DIAGNOSTIC ICD-9 DIAGNOSTIC PRIMARY ICD-9 CODES CODE CODE PRIMARY ICD-9 CODES CODE CODE Abetalipoproteinemia 272.5 E78.6 Hallervorden-Spatz Syndrome 333.0 G23.0 Acrocephalosyndactyly (Apert’s Syndrome) 755.55 Q87.0 Head Injury, unspecified – Age of onset: 959.01 S09.90XA Adrenaleukodystrophy 277.86 E71.529 Hemiplegia, unspecified 342.9 G81.90 Arginase Deficiency 270.6 E72.21 Holoprosencephaly 742.2 Q04.2 Agenesis of the Corpus Callosum 742.2 Q04.3 Homocystinuria 270.4 E72.11 Agenesis of Septum Pellucidum 742.2 Q04.3 Huntington’s Chorea 333.4 G10 Argyria/Pachygyria/Microgyria 742.2 Q04.3 Hurler’s Syndrome 277.5 E76.01 or 758.33 Aicardi Syndrome 333 G23.8 Hyperammonemia Syndrome 270.6 E72.4 Alcohol Embryo and Fetopathy 760.71 F84.5 I-Cell Disease 272.2 E77.0 Anencephaly 655.0 Q00.0 Idiopathic Torsion Dystonia 333.6 G24.1 Angelman Syndrome 759.89 Q93.5 Incontinentia Pigmenti 757.33 Q82.3 Asperger Syndrome 299.8 F84.5 Infantile Cerebral Palsy, unspecified 343.9 G80.9 Ataxia-Telangiectasia 334.8 G11.3 Intractable Seizure Disorder 345.1 G40.309 Autistic Disorder (Childhood Autism, Infantile 299.0 F84.0 Klinefelter’s Syndrome 758.7 Q98.4 Psychosis, Kanner’s Syndrome) Biotinidase Deficiency 277.6 D84.1 Krabbe Disease 333.0 E75.23 Canavan Disease 330.0 E75.29 Kugelberg-Welander Disease 335.11 G12.1 Carpenter Syndrome 759.89 Q87.0 Larsen’s Syndrome 755.8 Q74.8 Cerebral Palsy, unspecified 343.69 G80.9