GSK3 and Its Interactions with the PI3K/AKT/Mtor Signalling Network
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Gene Symbol Gene Description ACVR1B Activin a Receptor, Type IB
Table S1. Kinase clones included in human kinase cDNA library for yeast two-hybrid screening Gene Symbol Gene Description ACVR1B activin A receptor, type IB ADCK2 aarF domain containing kinase 2 ADCK4 aarF domain containing kinase 4 AGK multiple substrate lipid kinase;MULK AK1 adenylate kinase 1 AK3 adenylate kinase 3 like 1 AK3L1 adenylate kinase 3 ALDH18A1 aldehyde dehydrogenase 18 family, member A1;ALDH18A1 ALK anaplastic lymphoma kinase (Ki-1) ALPK1 alpha-kinase 1 ALPK2 alpha-kinase 2 AMHR2 anti-Mullerian hormone receptor, type II ARAF v-raf murine sarcoma 3611 viral oncogene homolog 1 ARSG arylsulfatase G;ARSG AURKB aurora kinase B AURKC aurora kinase C BCKDK branched chain alpha-ketoacid dehydrogenase kinase BMPR1A bone morphogenetic protein receptor, type IA BMPR2 bone morphogenetic protein receptor, type II (serine/threonine kinase) BRAF v-raf murine sarcoma viral oncogene homolog B1 BRD3 bromodomain containing 3 BRD4 bromodomain containing 4 BTK Bruton agammaglobulinemia tyrosine kinase BUB1 BUB1 budding uninhibited by benzimidazoles 1 homolog (yeast) BUB1B BUB1 budding uninhibited by benzimidazoles 1 homolog beta (yeast) C9orf98 chromosome 9 open reading frame 98;C9orf98 CABC1 chaperone, ABC1 activity of bc1 complex like (S. pombe) CALM1 calmodulin 1 (phosphorylase kinase, delta) CALM2 calmodulin 2 (phosphorylase kinase, delta) CALM3 calmodulin 3 (phosphorylase kinase, delta) CAMK1 calcium/calmodulin-dependent protein kinase I CAMK2A calcium/calmodulin-dependent protein kinase (CaM kinase) II alpha CAMK2B calcium/calmodulin-dependent -

Selective Targeting of Cyclin E1-Amplified High-Grade Serous Ovarian Cancer by Cyclin-Dependent Kinase 2 and AKT Inhibition
Published OnlineFirst September 23, 2016; DOI: 10.1158/1078-0432.CCR-16-0620 Biology of Human Tumors Clinical Cancer Research Selective Targeting of Cyclin E1-Amplified High-Grade Serous Ovarian Cancer by Cyclin- Dependent Kinase 2 and AKT Inhibition George Au-Yeung1,2, Franziska Lang1, Walid J. Azar1, Chris Mitchell1, Kate E. Jarman3, Kurt Lackovic3,4, Diar Aziz5, Carleen Cullinane1,6, Richard B. Pearson1,2,7, Linda Mileshkin2,8, Danny Rischin2,8, Alison M. Karst9, Ronny Drapkin10, Dariush Etemadmoghadam1,2,5, and David D.L. Bowtell1,2,7,11 Abstract Purpose: Cyclin E1 (CCNE1) amplification is associated with Results: We validate CDK2 as a therapeutic target by demon- primary treatment resistance and poor outcome in high-grade strating selective sensitivity to gene suppression. However, we found serous ovarian cancer (HGSC). Here, we explore approaches to that dinaciclib did not trigger amplicon-dependent sensitivity in a target CCNE1-amplified cancers and potential strategies to over- panel of HGSC cell lines. A high-throughput compound screen come resistance to targeted agents. identified synergistic combinations in CCNE1-amplified HGSC, Experimental Design: To examine dependency on CDK2 in including dinaciclib and AKT inhibitors. Analysis of genomic data CCNE1-amplified HGSC, we utilized siRNA and conditional from TCGA demonstrated coamplification of CCNE1 and AKT2. shRNA gene suppression, and chemical inhibition using dina- Overexpression of Cyclin E1 and AKT isoforms, in addition to ciclib, a small-molecule CDK2 inhibitor. High-throughput mutant TP53, imparted malignant characteristics in untransformed compound screening was used to identify selective synergistic fallopian tube secretory cells, the dominant site of origin of HGSC. -

In Vitro Pharmacological Profiling of R406 Identifies Molecular Targets
ORIGINAL ARTICLE In vitro pharmacological profiling of R406 identifies molecular targets underlying the clinical effects of fostamatinib Michael G. Rolf, Jon O. Curwen, Margaret Veldman-Jones, Cath Eberlein, Jianyan Wang, Alex Harmer, Caroline J. Hellawell & Martin Braddock AstraZeneca R&D Alderley Park, Macclesfield, Cheshire SK10 4TG, United Kingdom Keywords Abstract Blood pressure elevation, fostamatinib, in vitro pharmacological profiling, R406, SYK Off-target pharmacology may contribute to both adverse and beneficial effects of a new drug. In vitro pharmacological profiling is often applied early in drug Correspondence discovery; there are fewer reports addressing the relevance of broad profiles to Michael G. Rolf, AstraZeneca R&D Molndal,€ clinical adverse effects. Here, we have characterized the pharmacological profile Pepparedsleden 1, 431 83 Mo¨ lndal, Sweden. of the active metabolite of fostamatinib, R406, linking an understanding of drug Tel: +46 31 776 60 40; Fax: +46 31 776 37 selectivity to the increase in blood pressure observed in clinical studies. R406 60; E-mail: [email protected] was profiled in a broad range of in vitro assays to generate a comprehensive Funding Information pharmacological profile and key targets were further investigated using func- No funding information provided. tional and cellular assay systems. A combination of traditional literature searches and text-mining approaches established potential mechanistic links Received: 9 April 2015; Accepted: 14 July between the profile of R406 and clinical side effects. R406 was selective outside 2015 the kinase domain, with only antagonist activity at the adenosine A3 receptor in the range relevant to clinical effects. R406 was less selective in the kinase Pharma Res Per, 3(5), 2015, e00175, doi: 10.1002/prp2.175 domain, having activity at many protein kinases at therapeutically relevant con- centrations when tested in multiple in vitro systems. -

Kinome Profiling of Clinical Cancer Specimens
Published OnlineFirst March 23, 2010; DOI: 10.1158/0008-5472.CAN-09-3989 Review Cancer Research Kinome Profiling of Clinical Cancer Specimens Kaushal Parikh and Maikel P. Peppelenbosch Abstract Over the past years novel technologies have emerged to enable the determination of the transcriptome and proteome of clinical samples. These data sets will prove to be of significant value to our elucidation of the mechanisms that govern pathophysiology and may provide biological markers for future guidance in person- alized medicine. However, an equally important goal is to define those proteins that participate in signaling pathways during the disease manifestation itself or those pathways that are made active during successful clinical treatment of the disease: the main challenge now is the generation of large-scale data sets that will allow us to define kinome profiles with predictive properties on the outcome-of-disease and to obtain insight into tissue-specific analysis of kinase activity. This review describes the current techniques available to gen- erate kinome profiles of clinical tissue samples and discusses the future strategies necessary to achieve new insights into disease mechanisms and treatment targets. Cancer Res; 70(7); 2575–8. ©2010 AACR. Background the current state of the cell or tissue as characterized by pro- teomic and metabolic measurements (3). The past five years have seen an exponential increase in In this review we describe and evaluate the various tech- technological development to explore the genome and the niques that have recently become available to study the ki- proteome to understand the molecular basis of disease. How- nomes of clinical tissue samples. -
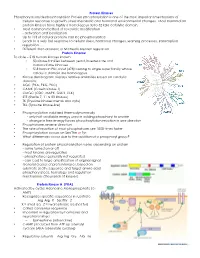
Protein Kinases Phosphorylation/Dephosphorylation Protein Phosphorylation Is One of the Most Important Mechanisms of Cellular Re
Protein Kinases Phosphorylation/dephosphorylation Protein phosphorylation is one of the most important mechanisms of cellular responses to growth, stress metabolic and hormonal environmental changes. Most mammalian protein kinases have highly a homologous 30 to 32 kDa catalytic domain. • Most common method of reversible modification - activation and localization • Up to 1/3 of cellular proteins can be phosphorylated • Leads to a very fast response to cellular stress, hormonal changes, learning processes, transcription regulation .... • Different than allosteric or Michealis Menten regulation Protein Kinome To date – 518 human kinases known • 50 kinase families between yeast, invertebrate and mammaliane kinomes • 518 human PKs, most (478) belong to single super family whose catalytic domain are homologous. • Kinase dendrogram displays relative similarities based on catalytic domains. • AGC (PKA, PKG, PKC) • CAMK (Casein kinase 1) • CMGC (CDC, MAPK, GSK3, CLK) • STE (Sterile 7, 11 & 20 kinases) • TK (Tryosine kinases memb and cyto) • TKL (Tyrosine kinase-like) • Phosphorylation stabilized thermodynamically - only half available energy used in adding phosphoryl to protein - change in free energy forces phosphorylation reaction in one direction • Phosphatases reverse direction • The rate of reaction of most phosphatases are 1000 times faster • Phosphorylation occurs on Ser/The or Tyr • What differences occur due to the addition of a phosphoryl group? • Regulation of protein phosphorylation varies depending on protein - some turned on or off -

Phospho-RPS6KA1(T359) Antibody Peptide Affinity Purified Rabbit Polyclonal Antibody (Pab) Catalog # Ap3497a
9765 Clairemont Mesa Blvd, Suite C San Diego, CA 92124 Tel: 858.875.1900 Fax: 858.622.0609 Phospho-RPS6KA1(T359) Antibody Peptide Affinity Purified Rabbit Polyclonal Antibody (Pab) Catalog # AP3497a Specification Phospho-RPS6KA1(T359) Antibody - Product Information Application WB, DB,E Primary Accession Q15418 Other Accession P10666, P10665, P18652 Reactivity Human Predicted Chicken, Xenopus Host Rabbit Clonality Polyclonal Isotype Rabbit Ig Calculated MW 82723 Phospho-RPS6KA1(T359) Antibody - Additional Information Gene ID 6195 Other Names Western blot analysis of extracts from Hela Ribosomal protein S6 kinase alpha-1, cells,untreated or treated with S6K-alpha-1, 90 kDa ribosomal protein S6 TPA,200nM,using phospho RPS6KA1-T359 (left) kinase 1, p90-RSK 1, p90RSK1, p90S6K, MAP or RPS6KA1 antibody(right) kinase-activated protein kinase 1a, MAPK-activated protein kinase 1a, MAPKAP kinase 1a, MAPKAPK-1a, Ribosomal S6 kinase 1, RSK-1, RPS6KA1, MAPKAPK1A, RSK1 Target/Specificity This RPS6KA1 Antibody is generated from rabbits immunized with a KLH conjugated synthetic phosphopeptide corresponding to amino acid residues surrounding T359 of human RPS6KA1. Dilution WB~~1:2000 DB~~1:500 Format Purified polyclonal antibody supplied in PBS with 0.09% (W/V) sodium azide. This antibody is purified through a protein A column, followed by peptide affinity purification. Dot blot analysis of anti-RPS6KA1-pT359 Pab (RB13385) on nitrocellulose membrane. 50ng Storage of Phospho-peptide or Non Phospho-peptide per Maintain refrigerated at 2-8°C for up to 6 dot were adsorbed. Antibody working months. For long term storage store at -20°C concentrations are 0.5ug per ml. in small aliquots to prevent freeze-thaw cycles. -

Novel Driver Strength Index Highlights Important Cancer Genes in TCGA Pancanatlas Patients
medRxiv preprint doi: https://doi.org/10.1101/2021.08.01.21261447; this version posted August 5, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. It is made available under a CC-BY-NC-ND 4.0 International license . Novel Driver Strength Index highlights important cancer genes in TCGA PanCanAtlas patients Aleksey V. Belikov*, Danila V. Otnyukov, Alexey D. Vyatkin and Sergey V. Leonov Laboratory of Innovative Medicine, School of Biological and Medical Physics, Moscow Institute of Physics and Technology, 141701 Dolgoprudny, Moscow Region, Russia *Corresponding author: [email protected] NOTE: This preprint reports new research that has not been certified by peer review and should not be used to guide clinical practice. 1 medRxiv preprint doi: https://doi.org/10.1101/2021.08.01.21261447; this version posted August 5, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. It is made available under a CC-BY-NC-ND 4.0 International license . Abstract Elucidating crucial driver genes is paramount for understanding the cancer origins and mechanisms of progression, as well as selecting targets for molecular therapy. Cancer genes are usually ranked by the frequency of mutation, which, however, does not necessarily reflect their driver strength. Here we hypothesize that driver strength is higher for genes that are preferentially mutated in patients with few driver mutations overall, because these few mutations should be strong enough to initiate cancer. -

Expressed Gene Fusions As Frequent Drivers of Poor Outcomes in Hormone Receptor–Positive Breast Cancer
Published OnlineFirst December 14, 2017; DOI: 10.1158/2159-8290.CD-17-0535 RESEARCH ARTICLE Expressed Gene Fusions as Frequent Drivers of Poor Outcomes in Hormone Receptor–Positive Breast Cancer Karina J. Matissek1,2, Maristela L. Onozato3, Sheng Sun1,2, Zongli Zheng2,3,4, Andrew Schultz1, Jesse Lee3, Kristofer Patel1, Piiha-Lotta Jerevall2,3, Srinivas Vinod Saladi1,2, Allison Macleay3, Mehrad Tavallai1,2, Tanja Badovinac-Crnjevic5, Carlos Barrios6, Nuran Beşe7, Arlene Chan8, Yanin Chavarri-Guerra9, Marcio Debiasi6, Elif Demirdögen10, Ünal Egeli10, Sahsuvar Gökgöz10, Henry Gomez11, Pedro Liedke6, Ismet Tasdelen10, Sahsine Tolunay10, Gustavo Werutsky6, Jessica St. Louis1, Nora Horick12, Dianne M. Finkelstein2,12, Long Phi Le2,3, Aditya Bardia1,2, Paul E. Goss1,2, Dennis C. Sgroi2,3, A. John Iafrate2,3, and Leif W. Ellisen1,2 ABSTRACT We sought to uncover genetic drivers of hormone receptor–positive (HR+) breast cancer, using a targeted next-generation sequencing approach for detecting expressed gene rearrangements without prior knowledge of the fusion partners. We identified inter- genic fusions involving driver genes, including PIK3CA, AKT3, RAF1, and ESR1, in 14% (24/173) of unselected patients with advanced HR+ breast cancer. FISH confirmed the corresponding chromo- somal rearrangements in both primary and metastatic tumors. Expression of novel kinase fusions in nontransformed cells deregulates phosphoprotein signaling, cell proliferation, and survival in three- dimensional culture, whereas expression in HR+ breast cancer models modulates estrogen-dependent growth and confers hormonal therapy resistance in vitro and in vivo. Strikingly, shorter overall survival was observed in patients with rearrangement-positive versus rearrangement-negative tumors. Cor- respondingly, fusions were uncommon (<5%) among 300 patients presenting with primary HR+ breast cancer. -

GSK-3 and Tau: a Key Duet in Alzheimer's Disease
cells Review GSK-3 and Tau: A Key Duet in Alzheimer’s Disease Carmen Laura Sayas 1,* and Jesús Ávila 2,3,* 1 Instituto de Tecnologías Biomédicas (ITB), Universidad de La Laguna (ULL), 38200 Tenerife, Spain 2 Centro de Biología Molecular Severo Ochoa (CBMSO), Consejo Superior de Investigaciones Científicas (CSIC) y la Universidad Autónoma de Madrid (UAM), 28049 Madrid, Spain 3 Centro de Investigación Biomédica en Red de Enfermedades Neurodegenerativas (CIBERNED), Valderrebollo 5, 28031 Madrid, Spain * Correspondence: [email protected] (C.L.S.); [email protected] (J.A.) Abstract: Glycogen synthase kinase-3 (GSK-3) is a ubiquitously expressed serine/threonine kinase with a plethora of substrates. As a modulator of several cellular processes, GSK-3 has a central position in cell metabolism and signaling, with important roles both in physiological and pathological conditions. GSK-3 has been associated with a number of human disorders, such as neurodegenerative diseases including Alzheimer’s disease (AD). GSK-3 contributes to the hyperphosphorylation of tau protein, the main component of neurofibrillary tangles (NFTs), one of the hallmarks of AD. GSK-3 is further involved in the regulation of different neuronal processes that are dysregulated during AD pathogenesis, such as the generation of amyloid-β (Aβ) peptide or Aβ-induced cell death, axonal transport, cholinergic function, and adult neurogenesis or synaptic function. In this review, we will summarize recent data about GSK-3 involvement in these processes contributing to AD pathology, mostly focusing on the crucial interplay between GSK-3 and tau protein. We further discuss the current development of potential AD therapies targeting GSK-3 or GSK-3-phosphorylated tau. -
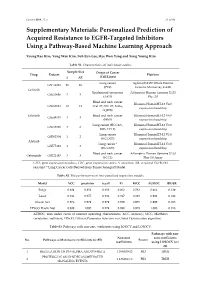
Personalized Prediction of Acquired Resistance to EGFR-Targeted Inhibitors Using a Pathway-Based Machine Learning Approach
Cancers 2019, 11, x S1 of S9 Supplementary Materials: Personalized Prediction of Acquired Resistance to EGFR-Targeted Inhibitors Using a Pathway-Based Machine Learning Approach Young Rae Kim, Yong Wan Kim, Suh Eun Lee, Hye Won Yang and Sung Young Kim Table S1. Characteristics of individual studies. Sample Size Origin of Cancer Drug Dataset Platform S AR (Cell Lines) Lung cancer Agilent-014850 Whole Human GSE34228 26 26 (PC9) Genome Microarray 4x44K Gefitinib Epidermoid carcinoma Affymetrix Human Genome U133 GSE10696 3 3 (A431) Plus 2.0 Head and neck cancer Illumina HumanHT-12 V4.0 GSE62061 12 12 (Cal-27, SSC-25, FaDu, expression beadchip SQ20B) Erlotinib Head and neck cancer Illumina HumanHT-12 V4.0 GSE49135 3 3 (HN5) expression beadchip Lung cancer (HCC827, Illumina HumanHT-12 V3.0 GSE38310 3 6 ER3, T15-2) expression beadchip Lung cancer Illumina HumanHT-12 V3.0 GSE62504 1 2 (HCC827) expression beadchip Afatinib Lung cancer * Illumina HumanHT-12 V4.0 GSE75468 1 3 (HCC827) expression beadchip Head and neck cancer Affymetrix Human Genome U133 Cetuximab GSE21483 3 3 (SCC1) Plus 2.0 Array GEO, gene expression omnibus; GSE, gene expression series; S, sensitive; AR, acquired EGFR-TKI resistant; * Lung Cancer Cells Derived from Tumor Xenograft Model. Table S2. The performances of four penalized regression models. Model ACC precision recall F1 MCC AUROC BRIER Ridge 0.889 0.852 0.958 0.902 0.782 0.964 0.129 Lasso 0.944 0.957 0.938 0.947 0.889 0.991 0.042 Elastic Net 0.978 0.979 0.979 0.979 0.955 0.999 0.023 EPSGO Elastic Net 0.989 1.000 0.979 0.989 0.978 1.000 0.018 AUROC, area under curve of receiver operating characteristic; ACC, accuracy; MCC, Matthews correlation coefficient; EPSGO, Efficient Parameter Selection via Global Optimization algorithm. -

Identification of Protein O-Glcnacylation Sites Using Electron Transfer Dissociation Mass Spectrometry on Native Peptides
Identification of protein O-GlcNAcylation sites using electron transfer dissociation mass spectrometry on native peptides Robert J. Chalkleya, Agnes Thalhammerb, Ralf Schoepfera,b, and A. L. Burlingamea,1 aDepartment of Pharmaceutical Chemistry, University of California, 600 16th Street, Genentech Hall, Suite N472, San Francisco, CA 94158; and bLaboratory for Molecular Pharmacology, and Department of Neuroscience, Physiology, and Pharmacology, University College London, Gower Street, London WC1E 6BT, United Kingdom Edited by James A. Wells, University of California, San Francisco, CA, and approved April 15, 2009 (received for review January 12, 2009) Protein O-GlcNAcylation occurs in all animals and plants and is and synaptic plasticity have been found O-GlcNAcylated (8). The implicated in modulation of a wide range of cytosolic and nuclear importance of O-GlcNAcylation is highlighted by the brain-specific protein functions, including gene silencing, nutrient and stress sens- OGT knockout mouse model that displays developmental defects ing, phosphorylation signaling, and diseases such as diabetes and that result in neonatal death (9). Most interestingly, in analogy to Alzheimer’s. The limiting factor impeding rapid progress in decipher- phosphorylation, synaptic activity leads to a dynamic modulation of ing the biological functions of protein O-GlcNAcylation has been the O-GlcNAc levels (10) and is also involved in long-term potentiation inability to easily identify exact residues of modification. We describe and memory (11). a robust, high-sensitivity strategy able to assign O-GlcNAcylation sites Phosphorylation is the most heavily studied regulatory PTM, and of native modified peptides using electron transfer dissociation mass proteomic approaches for global enrichment, detection, and char- spectrometry. -

Role of Glycogen Synthase Kinase 3B in Rapamycin-Mediated Cell Cycle
Research Article Role of Glycogen Synthase Kinase 3 B in Rapamycin-Mediated Cell Cycle Regulation and Chemosensitivity JinJiang Dong,1 Junying Peng,1 Haixia Zhang,1 Wallace H. Mondesire,1 Weiguo Jian,1 Gordon B. Mills,2 Mien-Chie Hung,1,3 and Funda Meric-Bernstam1 Departments of 1Surgical Oncology, 2Molecular Therapeutics, and 3Molecular and Cellular Oncology, University of Texas M.D. Anderson Cancer Center, Houston, Texas Abstract Introduction The mammalian target of rapamycin is a serine-threonine Rapamycin and its analogues are being actively investigated in kinase that regulates cell cycle progression. Rapamycin and clinical trials as novel targeted anticancer agents. The mammalian its analogues inhibit the mammalian target of rapamycin target of rapamycin (mTOR) is a serine-threonine kinase that and are being actively investigated in clinical trials as novel regulates cell cycle progression. The two best-studied targets of targeted anticancer agents. Although cyclin D1 is down- mTOR, eukaryotic initiation factor 4E-binding protein 1 and regulated by rapamycin, the role of this down-regulation in ribosomal p70 S6 kinase-1, are thought to modulate translation; rapamycin-mediated growth inhibition and the mechanism thus, rapamycin is thought to alter the translation of mRNA of cyclin D1 down-regulation are not well understood. Here, involved in control of the cell cycle. However, how rapamycin we show that overexpression of cyclin D1 partially over- blocks cell growth and proliferation is not well understood. comes rapamycin-induced cell cycle arrest and inhibition of Prior studies have suggested that cyclin D1 is a key target of anchorage-dependent growth in breast cancer cells.