Investigation Into the Rate-Determining Step Of
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Amplitaq and Amplitaq Gold DNA Polymerase
AmpliTaq and AmpliTaq Gold DNA Polymerase The Most Referenced Brand of DNA Polymerase in the World Date: 2005-05 Notes: Authors are listed alphabetically J. Biol. Chem. (223) Ezoe, S., I. Matsumura, et al. (2005). "GATA Transcription Factors Inhibit Cytokine-dependent Growth and Survival of a Hematopoietic Cell Line through the Inhibition of STAT3 Activity." J. Biol. Chem. 280(13): 13163-13170. http://www.jbc.org/cgi/content/abstract/280/13/13163 Although GATA-1 and GATA-2 were shown to be essential for the development of hematopoietic cells by gene targeting experiments, they were also reported to inhibit the growth of hematopoietic cells. Therefore, in this study, we examined the effects of GATA-1 and GATA-2 on cytokine signals. A tamoxifen-inducible form of GATA-1 (GATA-1/ERT) showed a minor inhibitory effect on interleukin-3 (IL-3)-dependent growth of an IL-3-dependent cell line Ba/F3. On the other hand, it drastically inhibited TPO-dependent growth and gp130-mediated growth/survival of Ba/F3. Similarly, an estradiol-inducible form of GATA-2 (GATA-2/ER) disrupted thrombopoietin (TPO)-dependent growth and gp130-mediated growth/survival of Ba/F3. As for this mechanism, we found that both GATA-1 and GATA-2 directly bound to STAT3 both in vitro and in vivo and inhibited its DNA-binding activity in gel shift assays and chromatin immunoprecipitation assays, whereas they hardly affected STAT5 activity. In addition, endogenous GATA-1 was found to interact with STAT3 in normal megakaryocytes, suggesting that GATA-1 may inhibit STAT3 activity in normal hematopoietic cells. -

Molecular Expression, Characterization and Mechanism of ALAS2 Gain-Of-Function Mutants Vassili Tchaikovskii, Robert J
Tchaikovskii et al. Molecular Medicine (2019) 25:4 Molecular Medicine https://doi.org/10.1186/s10020-019-0070-9 RESEARCHARTICLE Open Access Molecular expression, characterization and mechanism of ALAS2 gain-of-function mutants Vassili Tchaikovskii, Robert J. Desnick and David F. Bishop* Abstract Background: X-linked protoporphyria (XLP) (MIM 300752) is an erythropoietic porphyria due to gain-of-function mutations in the last exon (Ducamp et al., Hum Mol Genet 22:1280-88, 2013) of the erythroid-specific aminolevulinate synthase gene (ALAS2). Five ALAS2 exon 11 variants identified by the NHBLI Exome sequencing project (p.R559H, p.E565D, p.R572C, p.S573F and p.Y586F) were expressed, purified and characterized in order to assess their possible contribution to XLP. To further characterize the XLP gain-of-function region, five novel ALAS2 truncation mutations (p.P561X, p.V562X, p.H563X, p.E569X and p.F575X) were also expressed and studied. Methods: Site-directed mutagenesis was used to generate ALAS2 mutant clones and all were prokaryotically expressed, purified to near homogeneity and characterized by protein and enzyme kinetic assays. Standard deviations were calculated for 3 or more assay replicates. Results: The five ALAS2 single nucleotide variants had from 1.3- to 1.9-fold increases in succinyl-CoA Vmax and 2- to 3-fold increases in thermostability suggesting that most could be gain-of-function modifiers of porphyria instead of causes. One SNP (p.R559H) had markedly low purification yield indicating enzyme instability as the likely cause for XLSA in an elderly patient with x-linked sideroblastic anemia. The five novel ALAS2 truncation mutations had increased Vmax values for both succinyl-CoA and glycine substrates (1.4 to 5.6-fold over wild-type), while the Kms for both substrates were only modestly changed. -

CYP2A6) by P53
Transcriptional Regulation of Human Stress Responsive Cytochrome P450 2A6 (CYP2A6) by p53 Hao Hu M.Biotech. (Biotechnology) 2012 The University of Queensland B.B.A. 2009 University of Electronic Science and Technology of China B.Sc. (Pharmacy) 2009 University of Electronic Science and Technology of China A thesis submitted for the degree of Doctor of Philosophy at The University of Queensland in 2016 School of Medicine ABSTRACT Human cytochrome P450 (CYP) 2A6 is highly expressed in the liver and the encoding gene is regulated by various stress activated transcription factors, such as the nuclear factor (erythroid-derived 2)-like 2 (Nrf-2). Unlike the other xenobiotic metabolising CYP enzymes (XMEs), CYP2A6 only plays a minor role in xenobiotic metabolism. The CYP2A6 is highly induced by multiple forms of cellular stress conditions, where XMEs expression is normally inhibited. Recent findings suggest that the CYP2A6 plays an important role in regulating BR homeostasis. A computer based sequence analysis on the 3 kb proximate CYP2A6 promoter revealed several putative binding sites for p53, a protein that mediates regulation of antioxidant and apoptosis pathways. In this study, the role of p53 in CYP2A6 gene regulation is demonstrated. The site closest to transcription start site (TSS) is highly homologous with the p53 consensus sequence. The p53 responsiveness of this site was confirmed by transfections with various stepwise deleted of CYP2A6-5’-Luc constructs containing the putative p53RE. Deletion of the putative p53RE resulted in a total abolishment of p53 responsiveness of CYP2A6 promoter. Specific binding of p53 to the putative p53RE was detected by electrophoresis mobility shift assay. -

1 Metabolic Dysfunction Is Restricted to the Sciatic Nerve in Experimental
Page 1 of 255 Diabetes Metabolic dysfunction is restricted to the sciatic nerve in experimental diabetic neuropathy Oliver J. Freeman1,2, Richard D. Unwin2,3, Andrew W. Dowsey2,3, Paul Begley2,3, Sumia Ali1, Katherine A. Hollywood2,3, Nitin Rustogi2,3, Rasmus S. Petersen1, Warwick B. Dunn2,3†, Garth J.S. Cooper2,3,4,5* & Natalie J. Gardiner1* 1 Faculty of Life Sciences, University of Manchester, UK 2 Centre for Advanced Discovery and Experimental Therapeutics (CADET), Central Manchester University Hospitals NHS Foundation Trust, Manchester Academic Health Sciences Centre, Manchester, UK 3 Centre for Endocrinology and Diabetes, Institute of Human Development, Faculty of Medical and Human Sciences, University of Manchester, UK 4 School of Biological Sciences, University of Auckland, New Zealand 5 Department of Pharmacology, Medical Sciences Division, University of Oxford, UK † Present address: School of Biosciences, University of Birmingham, UK *Joint corresponding authors: Natalie J. Gardiner and Garth J.S. Cooper Email: [email protected]; [email protected] Address: University of Manchester, AV Hill Building, Oxford Road, Manchester, M13 9PT, United Kingdom Telephone: +44 161 275 5768; +44 161 701 0240 Word count: 4,490 Number of tables: 1, Number of figures: 6 Running title: Metabolic dysfunction in diabetic neuropathy 1 Diabetes Publish Ahead of Print, published online October 15, 2015 Diabetes Page 2 of 255 Abstract High glucose levels in the peripheral nervous system (PNS) have been implicated in the pathogenesis of diabetic neuropathy (DN). However our understanding of the molecular mechanisms which cause the marked distal pathology is incomplete. Here we performed a comprehensive, system-wide analysis of the PNS of a rodent model of DN. -

Supplementary Materials
1 Supplementary Materials: Supplemental Figure 1. Gene expression profiles of kidneys in the Fcgr2b-/- and Fcgr2b-/-. Stinggt/gt mice. (A) A heat map of microarray data show the genes that significantly changed up to 2 fold compared between Fcgr2b-/- and Fcgr2b-/-. Stinggt/gt mice (N=4 mice per group; p<0.05). Data show in log2 (sample/wild-type). 2 Supplemental Figure 2. Sting signaling is essential for immuno-phenotypes of the Fcgr2b-/-lupus mice. (A-C) Flow cytometry analysis of splenocytes isolated from wild-type, Fcgr2b-/- and Fcgr2b-/-. Stinggt/gt mice at the age of 6-7 months (N= 13-14 per group). Data shown in the percentage of (A) CD4+ ICOS+ cells, (B) B220+ I-Ab+ cells and (C) CD138+ cells. Data show as mean ± SEM (*p < 0.05, **p<0.01 and ***p<0.001). 3 Supplemental Figure 3. Phenotypes of Sting activated dendritic cells. (A) Representative of western blot analysis from immunoprecipitation with Sting of Fcgr2b-/- mice (N= 4). The band was shown in STING protein of activated BMDC with DMXAA at 0, 3 and 6 hr. and phosphorylation of STING at Ser357. (B) Mass spectra of phosphorylation of STING at Ser357 of activated BMDC from Fcgr2b-/- mice after stimulated with DMXAA for 3 hour and followed by immunoprecipitation with STING. (C) Sting-activated BMDC were co-cultured with LYN inhibitor PP2 and analyzed by flow cytometry, which showed the mean fluorescence intensity (MFI) of IAb expressing DC (N = 3 mice per group). 4 Supplemental Table 1. Lists of up and down of regulated proteins Accession No. -

Protein Identities in Evs Isolated from U87-MG GBM Cells As Determined by NG LC-MS/MS
Protein identities in EVs isolated from U87-MG GBM cells as determined by NG LC-MS/MS. No. Accession Description Σ Coverage Σ# Proteins Σ# Unique Peptides Σ# Peptides Σ# PSMs # AAs MW [kDa] calc. pI 1 A8MS94 Putative golgin subfamily A member 2-like protein 5 OS=Homo sapiens PE=5 SV=2 - [GG2L5_HUMAN] 100 1 1 7 88 110 12,03704523 5,681152344 2 P60660 Myosin light polypeptide 6 OS=Homo sapiens GN=MYL6 PE=1 SV=2 - [MYL6_HUMAN] 100 3 5 17 173 151 16,91913397 4,652832031 3 Q6ZYL4 General transcription factor IIH subunit 5 OS=Homo sapiens GN=GTF2H5 PE=1 SV=1 - [TF2H5_HUMAN] 98,59 1 1 4 13 71 8,048185945 4,652832031 4 P60709 Actin, cytoplasmic 1 OS=Homo sapiens GN=ACTB PE=1 SV=1 - [ACTB_HUMAN] 97,6 5 5 35 917 375 41,70973209 5,478027344 5 P13489 Ribonuclease inhibitor OS=Homo sapiens GN=RNH1 PE=1 SV=2 - [RINI_HUMAN] 96,75 1 12 37 173 461 49,94108966 4,817871094 6 P09382 Galectin-1 OS=Homo sapiens GN=LGALS1 PE=1 SV=2 - [LEG1_HUMAN] 96,3 1 7 14 283 135 14,70620005 5,503417969 7 P60174 Triosephosphate isomerase OS=Homo sapiens GN=TPI1 PE=1 SV=3 - [TPIS_HUMAN] 95,1 3 16 25 375 286 30,77169764 5,922363281 8 P04406 Glyceraldehyde-3-phosphate dehydrogenase OS=Homo sapiens GN=GAPDH PE=1 SV=3 - [G3P_HUMAN] 94,63 2 13 31 509 335 36,03039959 8,455566406 9 Q15185 Prostaglandin E synthase 3 OS=Homo sapiens GN=PTGES3 PE=1 SV=1 - [TEBP_HUMAN] 93,13 1 5 12 74 160 18,68541938 4,538574219 10 P09417 Dihydropteridine reductase OS=Homo sapiens GN=QDPR PE=1 SV=2 - [DHPR_HUMAN] 93,03 1 1 17 69 244 25,77302971 7,371582031 11 P01911 HLA class II histocompatibility antigen, -

Characterisation of Bilirubin Metabolic Pathway in Hepatic Mitochondria Siti Nur Fadzilah Muhsain M.Sc
Characterisation of Bilirubin Metabolic Pathway in Hepatic Mitochondria Siti Nur Fadzilah Muhsain M.Sc. (Medical Research) 2005 Universiti Sains Malaysia Postgrad. Dip. (Toxicology) 2003 University of Surrey B.Sc.(Biomed. Sc.) 2000 Universiti Putra Malaysia A thesis submitted for the degree of Doctor of Philosophy at The University of Queensland in 2014 School of Medicine ABSTRACT Bilirubin (BR), a toxic waste product of degraded haem, is a potent antioxidant at physiological concentrations. To achieve the maximum benefit of BR, its intracellular level needs to be carefully regulated. A system comprising of two enzymes, haem oxygenase-1 (HMOX1) and cytochrome P450 2A5 (CYP2A5) exists in the endoplasmic reticulum (ER), responsible for regulating BR homeostasis. This system is induced in response to oxidative stress. In this thesis, oxidative stress caused accumulation of these enzymes in mitochondria — major producers and targets of reactive oxygen species (ROS) — is demonstrated. To understand the significance of this intracellular targeting, properties of microsomal and mitochondrial BR metabolising enzymes were compared and the capacity of mitochondrial CYP2A5 to oxidise BR in response to oxidative stress is reported. Microsomes and mitochondrial fractions were isolated from liver homogenates of DBA/2J mice, administered with sub-toxic dose of pyrazole, an oxidant stressor. The purity of extracted organelles was determined by analysing the expressions and activities of their respective marker enzymes. HMOX1 and CYP2A5 were significantly increased in microsomes and even more so in mitochondria in response to pyrazole-induced oxidative stress. By contrast, the treatment did not increase either microsomes or mitochondrial Uridine-diphosphate-glucuronosyltransferase 1A1 (UGT1A1), the sole enzyme that catalyses BR elimination through glucuronidation. -

Targeting Proteins Self-Association in Drug Design
Targeting proteins self-association in drug design Léopold Thabault,1,2 Maxime Liberelle,1 Raphaël Frédérick1* 1 Louvain Drug Research Institute (LDRI), Université catholique de Louvain (UCLouvain), B-1200 Brussels, Belgium. 2 Pole of Pharmacology and Therapeutics, Institut de Recherche Expérimentale et Clinique (IREC), Université catholique de Louvain (UCLouvain),B-1200 Brussels, Belgium. * Corresponding Author : Prof. Raphaël Frédérick Université catholique de Louvain (UCLouvain) Louvain Drug Research Institute (LDRI) Tour Van Helmont, B1.73.10 73 avenue Mounier 1200 Brussels Belgium Phone : +32 2 764 73 41 1 Abstract Protein self-association is a universal phenomenon essential for stability and molecular recognition. Disrupting constitutive homomers constitutes an original and emerging strategy in drug design. Inhibition of homomeric proteins can be achieved through direct complex disruption, subunits intercalation or by promoting inactive oligomeric states. Targeting self- interaction grants several advantages over active site inhibition thanks to the stimulation of protein degradation, the enhancement of selectivity, a sub-stoichiometric inhibition and a by- pass of compensatory mechanisms. This new landscape in protein inhibition is driven by the development of biophysical and biochemical tools suited for the study of homomeric proteins, such as DSF, native MS, FRET spectroscopy, two-dimensional NMR and X-ray crystallography. The present review covers the different aspects of this new paradigm in drug design. Keywords Protein-protein interaction; Homo-oligomers Disruptor; Protein destabilisation; Allosteric inhibition; Self-association. Teaser Disrupting protein self-association bears great therapeutic promesses and could provide exciting alternatives to active-site inhibitors. 2 1. PPI as a well-accepted paradigm in drug design The human interactome is nowadays widely considered as one of the keys to cell mechanistic regulation. -

GATA Transcription Factors Directly Regulate the Parkinson's
GATA transcription factors directly regulate the Parkinson’s disease-linked gene ␣-synuclein Clemens R. Scherzer*†‡, Jeffrey A. Grass§, Zhixiang Liao*, Imelda Pepivani*, Bin Zheng*, Aron C. Eklund*, Paul A. Ney¶, Juliana Ngʈ, Meghan McGoldrick*, Brit Mollenhauer*, Emery H. Bresnick†§**, and Michael G. Schlossmacher*†ʈ†† *Center for Neurologic Diseases, Harvard Medical School and Brigham and Women’s Hospital, Cambridge, MA 02139; §Department of Pharmacology, University of Wisconsin School of Medicine and Public Health, Madison, WI 53706; ¶Department of Biochemistry, St. Jude Children’s Research Hospital, Memphis, TN 38105; and ʈDivision of Neurosciences, Ottawa Health Research Institute, University of Ottawa, Ottawa, ON, Canada K1H 8M5 Edited by Gregory A. Petsko, Brandeis University, Waltham, MA, and approved May 27, 2008 (received for review March 13, 2008) Increased ␣-synuclein gene (SNCA) dosage due to locus multipli- Results cation causes autosomal dominant Parkinson’s disease (PD). Vari- SNCA mRNA and Protein Are Highly Abundant in Human and Mouse ation in SNCA expression may be critical in common, genetically Erythroid Cells. Relative SNCA mRNA abundance was determined complex PD but the underlying regulatory mechanism is unknown. by comparing the SNCA mRNA abundance in each target tissue to We show that SNCA and the heme metabolism genes ALAS2, FECH, the calibrator Universal Human Reference RNA (Fig. 1A). SNCA and BLVRB form a block of tightly correlated gene expression in 113 mRNA abundance was high in whole blood from human donors samples of human blood, where SNCA naturally abounds (vali- (relative abundance 10 {range 6.6–15.3}) with very high levels in ؊ ؊ ؊ -؋ 10 11, 1.8 ؋ 10 10, and 6.6 ؋ 10 5). -

Heme Oxygenase-1 Regulates Mitochondrial Quality Control in the Heart
RESEARCH ARTICLE Heme oxygenase-1 regulates mitochondrial quality control in the heart Travis D. Hull,1,2 Ravindra Boddu,1,3 Lingling Guo,2,3 Cornelia C. Tisher,1,3 Amie M. Traylor,1,3 Bindiya Patel,1,4 Reny Joseph,1,3 Sumanth D. Prabhu,1,4,5 Hagir B. Suliman,6 Claude A. Piantadosi,7 Anupam Agarwal,1,3,5 and James F. George2,3,4 1Department of Medicine, 2Department of Surgery, 3Nephrology Research and Training Center, and 4Comprehensive Cardiovascular Center, University of Alabama at Birmingham, Birmingham, Alabama, USA. 5Department of Veterans Affairs, Birmingham, Alabama, USA. 6Department of Anesthesiology and 7Department of Pulmonary, Allergy and Critical Care, Duke University School of Medicine, Durham, North Carolina, USA. The cardioprotective inducible enzyme heme oxygenase-1 (HO-1) degrades prooxidant heme into equimolar quantities of carbon monoxide, biliverdin, and iron. We hypothesized that HO-1 mediates cardiac protection, at least in part, by regulating mitochondrial quality control. We treated WT and HO-1 transgenic mice with the known mitochondrial toxin, doxorubicin (DOX). Relative to WT mice, mice globally overexpressing human HO-1 were protected from DOX-induced dilated cardiomyopathy, cardiac cytoarchitectural derangement, and infiltration of CD11b+ mononuclear phagocytes. Cardiac-specific overexpression of HO-1 ameliorated DOX-mediated dilation of the sarcoplasmic reticulum as well as mitochondrial disorganization in the form of mitochondrial fragmentation and increased numbers of damaged mitochondria in autophagic vacuoles. HO-1 overexpression promotes mitochondrial biogenesis by upregulating protein expression of NRF1, PGC1α, and TFAM, which was inhibited in WT animals treated with DOX. Concomitantly, HO-1 overexpression inhibited the upregulation of the mitochondrial fission mediator Fis1 and resulted in increased expression of the fusion mediators, Mfn1 and Mfn2. -

CDH12 Cadherin 12, Type 2 N-Cadherin 2 RPL5 Ribosomal
5 6 6 5 . 4 2 1 1 1 2 4 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 1 2 2 A A A A A A A A A A A A A A A A A A A A C C C C C C C C C C C C C C C C C C C C R R R R R R R R R R R R R R R R R R R R B , B B B B B B B B B B B B B B B B B B B , 9 , , , , 4 , , 3 0 , , , , , , , , 6 2 , , 5 , 0 8 6 4 , 7 5 7 0 2 8 9 1 3 3 3 1 1 7 5 0 4 1 4 0 7 1 0 2 0 6 7 8 0 2 5 7 8 0 3 8 5 4 9 0 1 0 8 8 3 5 6 7 4 7 9 5 2 1 1 8 2 2 1 7 9 6 2 1 7 1 1 0 4 5 3 5 8 9 1 0 0 4 2 5 0 8 1 4 1 6 9 0 0 6 3 6 9 1 0 9 0 3 8 1 3 5 6 3 6 0 4 2 6 1 0 1 2 1 9 9 7 9 5 7 1 5 8 9 8 8 2 1 9 9 1 1 1 9 6 9 8 9 7 8 4 5 8 8 6 4 8 1 1 2 8 6 2 7 9 8 3 5 4 3 2 1 7 9 5 3 1 3 2 1 2 9 5 1 1 1 1 1 1 5 9 5 3 2 6 3 4 1 3 1 1 4 1 4 1 7 1 3 4 3 2 7 6 4 2 7 2 1 2 1 5 1 6 3 5 6 1 3 6 4 7 1 6 5 1 1 4 1 6 1 7 6 4 7 e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e e l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l l p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p p m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m m -
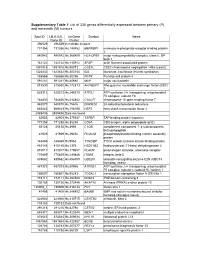
Supplementary Table 1 List of 335 Genes Differentially Expressed Between Primary (P) and Metastatic (M) Tumours
Supplementary Table 1 List of 335 genes differentially expressed between primary (P) and metastatic (M) tumours Spot ID I.M.A.G.E. UniGene Symbol Name Clone ID Cluster 296529 296529 In multiple clusters 731356 731356 Hs.140452 M6PRBP1 mannose-6-phosphate receptor binding protein 1 840942 840942 Hs.368409 HLA-DPB1 major histocompatibility complex, class II, DP beta 1 142122 142122 Hs.115912 AFAP actin filament associated protein 1891918 1891918 Hs.90073 CSE1L CSE1 chromosome segregation 1-like (yeast) 1323432 1323432 Hs.303154 IDS iduronate 2-sulfatase (Hunter syndrome) 788566 788566 Hs.80296 PCP4 Purkinje cell protein 4 591281 591281 Hs.80680 MVP major vault protein 815530 815530 Hs.172813 ARHGEF7 Rho guanine nucleotide exchange factor (GEF) 7 825312 825312 Hs.246310 ATP5J ATP synthase, H+ transporting, mitochondrial F0 complex, subunit F6 784830 784830 Hs.412842 C10orf7 chromosome 10 open reading frame 7 840878 840878 Hs.75616 DHCR24 24-dehydrocholesterol reductase 669443 669443 Hs.158195 HSF2 heat shock transcription factor 2 2485436 2485436 Data not found 82903 82903 Hs.370937 TAPBP TAP binding protein (tapasin) 771258 771258 Hs.85258 CD8A CD8 antigen, alpha polypeptide (p32) 85128 85128 Hs.8986 C1QB complement component 1, q subcomponent, beta polypeptide 41929 41929 Hs.39252 PICALM phosphatidylinositol binding clathrin assembly protein 148469 148469 Hs.9963 TYROBP TYRO protein tyrosine kinase binding protein 415145 415145 Hs.1376 HSD11B2 hydroxysteroid (11-beta) dehydrogenase 2 810017 810017 Hs.179657 PLAUR plasminogen activator,