Identification of Protein O-Glcnacylation Sites Using Electron Transfer Dissociation Mass Spectrometry on Native Peptides
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Kinome Profiling of Clinical Cancer Specimens
Published OnlineFirst March 23, 2010; DOI: 10.1158/0008-5472.CAN-09-3989 Review Cancer Research Kinome Profiling of Clinical Cancer Specimens Kaushal Parikh and Maikel P. Peppelenbosch Abstract Over the past years novel technologies have emerged to enable the determination of the transcriptome and proteome of clinical samples. These data sets will prove to be of significant value to our elucidation of the mechanisms that govern pathophysiology and may provide biological markers for future guidance in person- alized medicine. However, an equally important goal is to define those proteins that participate in signaling pathways during the disease manifestation itself or those pathways that are made active during successful clinical treatment of the disease: the main challenge now is the generation of large-scale data sets that will allow us to define kinome profiles with predictive properties on the outcome-of-disease and to obtain insight into tissue-specific analysis of kinase activity. This review describes the current techniques available to gen- erate kinome profiles of clinical tissue samples and discusses the future strategies necessary to achieve new insights into disease mechanisms and treatment targets. Cancer Res; 70(7); 2575–8. ©2010 AACR. Background the current state of the cell or tissue as characterized by pro- teomic and metabolic measurements (3). The past five years have seen an exponential increase in In this review we describe and evaluate the various tech- technological development to explore the genome and the niques that have recently become available to study the ki- proteome to understand the molecular basis of disease. How- nomes of clinical tissue samples. -
Protein Kinases Phosphorylation/Dephosphorylation Protein Phosphorylation Is One of the Most Important Mechanisms of Cellular Re
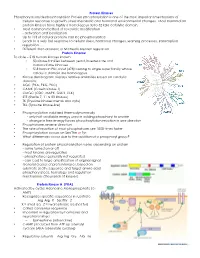
Protein Kinases Phosphorylation/dephosphorylation Protein phosphorylation is one of the most important mechanisms of cellular responses to growth, stress metabolic and hormonal environmental changes. Most mammalian protein kinases have highly a homologous 30 to 32 kDa catalytic domain. • Most common method of reversible modification - activation and localization • Up to 1/3 of cellular proteins can be phosphorylated • Leads to a very fast response to cellular stress, hormonal changes, learning processes, transcription regulation .... • Different than allosteric or Michealis Menten regulation Protein Kinome To date – 518 human kinases known • 50 kinase families between yeast, invertebrate and mammaliane kinomes • 518 human PKs, most (478) belong to single super family whose catalytic domain are homologous. • Kinase dendrogram displays relative similarities based on catalytic domains. • AGC (PKA, PKG, PKC) • CAMK (Casein kinase 1) • CMGC (CDC, MAPK, GSK3, CLK) • STE (Sterile 7, 11 & 20 kinases) • TK (Tryosine kinases memb and cyto) • TKL (Tyrosine kinase-like) • Phosphorylation stabilized thermodynamically - only half available energy used in adding phosphoryl to protein - change in free energy forces phosphorylation reaction in one direction • Phosphatases reverse direction • The rate of reaction of most phosphatases are 1000 times faster • Phosphorylation occurs on Ser/The or Tyr • What differences occur due to the addition of a phosphoryl group? • Regulation of protein phosphorylation varies depending on protein - some turned on or off -

GSK3 and Its Interactions with the PI3K/AKT/Mtor Signalling Network
Heriot-Watt University Research Gateway GSK3 and its interactions with the PI3K/AKT/mTOR signalling network Citation for published version: Hermida, MA, Kumar, JD & Leslie, NR 2017, 'GSK3 and its interactions with the PI3K/AKT/mTOR signalling network', Advances in Biological Regulation, vol. 65, pp. 5-15. https://doi.org/10.1016/j.jbior.2017.06.003 Digital Object Identifier (DOI): 10.1016/j.jbior.2017.06.003 Link: Link to publication record in Heriot-Watt Research Portal Document Version: Peer reviewed version Published In: Advances in Biological Regulation Publisher Rights Statement: © 2017 Elsevier B.V. General rights Copyright for the publications made accessible via Heriot-Watt Research Portal is retained by the author(s) and / or other copyright owners and it is a condition of accessing these publications that users recognise and abide by the legal requirements associated with these rights. Take down policy Heriot-Watt University has made every reasonable effort to ensure that the content in Heriot-Watt Research Portal complies with UK legislation. If you believe that the public display of this file breaches copyright please contact [email protected] providing details, and we will remove access to the work immediately and investigate your claim. Download date: 27. Sep. 2021 Accepted Manuscript GSK3 and its interactions with the PI3K/AKT/mTOR signalling network Miguel A. Hermida, J. Dinesh Kumar, Nick R. Leslie PII: S2212-4926(17)30124-0 DOI: 10.1016/j.jbior.2017.06.003 Reference: JBIOR 180 To appear in: Advances in Biological Regulation Received Date: 13 June 2017 Accepted Date: 23 June 2017 Please cite this article as: Hermida MA, Dinesh Kumar J, Leslie NR, GSK3 and its interactions with the PI3K/AKT/mTOR signalling network, Advances in Biological Regulation (2017), doi: 10.1016/ j.jbior.2017.06.003. -

Modified Recipe to Inhibit GSK-3 for the Living Fungal Biomaterial Manufacture
bioRxiv preprint doi: https://doi.org/10.1101/496265; this version posted December 13, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 1 Modified recipe to inhibit GSK-3 for the living fungal 2 biomaterial manufacture 1¶ 1¶ 1 1 1 3 Jinhui Chang , Po Lam Chan , Yichun Xie , Man Kit Cheung , Ka Lee Ma , Hoi Shan 1* 4 Kwan 1 5 School of Life Sciences, The Chinese University of Hong Kong, Shatin, New Territories, 6 Hong Kong 7 *Corresponding author: E-mail: [email protected] ¶ 8 These authors contributed equally to this work. 9 10 11 bioRxiv preprint doi: https://doi.org/10.1101/496265; this version posted December 13, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 12 Abstract 13 Living fungal mycelium with suppressed or abolished fruit-forming ability is a self-healing 14 substance particularly valuable biomaterial for further engineering and development in 15 applications such as monitoring/sensing environmental changes and secreting signals. The 16 ability to suppress fungal fruiting is also a useful tool for maintaining stability (e.g., shape, 17 form) of a mycelium-based biomaterial with ease and lower cost. 18 The objective of this present study is to provide a biochemical solution to regulate the fruiting 19 body formation to replace heat killing of mycelium during production. -

The Discovery of Novel Gsk3 Substrates and Their Role in the Brain
THE DISCOVERY OF NOVEL GSK3 SUBSTRATES AND THEIR ROLE IN THE BRAIN James Robinson A DISSERTATION IN FULFILMENT OF THE REQUIREMENTS FOR THE DEGREE OF DOCTOR OF PHILOSOPHY St. Vincent’s Clinical School Facility of Medicine The University of New South Wales May 2015 ORIGINALITY STATEMENT ‘I hereby declare that this submission is my own work and to the best of my knowledge it contains no materials previously published or written by another person, or substantial proportions of material which have been accepted for the award of any other degree or diploma at UNSW or any other educational institution, except where due acknowledgement is made in the thesis. Any contribution made to the research by others, with whom I have worked at UNSW or elsewhere, is explicitly acknowledged in the thesis. I also declare that the intellectual content of this thesis is the product of my own work, except to the extent that assistance from others in the project's design and conception or in style, presentation and linguistic expression is acknowledged.’ Signed ……………………………………………… Date ………………………………………………… ii |Page Abstract Bipolar Disorder (BD) is a debilitating disease that dramatically impairs people’s lives and severe cases can lead to exclusion from society and suicide. There are no clear genetic or environmental causes, and current treatments suffer from limiting side effects. Therefore, alternative intervention strategies are urgently required. Our approach is to determine mechanisms of action of current drug therapies, in the hope that they will lead to discovery of next generation therapeutic targets. Lithium has been the mainstay treatment for BD for over 50 years, although its mechanism of action is not yet clear. -

Phosphorylation-Dependent Sub-Functionalization of the Calcium-Dependent Protein Kinase CPK28
bioRxiv preprint doi: https://doi.org/10.1101/2020.10.16.338442; this version posted October 17, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Phosphorylation-dependent sub-functionalization of 2 the calcium-dependent protein kinase CPK28 3 4 5 Melissa Bredow1,#, Kyle W. Bender2,#,a, Alexandra Johnson Dingee1,b, Danalyn R. 6 Holmes1,c, Alysha Thomson1,d, Danielle Ciren1,e, Cailun A. S. Tanney1,f, Katherine E. 7 Dunning1,3, Marco Trujillo3, Steven C. Huber2, and Jacqueline Monaghan1,* 8 9 10 1 Department of Biology, Queen’s University, Kingston, Canada 11 2 Department of Plant Biology, School of Integrative Biology, University of Illinois- 12 Urbana-Champaign, Urbana-Champaign, United States of America 13 3 Department of Cell Biology, University of Freiburg, Freiburg, Germany 14 15 # These authors contributed equally to this work 16 * Corresponding author: [email protected] 17 18 19 a Current address: University of Zurich, Department of Plant and Microbial Biology, 20 Zurich, Switzerland 21 b Current address: Faculty of Law, Western University, London, Canada 22 c Current address: Center for Plant Molecular Biology, University of Tuebingen, 23 Tuebingen, Germany 24 d Current address: Faculty of Medicine, McMaster University, Hamilton, Canada 25 e Current address: Cold Spring Harbor Laboratory, Cold Spring Harbor, United States of 26 America 27 f Current address: Department of Plant Science, McGill University, Montreal, Canada 28 29 30 Keywords: calcium-dependent protein kinases, calcium-dependent protein kinase 28, 31 botrytis-induced kinase 1, phosphorylation, immune signaling, stem elongation, 32 biochemical mechanism 33 34 35 36 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.10.16.338442; this version posted October 17, 2020. -

Functional Interrogation of the Kinome Using Nucleotide Acyl Phosphates Matthew P
350 Biochemistry 2007, 46, 350-358 Functional Interrogation of the Kinome Using Nucleotide Acyl Phosphates Matthew P. Patricelli,‡ A. Katrin Szardenings,‡ Marek Liyanage, Tyzoon K. Nomanbhoy, Min Wu, Helge Weissig, Arwin Aban, Doris Chun, Stephen Tanner, and John W. Kozarich* ActiVX Biosciences, 11025 North Torrey Pines Road, La Jolla, California 92037 ReceiVed October 15, 2006; ReVised Manuscript ReceiVed December 2, 2006 ABSTRACT: The central role of protein kinases in signal transduction pathways has generated intense interest in targeting these enzymes for a wide range of therapeutic indications. Here we report a method for identifying and quantifying protein kinases in any biological sample or tissue from any species. The procedure relies on acyl phosphate-containing nucleotides, prepared from a biotin derivative and ATP or ADP. The acyl phosphate probes react selectively and covalently at the ATP binding sites of at least 75% of the known human protein kinases. Biotinylated peptide fragments from labeled proteomes are captured and then sequenced and identified using a mass spectrometry-based analysis platform to determine the kinases present and their relative levels. Further, direct competition between the probes and inhibitors can be assessed to determine inhibitor potency and selectivity against native protein kinases, as well as hundreds of other ATPases. The ability to broadly profile kinase activities in native proteomes offers an exciting prospect for both target discovery and inhibitor selectivity profiling. Protein kinases represent the single largest mammalian for protein kinase inhibitors, highlight the need for improved enzyme family with more than 500 members in the human methods for the comprehensive screening of protein kinases proteome. -

Kinome-Wide Selectivity Profiling of ATP-Competitive Mammalian Target of Rapamycin (Mtor) Inhibitors and Characterization Of
Supplemental Material can be found at: http://www.jbc.org/content/suppl/2012/01/05/M111.304485.DC1.html THE JOURNAL OF BIOLOGICAL CHEMISTRY VOL. 287, NO. 13, pp. 9742–9752, March 23, 2012 © 2012 by The American Society for Biochemistry and Molecular Biology, Inc. Published in the U.S.A. Kinome-wide Selectivity Profiling of ATP-competitive Mammalian Target of Rapamycin (mTOR) Inhibitors and Characterization of Their Binding Kinetics*□S Received for publication, September 15, 2011, and in revised form, December 29, 2011 Published, JBC Papers in Press, January 5, 2012, DOI 10.1074/jbc.M111.304485 Qingsong Liu,a Sivapriya Kirubakaran,a Wooyoung Hur,a Mario Niepel,b Kenneth Westover,c Carson C. Thoreen,a1 Jinhua Wang,a Jing Ni,a Matthew P. Patricelli,d Kurt Vogel,e Steve Riddle,e David L. Waller,a Ryan Traynor,f Takaomi Sanda,g2 Zheng Zhao,h Seong A. Kang,i,j Jean Zhao,a A. Thomas Look,g Peter K. Sorger,b David M. Sabatini,i,j,k and Nathanael S. Graya3 From the aDepartment of Cancer Biology, Dana Farber Cancer Institute, Department of Biological Chemistry and Molecular Pharmacology, bCenter for Cell Decision Processes, and Department of Systems Biology, Harvard Medical School, Boston, Massachusetts 02115, the cHarvard Radiation Oncology Program, Boston, Massachusetts 02115, the dActivX Biosciences, Inc., La Jolla, California 92037, eInvitrogen Corp., Madison, Wisconsin 53719, the fNational Centre for Protein Kinase Profiling, Dundee Division of Signal Transduction Therapy, College of Life Sciences, University of Dundee, Dundee DD1 5EH, Scotland, United Downloaded from Kingdom, the gDepartment of Pediatric Oncology, Dana-Farber Cancer Institute, Boston, Massachusetts 02215, the hHigh Magnetic Field Laboratory, Chinese Academy of Science, P. -

Page 1 Exploring the Understudied Human Kinome For
bioRxiv preprint doi: https://doi.org/10.1101/2020.04.02.022277; this version posted June 30, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. Exploring the understudied human kinome for research and therapeutic opportunities Nienke Moret1,2,*, Changchang Liu1,2,*, Benjamin M. Gyori2, John A. Bachman,2, Albert Steppi2, Rahil Taujale3, Liang-Chin Huang3, Clemens Hug2, Matt Berginski1,4,5, Shawn Gomez1,4,5, Natarajan Kannan,1,3 and Peter K. Sorger1,2,† *These authors contributed equally † Corresponding author 1The NIH Understudied Kinome Consortium 2Laboratory of Systems Pharmacology, Department of Systems Biology, Harvard Program in Therapeutic Science, Harvard Medical School, Boston, Massachusetts 02115, USA 3 Institute of Bioinformatics, University of Georgia, Athens, GA, 30602 USA 4 Department of Pharmacology, The University of North Carolina at Chapel Hill, Chapel Hill, NC 27599, USA 5 Joint Department of Biomedical Engineering at the University of North Carolina at Chapel Hill and North Carolina State University, Chapel Hill, NC 27599, USA Key Words: kinase, human kinome, kinase inhibitors, drug discovery, cancer, cheminformatics, † Peter Sorger Warren Alpert 432 200 Longwood Avenue Harvard Medical School, Boston MA 02115 [email protected] cc: [email protected] 617-432-6901 ORCID Numbers Peter K. Sorger 0000-0002-3364-1838 Nienke Moret 0000-0001-6038-6863 Changchang Liu 0000-0003-4594-4577 Ben Gyori 0000-0001-9439-5346 John Bachman 0000-0001-6095-2466 Albert Steppi 0000-0001-5871-6245 Page 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.04.02.022277; this version posted June 30, 2020. -

Kinome Profiling to Predict Sensitivity to MAPK Inhibition in Melanoma
cancers Article Kinome Profiling to Predict Sensitivity to MAPK Inhibition in Melanoma and to Provide New Insights into Intrinsic and Acquired Mechanism of Resistance 1, , 2, 1 3 Mohamad Krayem * y , Philippe Aftimos y, Ahmad Najem , Tim van den Hooven , Adriënne van den Berg 3, Liesbeth Hovestad-Bijl 3, Rik de Wijn 3, Riet Hilhorst 3 , Rob Ruijtenbeek 3, Malak Sabbah 1, Joseph Kerger 2, Ahmad Awada 1,2 , Fabrice Journe 1 and Ghanem E. Ghanem 1 1 Laboratory of Oncology and Experimental Surgery, Institut Jules Bordet, Université Libre de Bruxelles, 1000 Brussels, Belgium; [email protected] (A.N.); [email protected] (M.S.); [email protected] (A.A.); [email protected] (F.J.); [email protected] (G.E.G.) 2 Medical Oncology Clinic, Institut Jules Bordet, Université Libre de Bruxelles, 1000 Brussels, Belgium; [email protected] (P.A.); [email protected] (J.K.) 3 PamGene International BV, 5211HH ’s-Hertogenbosch, The Netherlands; [email protected] (T.v.d.H.); [email protected] (A.v.d.B.); [email protected] (L.H.-B.); [email protected] (R.d.W.); [email protected] (R.H.); [email protected] (R.R.) * Correspondence: [email protected] Both authors equally contributed to this manuscript. y Received: 20 January 2020; Accepted: 20 February 2020; Published: 22 February 2020 Abstract: Mitogen-activated protein kinase (MAPK) inhibition with the combination of BRAF (Rapidly Accelerated Fibrosarcoma) and MEK (Mitogen-activated protein kinase kinase) inhibitors has become the standard of first-line therapy of metastatic melanoma harbouring BRAF V600 mutations. However, about half of the patients present with primary resistance while the remaining develop secondary resistance under prolonged treatment. -

Towards Application of Small Molecules in Dentin Repair
International Journal of Molecular Sciences Article Drug Repurposing in Dentistry: Towards Application of Small Molecules in Dentin Repair Anahid A. Birjandi, Fernanda R. Suzano and Paul T. Sharpe * Centre for Craniofacial and Regenerative Biology, Guy’s Hospital, King’s College London, London SE1 9RT, UK; [email protected] (A.A.B.); [email protected] (F.R.S.) * Correspondence: [email protected] Received: 30 July 2020; Accepted: 29 August 2020; Published: 2 September 2020 Abstract: One of the main goals of dentistry is the natural preservation of the tooth structure following damage. This is particularly implicated in deep dental cavities affecting dentin and pulp, where odontoblast survival is jeopardized. This activates pulp stem cells and differentiation of new odontoblast-like cells, accompanied by increased Wnt signaling. Our group has shown that delivery of small molecule inhibitors of GSK3 stimulates Wnt/β-catenin signaling in the tooth cavity with pulp exposure and results in effective promotion of dentin repair. Small molecules are a good therapeutic option due to their ability to pass across cell membranes and reach target. Here, we investigate a range of non-GSK3 target small molecules that are currently used for treatment of various medical conditions based on other kinase inhibitory properties. We analyzed the ability of these drugs to stimulate Wnt signaling activity by off-target inhibition of GSK3. Our results show that a c-Met inhibitor, has the ability to stimulate Wnt/β-catenin pathway in dental pulp cells in vitro at low concentrations. This work is an example of drug repurposing for dentistry and suggests a candidate drug to be tested in vivo for natural dentin repair. -

The Evolving World of Pseudoenzymes: Proteins, Prejudice and Zombies Patrick A
Eyers and Murphy BMC Biology (2016) 14:98 DOI 10.1186/s12915-016-0322-x QUESTION AND ANSWER Open Access The evolving world of pseudoenzymes: proteins, prejudice and zombies Patrick A. Eyers1* and James M. Murphy2,3* clear patterns of amino acid conservation in classic Abstract enzyme families (for example amongst the protein Pseudoenzymes are catalytically deficient variants of kinases [2]). These patterns often explain how an enzymes that are represented in all major enzyme enzyme works or is likely to be regulated [3, 4]. They families. Their regulatory functions in signalling mayinclude,forexample,thepresenceofametal- pathways are shedding new light on the non-catalytic binding motif or a conserved network of amino acids functions of active enzymes, and are suggesting new that form a catalytic signature. ways to target cellular signalling mechanisms with What has become apparent now that thousands and drugs. thousands of genomes have been sequenced is that they contain, within every enzyme family where we look, di- vergent examples of proteins that are clearly part of What is a pseudoenzyme? large enzyme families but for which basic enzyme func- A pseudoenzyme contains a protein domain with a fold tion (increasing the rate of a chemical reaction) is pre- that resembles, or is predicted to resemble, a catalytically dicted to be lacking. These ‘dead’ enzymes have amino active, conventional protein domain counterpart. How- acid sequence deviations from the catalytic residues of ever, the pseudoenzyme possesses vestigial or zero cata- counterparts that would be predicted to confer enzyme lytic activity owing to the absence of key catalytic amino function—or, in an experimentally ideal world, where acids or motifs.