ARID1B Is a Specific Vulnerability in ARID1A-Mutant Cancers The
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Genetic Screening Identifies a Component of the SWI/SNF Complex, Arid1b As a Senescence Regulator
A genetic screening identifies a component of the SWI/SNF complex, Arid1b as a senescence regulator Sadaf Khan A thesis submitted to Imperial College London for the degree of Doctor in Philosophy MRC Clinical Sciences Centre Imperial College London, School of Medicine July 2013 Statement of originality All experiments included in this thesis were performed by myself unless otherwise stated. Copyright Declaration The copyright of this thesis rests with the author and is made available under a Creative Commons Attribution Non-Commercial No Derivatives license. Researchers are free to copy, distribute or transmit the thesis on the condition that they attribute it, that they do not use it for commercial purposes and that they do not alter, transform or build upon it. For any reuse or redistribution, researchers must make clear to others the license terms of this work. 2 Abstract Senescence is an important tumour suppressor mechanism, which prevents the proliferation of stressed or damaged cells. The use of RNA interference to identify genes with a role in senescence is an important tool in the discovery of novel cancer genes. In this work, a protocol was established for conducting bypass of senescence screenings, using shRNA libraries together with next-generation sequencing. Using this approach, the SWI/SNF subunit Arid1b was identified as a regulator of cellular lifespan in MEFs. SWI/SNF is a large multi-subunit complex that remodels chromatin. Mutations in SWI/SNF proteins are frequently associated with cancer, suggesting that SWI/SNF components are tumour suppressors. Here the role of ARID1B during senescence was investigated. Depletion of ARID1B extends the proliferative capacity of primary mouse and human fibroblasts. -

The Genetic Basis of Hepatosplenic T-Cell Lymphoma
Published OnlineFirst January 25, 2017; DOI: 10.1158/2159-8290.CD-16-0330 RESEARCH BRIEF The Genetic Basis of Hepatosplenic T-cell Lymphoma Matthew McKinney 1 , Andrea B. Moffi tt 2 , Philippe Gaulard 3 , Marion Travert 3 , Laurence De Leval4 , Alina Nicolae 5 , Mark Raffeld 5 , Elaine S. Jaffe 5 , Stefania Pittaluga 5 , Liqiang Xi 5 , Tayla Heavican 6 , Javeed Iqbal 6 , Karim Belhadj 3 , Marie Helene Delfau-Larue 3 , Virginie Fataccioli 3 , Magdalena B. Czader7 , Izidore S. Lossos 8 , Jennifer R. Chapman-Fredricks 8 , Kristy L. Richards 9 , Yuri Fedoriw 9 , Sarah L. Ondrejka10 , Eric D. Hsi 10 , Lawrence Low 11 , Dennis Weisenburger 11 , Wing C. Chan 11 , Neha Mehta-Shah12 , Steven Horwitz 12 , Leon Bernal-Mizrachi 13 , Christopher R. Flowers 13 , Anne W. Beaven 1 , Mayur Parihar 14 , Lucile Baseggio 15 , Marie Parrens 16 , Anne Moreau 17 , Pierre Sujobert 18 , Monika Pilichowska 19 , Andrew M. Evens 19 , Amy Chadburn 20 , Rex K.H. Au-Yeung 21 , Gopesh Srivastava 21 , William W. L. Choi 21 , John R. Goodlad 22 , Igor Aurer 23 , Sandra Basic-Kinda 23 , Randy D. Gascoyne 24 , Nicholas S. Davis 1 , Guojie Li 1 , Jenny Zhang 1 , Deepthi Rajagopalan 1 , Anupama Reddy 1 , Cassandra Love 1 , Shawn Levy 25 , Yuan Zhuang 1 , Jyotishka Datta 26 , David B. Dunson 26 , and Sandeep S. Davé 1 , 2 ABSTRACT Hepatosplenic T-cell lymphoma (HSTL) is a rare and lethal lymphoma; the genetic drivers of this disease are unknown. Through whole-exome sequencing of 68 HSTLs, we defi ne recurrently mutated driver genes and copy-number alterations in the disease. -

MECHANISMS in ENDOCRINOLOGY: Novel Genetic Causes of Short Stature
J M Wit and others Genetics of short stature 174:4 R145–R173 Review MECHANISMS IN ENDOCRINOLOGY Novel genetic causes of short stature 1 1 2 2 Jan M Wit , Wilma Oostdijk , Monique Losekoot , Hermine A van Duyvenvoorde , Correspondence Claudia A L Ruivenkamp2 and Sarina G Kant2 should be addressed to J M Wit Departments of 1Paediatrics and 2Clinical Genetics, Leiden University Medical Center, PO Box 9600, 2300 RC Leiden, Email The Netherlands [email protected] Abstract The fast technological development, particularly single nucleotide polymorphism array, array-comparative genomic hybridization, and whole exome sequencing, has led to the discovery of many novel genetic causes of growth failure. In this review we discuss a selection of these, according to a diagnostic classification centred on the epiphyseal growth plate. We successively discuss disorders in hormone signalling, paracrine factors, matrix molecules, intracellular pathways, and fundamental cellular processes, followed by chromosomal aberrations including copy number variants (CNVs) and imprinting disorders associated with short stature. Many novel causes of GH deficiency (GHD) as part of combined pituitary hormone deficiency have been uncovered. The most frequent genetic causes of isolated GHD are GH1 and GHRHR defects, but several novel causes have recently been found, such as GHSR, RNPC3, and IFT172 mutations. Besides well-defined causes of GH insensitivity (GHR, STAT5B, IGFALS, IGF1 defects), disorders of NFkB signalling, STAT3 and IGF2 have recently been discovered. Heterozygous IGF1R defects are a relatively frequent cause of prenatal and postnatal growth retardation. TRHA mutations cause a syndromic form of short stature with elevated T3/T4 ratio. Disorders of signalling of various paracrine factors (FGFs, BMPs, WNTs, PTHrP/IHH, and CNP/NPR2) or genetic defects affecting cartilage extracellular matrix usually cause disproportionate short stature. -

1714 Gene Comprehensive Cancer Panel Enriched for Clinically Actionable Genes with Additional Biologically Relevant Genes 400-500X Average Coverage on Tumor
xO GENE PANEL 1714 gene comprehensive cancer panel enriched for clinically actionable genes with additional biologically relevant genes 400-500x average coverage on tumor Genes A-C Genes D-F Genes G-I Genes J-L AATK ATAD2B BTG1 CDH7 CREM DACH1 EPHA1 FES G6PC3 HGF IL18RAP JADE1 LMO1 ABCA1 ATF1 BTG2 CDK1 CRHR1 DACH2 EPHA2 FEV G6PD HIF1A IL1R1 JAK1 LMO2 ABCB1 ATM BTG3 CDK10 CRK DAXX EPHA3 FGF1 GAB1 HIF1AN IL1R2 JAK2 LMO7 ABCB11 ATR BTK CDK11A CRKL DBH EPHA4 FGF10 GAB2 HIST1H1E IL1RAP JAK3 LMTK2 ABCB4 ATRX BTRC CDK11B CRLF2 DCC EPHA5 FGF11 GABPA HIST1H3B IL20RA JARID2 LMTK3 ABCC1 AURKA BUB1 CDK12 CRTC1 DCUN1D1 EPHA6 FGF12 GALNT12 HIST1H4E IL20RB JAZF1 LPHN2 ABCC2 AURKB BUB1B CDK13 CRTC2 DCUN1D2 EPHA7 FGF13 GATA1 HLA-A IL21R JMJD1C LPHN3 ABCG1 AURKC BUB3 CDK14 CRTC3 DDB2 EPHA8 FGF14 GATA2 HLA-B IL22RA1 JMJD4 LPP ABCG2 AXIN1 C11orf30 CDK15 CSF1 DDIT3 EPHB1 FGF16 GATA3 HLF IL22RA2 JMJD6 LRP1B ABI1 AXIN2 CACNA1C CDK16 CSF1R DDR1 EPHB2 FGF17 GATA5 HLTF IL23R JMJD7 LRP5 ABL1 AXL CACNA1S CDK17 CSF2RA DDR2 EPHB3 FGF18 GATA6 HMGA1 IL2RA JMJD8 LRP6 ABL2 B2M CACNB2 CDK18 CSF2RB DDX3X EPHB4 FGF19 GDNF HMGA2 IL2RB JUN LRRK2 ACE BABAM1 CADM2 CDK19 CSF3R DDX5 EPHB6 FGF2 GFI1 HMGCR IL2RG JUNB LSM1 ACSL6 BACH1 CALR CDK2 CSK DDX6 EPOR FGF20 GFI1B HNF1A IL3 JUND LTK ACTA2 BACH2 CAMTA1 CDK20 CSNK1D DEK ERBB2 FGF21 GFRA4 HNF1B IL3RA JUP LYL1 ACTC1 BAG4 CAPRIN2 CDK3 CSNK1E DHFR ERBB3 FGF22 GGCX HNRNPA3 IL4R KAT2A LYN ACVR1 BAI3 CARD10 CDK4 CTCF DHH ERBB4 FGF23 GHR HOXA10 IL5RA KAT2B LZTR1 ACVR1B BAP1 CARD11 CDK5 CTCFL DIAPH1 ERCC1 FGF3 GID4 HOXA11 IL6R KAT5 ACVR2A -

Genome-Wide DNA Methylation Analysis Reveals Molecular Subtypes of Pancreatic Cancer
www.impactjournals.com/oncotarget/ Oncotarget, 2017, Vol. 8, (No. 17), pp: 28990-29012 Research Paper Genome-wide DNA methylation analysis reveals molecular subtypes of pancreatic cancer Nitish Kumar Mishra1 and Chittibabu Guda1,2,3,4 1Department of Genetics, Cell Biology and Anatomy, University of Nebraska Medical Center, Omaha, NE, 68198, USA 2Bioinformatics and Systems Biology Core, University of Nebraska Medical Center, Omaha, NE, 68198, USA 3Department of Biochemistry and Molecular Biology, University of Nebraska Medical Center, Omaha, NE, 68198, USA 4Fred and Pamela Buffet Cancer Center, University of Nebraska Medical Center, Omaha, NE, 68198, USA Correspondence to: Chittibabu Guda, email: [email protected] Keywords: TCGA, pancreatic cancer, differential methylation, integrative analysis, molecular subtypes Received: October 20, 2016 Accepted: February 12, 2017 Published: March 07, 2017 Copyright: Mishra et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC-BY), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. ABSTRACT Pancreatic cancer (PC) is the fourth leading cause of cancer deaths in the United States with a five-year patient survival rate of only 6%. Early detection and treatment of this disease is hampered due to lack of reliable diagnostic and prognostic markers. Recent studies have shown that dynamic changes in the global DNA methylation and gene expression patterns play key roles in the PC development; hence, provide valuable insights for better understanding the initiation and progression of PC. In the current study, we used DNA methylation, gene expression, copy number, mutational and clinical data from pancreatic patients. -

MDM4 Is Targeted by 1Q Gain and Drives Disease in Burkitt Lymphoma
Published OnlineFirst April 18, 2019; DOI: 10.1158/0008-5472.CAN-18-3438 Cancer Translational Science Research MDM4 Is Targeted by 1q Gain and Drives Disease in Burkitt Lymphoma Jennifer Hullein€ 1,2, Mikołaj Słabicki1, Maciej Rosolowski3, Alexander Jethwa1,2, Stefan Habringer4, Katarzyna Tomska1, Roma Kurilov5, Junyan Lu6, Sebastian Scheinost1, Rabea Wagener7,8, Zhiqin Huang9, Marina Lukas1, Olena Yavorska6, Hanne Helfrich10,Rene Scholtysik11, Kyle Bonneau12, Donato Tedesco12,RalfKuppers€ 11, Wolfram Klapper13, Christiane Pott14, Stephan Stilgenbauer10, Birgit Burkhardt15, Markus Lof€ fler3, Lorenz H. Trumper€ 16, Michael Hummel17, Benedikt Brors5, Marc Zapatka9, Reiner Siebert7,8, Markus Kreuz3, Ulrich Keller4,18, Wolfgang Huber6, and Thorsten Zenz1,19 Abstract Oncogenic MYC activation promotes proliferation in growth in a xenograft model in a p53-dependent manner. Burkitt lymphoma, but also induces cell-cycle arrest and Small molecule inhibition of the MDM4–p53 interaction apoptosis mediated by p53, a tumor suppressor that is was effective only in TP53wt Burkitt lymphoma cell lines. mutated in 40% of Burkitt lymphoma cases. To identify Moreover, primary TP53wt Burkitt lymphoma samples fre- molecular dependencies in Burkitt lymphoma, we per- quently acquired gains of chromosome 1q, which includes formed RNAi-based, loss-of-function screening in eight the MDM4 locus, and showed elevated MDM4 mRNA levels. Burkitt lymphoma cell lines and integrated non-Burkitt 1q gain was associated with TP53wt across 789 cancer cell lymphoma RNAi screens and genetic data. We identified lines and MDM4 was essential in the TP53wt-context in 216 76 genes essential to Burkitt lymphoma, including genes cell lines representing 19 cancer entities from the Achilles associated with hematopoietic cell differentiation (FLI1, Project. -
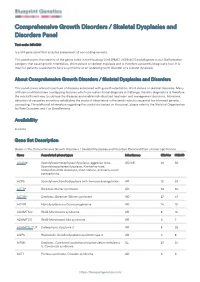
Blueprint Genetics Comprehensive Growth Disorders / Skeletal
Comprehensive Growth Disorders / Skeletal Dysplasias and Disorders Panel Test code: MA4301 Is a 374 gene panel that includes assessment of non-coding variants. This panel covers the majority of the genes listed in the Nosology 2015 (PMID: 26394607) and all genes in our Malformation category that cause growth retardation, short stature or skeletal dysplasia and is therefore a powerful diagnostic tool. It is ideal for patients suspected to have a syndromic or an isolated growth disorder or a skeletal dysplasia. About Comprehensive Growth Disorders / Skeletal Dysplasias and Disorders This panel covers a broad spectrum of diseases associated with growth retardation, short stature or skeletal dysplasia. Many of these conditions have overlapping features which can make clinical diagnosis a challenge. Genetic diagnostics is therefore the most efficient way to subtype the diseases and enable individualized treatment and management decisions. Moreover, detection of causative mutations establishes the mode of inheritance in the family which is essential for informed genetic counseling. For additional information regarding the conditions tested on this panel, please refer to the National Organization for Rare Disorders and / or GeneReviews. Availability 4 weeks Gene Set Description Genes in the Comprehensive Growth Disorders / Skeletal Dysplasias and Disorders Panel and their clinical significance Gene Associated phenotypes Inheritance ClinVar HGMD ACAN# Spondyloepimetaphyseal dysplasia, aggrecan type, AD/AR 20 56 Spondyloepiphyseal dysplasia, Kimberley -

Supplementary Table 1A. Genes Significantly Altered in A4573 ESFT
Supplementary Table 1A. Genes significantly altered in A4573 ESFT cells following BMI-1knockdown genesymbol genedescription siControl siBMI1 FC Direction P-value AASS aminoadipate-semialdehyde synthase | tetra-peptide repeat homeobox-like6.68 7.24 1.5 Up 0.007 ABCA2 ATP-binding cassette, sub-family A (ABC1), member 2 | neural5.44 proliferation,6.3 differentiation1.8 and Upcontrol, 1 0.006 ABHD4 abhydrolase domain containing 4 7.51 6.69 1.8 Down 0.002 ACACA acetyl-Coenzyme A carboxylase alpha | peroxiredoxin 5 | similar6.2 to High mobility7.26 group2.1 protein UpB1 (High mobility0.009 group protein 1) (HMG-1) (Amphoterin) (Heparin-binding protein p30) | Coenzyme A synthase ACAD9 acyl-Coenzyme A dehydrogenase family, member 9 9.25 8.59 1.6 Down 0.008 ACBD3 acyl-Coenzyme A binding domain containing 3 7.89 8.53 1.6 Up 0.008 ACCN2 amiloride-sensitive cation channel 2, neuronal 5.47 6.28 1.8 Up 0.005 ACIN1 apoptotic chromatin condensation inducer 1 7.15 7.79 1.6 Up 0.008 ACPL2 acid phosphatase-like 2 6.04 7.6 2.9 Up 0.000 ACSL4 acyl-CoA synthetase long-chain family member 4 6.72 5.8 1.9 Down 0.001 ACTA2 actin, alpha 2, smooth muscle, aorta 9.18 8.44 1.7 Down 0.003 ACYP1 acylphosphatase 1, erythrocyte (common) type 7.09 7.66 1.5 Up 0.009 ADA adenosine deaminase 6.34 7.1 1.7 Up 0.009 ADAL adenosine deaminase-like 7.88 6.89 2.0 Down 0.006 ADAMTS1 ADAM metallopeptidase with thrombospondin type 1 motif, 1 6.57 7.65 2.1 Up 0.000 ADARB1 adenosine deaminase, RNA-specific, B1 (RED1 homolog rat) 6.49 7.13 1.6 Up 0.008 ADCY9 adenylate cyclase 9 6.5 7.18 -

Extreme Intratumour Heterogeneity and Driver Evolution in Mismatch Repair Deficient Gastro- Oesophageal Cancer
ARTICLE Corrected: Author correction https://doi.org/10.1038/s41467-019-13915-7 OPEN Extreme intratumour heterogeneity and driver evolution in mismatch repair deficient gastro- oesophageal cancer Katharina von Loga 1,6,8, Andrew Woolston 1,8, Marco Punta2, Louise J. Barber1, Beatrice Griffiths1, Maria Semiannikova1, Georgia Spain1, Benjamin Challoner 1, Kerry Fenwick3, Ronald Simon4, Andreas Marx4,7, Guido Sauter4, Stefano Lise 2, Nik Matthews3 & Marco Gerlinger 1,5* 1234567890():,; Mismatch repair deficient (dMMR) gastro-oesophageal adenocarcinomas (GOAs) show better outcomes than their MMR-proficient counterparts and high immunotherapy sensi- tivity. The hypermutator-phenotype of dMMR tumours theoretically enables high evolvability but their evolution has not been investigated. Here we apply multi-region exome sequencing (MSeq) to four treatment-naive dMMR GOAs. This reveals extreme intratumour hetero- geneity (ITH), exceeding ITH in other cancer types >20-fold, but also long phylogenetic trunks which may explain the exquisite immunotherapy sensitivity of dMMR tumours. Sub- clonal driver mutations are common and parallel evolution occurs in RAS, PIK3CA, SWI/SNF- complex genes and in immune evasion regulators. MSeq data and evolution analysis of single region-data from 64 MSI GOAs show that chromosome 8 gains are early genetic events and that the hypermutator-phenotype remains active during progression. MSeq may be necessary for biomarker development in these heterogeneous cancers. Comparison with other MSeq- analysed tumour types reveals mutation rates and their timing to determine phylogenetic tree morphologies. 1 Translational Oncogenomics Laboratory, Centre for Evolution and Cancer, The Institute of Cancer Research, London SW3 6JB, United Kingdom. 2 Bioinformatics Core, Centre for Evolution and Cancer, The Institute of Cancer Research, London SM2 5NG, United Kingdom. -

ARID1A Mutations in Cancer: Another Epigenetic Tumor Suppressor?
Published OnlineFirst December 3, 2012; DOI: 10.1158/2159-8290.CD-12-0361 MINI REVIEW ARID1A Mutations in Cancer: Another Epigenetic Tumor Suppressor? Jennifer N. Wu and Charles W.M. Roberts ABSTRACT Although disordered chromatin organization has long been recognized as a fea- ture of cancer, the molecular underpinnings of chromatin structure, epigenetic regulation, and their relationships to transcription are only beginning to be understood. Cancer genome sequencing studies have revealed a novel theme: frequent mutation of epigenetic regulators. Among these, the ARID1A/BAF250A subunit of the SWI/SNF (BRG1-associated factors) chromatin remod- eling complex has emerged as recurrently mutated in a broad array of tumor types. We review the genomic and functional data supporting classifi cation of ARID1A as a tumor suppressor. Signifi cance: Mutations in chromatin remodeling complex genes are increasingly recognized in many cancer types. However, the mechanisms by which chromatin remodeling complexes contribute to gene expression and the cancer phenotype are poorly understood. Understanding how mutation of chroma- tin remodelers facilitates transformation may offer the potential for development and implementation of novel therapies for cancer. Cancer Discov; 3(1); 35–43. ©2012 AACR. INTRODUCTION risk of developing malignant rhabdoid tumors at an especially young age. Epigenetic regulators impose upon the genetic code a Subsequent to these discoveries, implementation of next- chromatin structure characterized by chromatin accessibil- generation sequencing technologies have revealed frequent ity, nucleosome position, and histone modifi cations. These and recurrent mutations in a wide variety of epigenetic features of chromatin structure modulate gene expres- modulators, including mediators of DNA methylation (e.g., sion, thereby affecting both the identity and function of DNMT3a) and covalent modifi ers of histones (e.g., MLL, MLL3, cells. -

Perkinelmer Genomics to Request the Saliva Swab Collection Kit for Patients That Cannot Provide a Blood Sample As Whole Blood Is the Preferred Sample
Autism and Intellectual Disability TRIO Panel Test Code TR002 Test Summary This test analyzes 2429 genes that have been associated with Autism and Intellectual Disability and/or disorders associated with Autism and Intellectual Disability with the analysis being performed as a TRIO Turn-Around-Time (TAT)* 3 - 5 weeks Acceptable Sample Types Whole Blood (EDTA) (Preferred sample type) DNA, Isolated Dried Blood Spots Saliva Acceptable Billing Types Self (patient) Payment Institutional Billing Commercial Insurance Indications for Testing Comprehensive test for patients with intellectual disability or global developmental delays (Moeschler et al 2014 PMID: 25157020). Comprehensive test for individuals with multiple congenital anomalies (Miller et al. 2010 PMID 20466091). Patients with autism/autism spectrum disorders (ASDs). Suspected autosomal recessive condition due to close familial relations Previously negative karyotyping and/or chromosomal microarray results. Test Description This panel analyzes 2429 genes that have been associated with Autism and ID and/or disorders associated with Autism and ID. Both sequencing and deletion/duplication (CNV) analysis will be performed on the coding regions of all genes included (unless otherwise marked). All analysis is performed utilizing Next Generation Sequencing (NGS) technology. CNV analysis is designed to detect the majority of deletions and duplications of three exons or greater in size. Smaller CNV events may also be detected and reported, but additional follow-up testing is recommended if a smaller CNV is suspected. All variants are classified according to ACMG guidelines. Condition Description Autism Spectrum Disorder (ASD) refers to a group of developmental disabilities that are typically associated with challenges of varying severity in the areas of social interaction, communication, and repetitive/restricted behaviors. -

SUPPLEMENTARY APPENDIX Derepression of Retroelements in Acute Myeloid Leukemia with 3Q Aberrations
SUPPLEMENTARY APPENDIX Derepression of retroelements in acute myeloid leukemia with 3q aberrations Jagoda Mika, 1 Sophie Ottema, 2 Sandra Kiehlmeier, 1 Sabrina Kruse, 1 Leonie Smeenk, 2 Judith Müller, 1 Sabrina Schweiggert, 1 Carl Herrmann, 3 Mathijs Sanders, 2 Ruud Delwel 2 and Stefan Gröschel 1,4,5 1Molecular Leukemogenesis Group, German Cancer Research Center, Heidelberg, Germany; 2Department of Hematology, Erasmus University Medical Cen - ter, Oncode Institute, Rotterdam, the Netherlands; 3Health Data Science Unit, Medical Faculty Heidelberg and BioQuant, Heidelberg, Germany; 4Internal Medi - cine V, Heidelberg University Hospital, Heidelberg, Germany and 5Oncology Center Worms, Worms, Germany Correspondence: STEFAN GROESCHEL - [email protected] doi:10.3324/haematol.2020.277400 Supplementary Information Supplementary figures A break-RE G2DHE break-RE G2DHE 12345123ctr121ctr [kb] 3 2 1.5 1 0.7 0.5 3071 MOLM-1 B del NTC WT del A del B NTC WT 1212bulk 1234 12123bulk [kb] [kb] 3 3 2 2 1.5 1.5 1 1 0.7 0.7 0.5 0.5 MOLM-1 UCSD-AML1 Fig. S1. PCR validation of expanded single clones upon CRISPR-Cas9 targeting. (A) PCR spanning the insertion sites in K562 shows successful knock-in of desired sequences (break-RE: breakpoint-RE) by the presence of higher-running bands on an agarose gel in respect to the control bands. (B) Rearranged allele-specific PCR of the MOLM-1 and UCSD- AML1 deletion clones (del, del A, del B) confirms the presence of desired deletions with smaller amplicons in comparison to control samples (NTC: nontargeting control, WT: wild- type). Number over each lane corresponds to a different single clone within a given condition.