Supporting Figure 12
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Ovulation-Selective Genes: the Generation and Characterization of an Ovulatory-Selective Cdna Library
531 Ovulation-selective genes: the generation and characterization of an ovulatory-selective cDNA library A Hourvitz1,2*, E Gershon2*, J D Hennebold1, S Elizur2, E Maman2, C Brendle1, E Y Adashi1 and N Dekel2 1Division of Reproductive Sciences, Department of Obstetrics and Gynecology, University of Utah Health Sciences Center, Salt Lake City, Utah 84132, USA 2Department of Biological Regulation, Weizmann Institute of Science, Rehovot, Israel (Requests for offprints should be addressed to N Dekel; Email: [email protected]) *(A Hourvitz and E Gershon contributed equally to this paper) (J D Hennebold is now at Division of Reproductive Sciences, Oregon National Primate Research Center, Oregon Health and Science University, Beaverton, Oregon 97006, USA) Abstract Ovulation-selective/specific genes, that is, genes prefer- (FAE-1) homolog, found to be localized to the inner entially or exclusively expressed during the ovulatory periantral granulosa and to the cumulus granulosa cells of process, have been the subject of growing interest. We antral follicles. The FAE-1 gene is a -ketoacyl-CoA report herein studies on the use of suppression subtractive synthase belonging to the fatty acid elongase (ELO) hybridization (SSH) to construct a ‘forward’ ovulation- family, which catalyzes the initial step of very long-chain selective/specific cDNA library. In toto, 485 clones were fatty acid synthesis. All in all, the present study accom- sequenced and analyzed for homology to known genes plished systematic identification of those hormonally with the basic local alignment tool (BLAST). Of those, regulated genes that are expressed in the ovary in an 252 were determined to be nonredundant. -

ATP-Induced Focal Adhesion Kinase Activity Is Negatively Modulated by Phospholipase D2 in PC12 Cells
EXPERIMENTAL and MOLECULAR MEDICINE, Vol. 33, No. 3, 150-155, September 2001 ATP-induced focal adhesion kinase activity is negatively modulated by phospholipase D2 in PC12 cells Yoe-Sik Bae1 and Sung Ho Ryu1,2 Introduction 1 Division of Molecular and Life Sciences, Pohang University of Purinergic receptors have been reported to play impor- Science and Technology, Pohang 790-784, Korea tant roles on the regulation of neuronal cell functions 2 Corresponding author: Tel, +82-54-279-2292; (Communi et al., 2000; Di Iorio et al., 1998). ATP, a Fax, +82-54-279-2199; E-mail, [email protected] ligand for the receptors modulate various cellular re- sponses such as mitogenic and morphogenic activity in Accepted 18 September 2001 PC12 rat pheochromocytoma cells (Neary et al., 1996; Soltoff et al., 1998; Schindelholz et al., 2000). Stimu- Abbreviations: Fak, focal adhesion kinase; PLD, phospholipase D; lation of cells with ATP induces tyrosine phosphorylation PA, phosphatidic acid; PC, phosphatidylcholine; DAG, diacylglyc- of several cytoskeletal proteins and focal adhesion erol; PBt, phosphatidylbutanol; PKC, protein kinase C; PAP, phos- molecules such as focal adhesion kinase (Fak), proline- phatidic acid phosphohydrolase rich tyrosine kinase (Pyk2), and paxillin (Soltoff et al., 1998; Schindelholz et al., 2000). Since these cytosk- eleton-associated proteins have been regarded as important factors for the regulation of neuronal cell Abstract functions, the study on the regulatory mechanism for the proteins remains an important issue. Extracellular ATP has been known to modulate vari- Phospholipase D (PLD) catalyzes the hydrolysis of ous cellular responses including mitogenesis, secre- phosphatidylcholine (PC) into phosphatidic acid (PA) tion and morphogenic activity in neuronal cells. -

Datasheet for Protein Phosphatase 1 (PP1) (P0754; Lot 0121306)
Supplied in: 200 mM NaCl, 50 mM HEPES Unit Definition: One unit is defined as Notes On Use: Avoid freeze/thaw cycles. Can be Protein the amount of enzyme that hydrolyzes 1 nmol of stored for 1 week or less at –20°C. (pH 7.0 @ 25°C), 1 mM MnCl2, 0.1 mM EGTA, Phosphatase 1 2.5 mM dithiothreitol, 0.025% Tween-20 and p-Nitrophenyl Phosphate (50 mM) (NEB #P0757) 50% glycerol. Store at –70°C in 1 minute at 30°C in a total reaction volume of The following information can be used as (PP1) 50 µl. suggested initial conditions for dephosphorylation 1-800-632-7799 Applications: PP1 can be used to release of proteins with PP1. [email protected] phosphate groups from phosphorylated serine, Specific Activity: ~ 80,000 units/mg. www.neb.com 0.1 unit of PP1 removes ~100% of phosphates P0754S 012130614061 threonine and tyrosine residues in proteins. Note that different proteins are dephosphorylated at Molecular Weight: 37.5 kDa. (0.5 nmol) from phosphoserine/threonine different rates. residues in phosphorylase a as well as in P0754S r y Purity: PP1 has been purified to > 90% phosphorylated myelin basic protein (phospho- 100 units 2,500 U/ml Lot: 0121306 Reagents Supplied with Enzyme: homogeneity as determined by SDS-PAGE and MyBP, 18.5 kDa) in 30 minutes in a 50 µl reaction. RECOMBINANT Store at –70°C Exp: 6/14 10X NEBuffer for Protein MetalloPhosphatases Coomassie Blue staining. The concentration of phospho-MyBP is 10 µM (PMP) with respect to phosphate. Description: Protein Phosphatase 1 (PP1) is a 10X MnCl2 (10 mM) Quality Assurance: PP1 contains no detectable Mn2+-dependent protein phosphatase with activity protease activity. -

Role of Phospholipases in Adrenal Steroidogenesis
229 1 W B BOLLAG Phospholipases in adrenal 229:1 R29–R41 Review steroidogenesis Role of phospholipases in adrenal steroidogenesis Wendy B Bollag Correspondence should be addressed Charlie Norwood VA Medical Center, One Freedom Way, Augusta, GA, USA to W B Bollag Department of Physiology, Medical College of Georgia, Augusta University (formerly Georgia Regents Email University), Augusta, GA, USA [email protected] Abstract Phospholipases are lipid-metabolizing enzymes that hydrolyze phospholipids. In some Key Words cases, their activity results in remodeling of lipids and/or allows the synthesis of other f adrenal cortex lipids. In other cases, however, and of interest to the topic of adrenal steroidogenesis, f angiotensin phospholipases produce second messengers that modify the function of a cell. In this f intracellular signaling review, the enzymatic reactions, products, and effectors of three phospholipases, f phospholipids phospholipase C, phospholipase D, and phospholipase A2, are discussed. Although f signal transduction much data have been obtained concerning the role of phospholipases C and D in regulating adrenal steroid hormone production, there are still many gaps in our knowledge. Furthermore, little is known about the involvement of phospholipase A2, Endocrinology perhaps, in part, because this enzyme comprises a large family of related enzymes of that are differentially regulated and with different functions. This review presents the evidence supporting the role of each of these phospholipases in steroidogenesis in the Journal Journal of Endocrinology adrenal cortex. (2016) 229, R1–R13 Introduction associated GTP-binding protein exchanges a bound GDP for a GTP. The G protein with GTP bound can then Phospholipids serve a structural function in the cell in that activate the enzyme, phospholipase C (PLC), that cleaves they form the lipid bilayer that maintains cell integrity. -

Determining HDAC8 Substrate Specificity by Noah Ariel Wolfson A
Determining HDAC8 substrate specificity by Noah Ariel Wolfson A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy (Biological Chemistry) in the University of Michigan 2014 Doctoral Committee: Professor Carol A. Fierke, Chair Professor Robert S. Fuller Professor Anna K. Mapp Associate Professor Patrick J. O’Brien Associate Professor Raymond C. Trievel Dedication My thesis is dedicated to all my family, mentors, and friends who made getting to this point possible. ii Table of Contents Dedication ....................................................................................................................................... ii List of Figures .............................................................................................................................. viii List of Tables .................................................................................................................................. x List of Appendices ......................................................................................................................... xi Abstract ......................................................................................................................................... xii Chapter 1 HDAC8 substrates: Histones and beyond ...................................................................... 1 Overview ..................................................................................................................................... 1 HDAC introduction -

Transcriptional Analysis of Sodium Valproate in a Serotonergic Cell Line Reveals Gene Regulation Through Both HDAC Inhibition-Dependent and Independent Mechanisms
bioRxiv preprint doi: https://doi.org/10.1101/837732; this version posted November 12, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Transcriptional analysis of sodium valproate in a serotonergic cell line reveals gene regulation through both HDAC inhibition-dependent and independent mechanisms Priyanka Sinha1,2, Simone Cree1,2, Allison L. Miller1,2, John F. Pearson1,2,3, Martin A. Kennedy1,2. 1Gene Structure and Function Laboratory, Department of Pathology and Biomedical Science, University of Otago, Christchurch, New Zealand. 2Carney Centre for Pharmacogenomics, University of Otago, Christchurch, New Zealand. 3Biostatistics and Computational Biology Unit, University of Otago, Christchurch, New Zealand. Correspondence to: Prof. M. A. Kennedy Department of Pathology and Biomedical Science University of Otago, Christchurch Christchurch, New Zealand Email: [email protected] Keywords: RNA-Seq, NanoString, lithium, valproate, HDAC inhibitor, mood stabilizer 1 bioRxiv preprint doi: https://doi.org/10.1101/837732; this version posted November 12, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Abstract Sodium valproate (VPA) is a histone deacetylase (HDAC) inhibitor, widely prescribed in the treatment of bipolar disorder, and yet the precise modes of therapeutic action for this drug are not fully understood. -
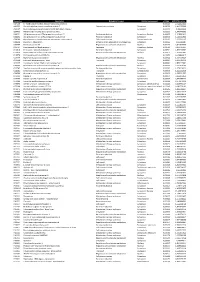
Accession Description Biological Process Cellular Component P-Value SCZ/CTRL Ration A4FU69 EF-Hand Calcium-Binding Domain-Contai
Accession Description Biological Process Cellular component p-Value SCZ/CTRL ration A4FU69 EF-hand calcium-binding domain-containing protein 5 0,001101 2,724427411 A4UGR9 Xin actin-binding repeat-containing protein 2 Cytoskeletal anchoring Cytoplasm 0,006756 1,413953388 A6NCE7 Microtubule-associated proteins 1A/1B light chain 3 beta 2 Cytoplasm 0,001417 1,99612969 B2RPK0 Putative high mobility group protein B1-like 1 0,032314 1,590743352 O00231 26S proteasome non-ATPase regulatory subunit 11 Protein metabolism Cytoplasm; Nucleus 0,000029 1,428664042 O00232 26S proteasome non-ATPase regulatory subunit 12 Protein metabolism Cytoplasm 0,008566 1,544545922 O00264 Membrane-associated progesterone receptor component 1 Cell communication Plasma membrane 0,001459 2,322924147 O00429 Dynamin-1-like protein Mitochondrion organization and biogenesis Cytoplasm 0,006560 0,06391487 O00567 Nucleolar protein 56 Regulation of nucleotide metabolism Nucleus 0,007330 3,842896338 O14737 Programmed cell death protein 5 Apoptosis Cytoplasm; Nucleus 0,006358 3,836727237 O14818 Proteasome subunit alpha type-7 Protein metabolism Cytoplasm 0,030521 1,893928387 O14979 Heterogeneous nuclear ribonucleoprotein D-like Regulation of nucleotide metabolism Nucleus 0,000637 2,150005885 O15078 Centrosomal protein of 290 kDa 0,015359 14,26648619 O15347 High mobility group protein B3 Regulation of nucleotide metabolism Nucleus 0,005500 1,364309014 O15540 Fatty acid-binding protein_ brain Transport Cytoplasm 0,000087 3,125786118 O43237 Cytoplasmic dynein 1 light intermediate -

The Regulatory Roles of Phosphatases in Cancer
Oncogene (2014) 33, 939–953 & 2014 Macmillan Publishers Limited All rights reserved 0950-9232/14 www.nature.com/onc REVIEW The regulatory roles of phosphatases in cancer J Stebbing1, LC Lit1, H Zhang, RS Darrington, O Melaiu, B Rudraraju and G Giamas The relevance of potentially reversible post-translational modifications required for controlling cellular processes in cancer is one of the most thriving arenas of cellular and molecular biology. Any alteration in the balanced equilibrium between kinases and phosphatases may result in development and progression of various diseases, including different types of cancer, though phosphatases are relatively under-studied. Loss of phosphatases such as PTEN (phosphatase and tensin homologue deleted on chromosome 10), a known tumour suppressor, across tumour types lends credence to the development of phosphatidylinositol 3--kinase inhibitors alongside the use of phosphatase expression as a biomarker, though phase 3 trial data are lacking. In this review, we give an updated report on phosphatase dysregulation linked to organ-specific malignancies. Oncogene (2014) 33, 939–953; doi:10.1038/onc.2013.80; published online 18 March 2013 Keywords: cancer; phosphatases; solid tumours GASTROINTESTINAL MALIGNANCIES abs in sera were significantly associated with poor survival in Oesophageal cancer advanced ESCC, suggesting that they may have a clinical utility in Loss of PTEN (phosphatase and tensin homologue deleted on ESCC screening and diagnosis.5 chromosome 10) expression in oesophageal cancer is frequent, Cao et al.6 investigated the role of protein tyrosine phosphatase, among other gene alterations characterizing this disease. Zhou non-receptor type 12 (PTPN12) in ESCC and showed that PTPN12 et al.1 found that overexpression of PTEN suppresses growth and protein expression is higher in normal para-cancerous tissues than induces apoptosis in oesophageal cancer cell lines, through in 20 ESCC tissues. -

Histone Deacetylase 3 As a Novel Therapeutic Target in Multiple Myeloma
Leukemia (2014) 28, 680–689 & 2014 Macmillan Publishers Limited All rights reserved 0887-6924/14 www.nature.com/leu ORIGINAL ARTICLE Histone deacetylase 3 as a novel therapeutic target in multiple myeloma J Minami1,4, R Suzuki1,4, R Mazitschek2, G Gorgun1, B Ghosh2,3, D Cirstea1,YHu1, N Mimura1, H Ohguchi1, F Cottini1, J Jakubikova1, NC Munshi1, SJ Haggarty3, PG Richardson1, T Hideshima1 and KC Anderson1 Histone deacetylases (HDACs) represent novel molecular targets for the treatment of various types of cancers, including multiple myeloma (MM). Many HDAC inhibitors have already shown remarkable antitumor activities in the preclinical setting; however, their clinical utility is limited because of unfavorable toxicities associated with their broad range HDAC inhibitory effects. Isoform- selective HDAC inhibition may allow for MM cytotoxicity without attendant side effects. In this study, we demonstrated that HDAC3 knockdown and a small-molecule HDAC3 inhibitor BG45 trigger significant MM cell growth inhibition via apoptosis, evidenced by caspase and poly (ADP-ribose) polymerase cleavage. Importantly, HDAC3 inhibition downregulates phosphorylation (tyrosine 705 and serine 727) of signal transducers and activators of transcription 3 (STAT3). Neither interleukin-6 nor bone marrow stromal cells overcome this inhibitory effect of HDAC3 inhibition on phospho-STAT3 and MM cell growth. Moreover, HDAC3 inhibition also triggers hyperacetylation of STAT3, suggesting crosstalk signaling between phosphorylation and acetylation of STAT3. Importantly, inhibition of HDAC3, but not HDAC1 or 2, significantly enhances bortezomib-induced cytotoxicity. Finally, we confirm that BG45 alone and in combination with bortezomib trigger significant tumor growth inhibition in vivo in a murine xenograft model of human MM. -

Inhibition of Protein Phosphatase 1 Translation by Preventing JNK
Active ERK Contributes to Protein Translation by Preventing JNK-Dependent Inhibition of Protein Phosphatase 1 This information is current as Martha M. Monick, Linda S. Powers, Thomas J. Gross, of September 28, 2021. Dawn M. Flaherty, Christopher W. Barrett and Gary W. Hunninghake J Immunol 2006; 177:1636-1645; ; doi: 10.4049/jimmunol.177.3.1636 http://www.jimmunol.org/content/177/3/1636 Downloaded from References This article cites 59 articles, 43 of which you can access for free at: http://www.jimmunol.org/content/177/3/1636.full#ref-list-1 http://www.jimmunol.org/ Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication by guest on September 28, 2021 *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2006 by The American Association of Immunologists All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Active ERK Contributes to Protein Translation by Preventing JNK-Dependent Inhibition of Protein Phosphatase 11 Martha M. Monick,2 Linda S. Powers, Thomas J. Gross, Dawn M. Flaherty, Christopher W. -

Human Induced Pluripotent Stem Cell–Derived Podocytes Mature Into Vascularized Glomeruli Upon Experimental Transplantation
BASIC RESEARCH www.jasn.org Human Induced Pluripotent Stem Cell–Derived Podocytes Mature into Vascularized Glomeruli upon Experimental Transplantation † Sazia Sharmin,* Atsuhiro Taguchi,* Yusuke Kaku,* Yasuhiro Yoshimura,* Tomoko Ohmori,* ‡ † ‡ Tetsushi Sakuma, Masashi Mukoyama, Takashi Yamamoto, Hidetake Kurihara,§ and | Ryuichi Nishinakamura* *Department of Kidney Development, Institute of Molecular Embryology and Genetics, and †Department of Nephrology, Faculty of Life Sciences, Kumamoto University, Kumamoto, Japan; ‡Department of Mathematical and Life Sciences, Graduate School of Science, Hiroshima University, Hiroshima, Japan; §Division of Anatomy, Juntendo University School of Medicine, Tokyo, Japan; and |Japan Science and Technology Agency, CREST, Kumamoto, Japan ABSTRACT Glomerular podocytes express proteins, such as nephrin, that constitute the slit diaphragm, thereby contributing to the filtration process in the kidney. Glomerular development has been analyzed mainly in mice, whereas analysis of human kidney development has been minimal because of limited access to embryonic kidneys. We previously reported the induction of three-dimensional primordial glomeruli from human induced pluripotent stem (iPS) cells. Here, using transcription activator–like effector nuclease-mediated homologous recombination, we generated human iPS cell lines that express green fluorescent protein (GFP) in the NPHS1 locus, which encodes nephrin, and we show that GFP expression facilitated accurate visualization of nephrin-positive podocyte formation in -

HDAC3: Taking the SMRT-N-Correct Road to Repression
Oncogene (2007) 26, 5439–5449 & 2007 Nature Publishing Group All rights reserved 0950-9232/07 $30.00 www.nature.com/onc REVIEW HDAC3: taking the SMRT-N-CoRrect road to repression P Karagianni and J Wong1 Department of Molecular and Cellular Biology, Baylor College of Medicine, One Baylor Plaza, Houston, TX, USA Known histone deacetylases (HDACs) are divided into repression when targeted to promoters, as well as different classes, and HDAC3 belongs to Class I. Through physical association with the DNA-binding factor forming multiprotein complexes with the corepressors YY1 (Yang et al., 1997; Dangond et al., 1998; Emiliani SMRT and N-CoR, HDAC3 regulates the transcription et al., 1998). Together, these findings suggested that this of a plethora of genes. A growing list of nonhistone ubiquitously expressed protein could be involved in the substrates extends the role of HDAC3 beyond transcrip- regulation of mammalian gene expression. On the other tional repression. Here, we review data on the composi- hand, HDAC3 contains an intriguingly variable C tion, regulation and mechanism of action of the SMRT/ terminus, with no apparent similarity with other N-CoR-HDAC3 complexes and provide several examples HDACs. This observation led to the hypothesis that of nontranscriptional functions, to illustrate the wide HDAC3 may have some unique properties and may variety of physiological processes affected by this deacety- not be completely redundant with the other HDACs lase. Furthermore, we discuss the implication of HDAC3 (Yang et al., 1997). This is also suggested by the in cancer, focusing on leukemia. We conclude with some differential localization of HDAC3, which, unlike the thoughts about the potential therapeutic efficacies of predominantly nuclear HDACs 1 and 2, can be found in HDAC3 activity modulation.