Associated R-Loops at Near-Nucleotide Resolution Using Bisdrip-Seq Jason G Dumelie, Samie R Jaffrey*
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Recombinant Human ARFIP2 Protein Catalog Number: ATGP1695
Recombinant human ARFIP2 protein Catalog Number: ATGP1695 PRODUCT INPORMATION Expression system E.coli Domain 1-341aa UniProt No. P53365 NCBI Accession No. NP_036534 Alternative Names Arfaptin 2, POR1 PRODUCT SPECIFICATION Molecular Weight 40.2 kDa (364aa) confirmed by MALDI-TOF Concentration 0.25mg/ml (determined by Bradford assay) Formulation Liquid in. 20mM Tris-HCl buffer (pH 8.0) containing 0.2M NaCl, 40% glycerol, 1mM DTT Purity > 90% by SDS-PAGE Tag His-Tag Application SDS-PAGE Storage Condition Can be stored at +2C to +8C for 1 week. For long term storage, aliquot and store at -20C to -80C. Avoid repeated freezing and thawing cycles. BACKGROUND Description Arfaptin 2, also known as ARFIP2, is a Rac1 binding protein necessary for Rac-mediated actin polymerization and the subsequent formation of membrane ruffles and lamellipodia. ARFIP2 has also been shown to interact with the ADP ribosylation factor ARF6, a GTPase that associates with the plasma membrane and intracellular endosome vesicles, in a GTP dependent manner. Arfaptin 2 also regulates the aggregation of mutant Huntingtin protein by possibly impairing proteasome function. Expression of ARFIP2 was shown to be increased at sites of neurodegeneration. Recombinant human ARFIP2 protein, fused to His-tag at N-terminus, was expressed in E. coli 1 Recombinant human ARFIP2 protein Catalog Number: ATGP1695 and purified by using conventional chromatography techniques. Amino acid Sequence MGSSHHHHHH SSGLVPRGSH MGSMTDGILG KAATMEIPIH GNGEARQLPE DDGLEQDLQQ VMVSGPNLNE TSIVSGGYGG SGDGLIPTGS GRHPSHSTTP SGPGDEVARG IAGEKFDIVK KWGINTYKCT KQLLSERFGR GSRTVDLELE LQIELLRETK RKYESVLQLG RALTAHLYSL LQTQHALGDA FADLSQKSPE LQEEFGYNAE TQKLLCKNGE TLLGAVNFFV SSINTLVTKT MEDTLMTVKQ YEAARLEYDA YRTDLEELSL GPRDAGTRGR LESAQATFQA HRDKYEKLRG DVAIKLKFLE ENKIKVMHKQ LLLFHNAVSA YFAGNQKQLE QTLQQFNIKL RPPGAEKPSW LEEQ General References D'Souza Schorey C., et al. -

Download Download
Supplementary Figure S1. Results of flow cytometry analysis, performed to estimate CD34 positivity, after immunomagnetic separation in two different experiments. As monoclonal antibody for labeling the sample, the fluorescein isothiocyanate (FITC)- conjugated mouse anti-human CD34 MoAb (Mylteni) was used. Briefly, cell samples were incubated in the presence of the indicated MoAbs, at the proper dilution, in PBS containing 5% FCS and 1% Fc receptor (FcR) blocking reagent (Miltenyi) for 30 min at 4 C. Cells were then washed twice, resuspended with PBS and analyzed by a Coulter Epics XL (Coulter Electronics Inc., Hialeah, FL, USA) flow cytometer. only use Non-commercial 1 Supplementary Table S1. Complete list of the datasets used in this study and their sources. GEO Total samples Geo selected GEO accession of used Platform Reference series in series samples samples GSM142565 GSM142566 GSM142567 GSM142568 GSE6146 HG-U133A 14 8 - GSM142569 GSM142571 GSM142572 GSM142574 GSM51391 GSM51392 GSE2666 HG-U133A 36 4 1 GSM51393 GSM51394 only GSM321583 GSE12803 HG-U133A 20 3 GSM321584 2 GSM321585 use Promyelocytes_1 Promyelocytes_2 Promyelocytes_3 Promyelocytes_4 HG-U133A 8 8 3 GSE64282 Promyelocytes_5 Promyelocytes_6 Promyelocytes_7 Promyelocytes_8 Non-commercial 2 Supplementary Table S2. Chromosomal regions up-regulated in CD34+ samples as identified by the LAP procedure with the two-class statistics coded in the PREDA R package and an FDR threshold of 0.5. Functional enrichment analysis has been performed using DAVID (http://david.abcc.ncifcrf.gov/) -

Supplemental Information
Supplemental information Dissection of the genomic structure of the miR-183/96/182 gene. Previously, we showed that the miR-183/96/182 cluster is an intergenic miRNA cluster, located in a ~60-kb interval between the genes encoding nuclear respiratory factor-1 (Nrf1) and ubiquitin-conjugating enzyme E2H (Ube2h) on mouse chr6qA3.3 (1). To start to uncover the genomic structure of the miR- 183/96/182 gene, we first studied genomic features around miR-183/96/182 in the UCSC genome browser (http://genome.UCSC.edu/), and identified two CpG islands 3.4-6.5 kb 5’ of pre-miR-183, the most 5’ miRNA of the cluster (Fig. 1A; Fig. S1 and Seq. S1). A cDNA clone, AK044220, located at 3.2-4.6 kb 5’ to pre-miR-183, encompasses the second CpG island (Fig. 1A; Fig. S1). We hypothesized that this cDNA clone was derived from 5’ exon(s) of the primary transcript of the miR-183/96/182 gene, as CpG islands are often associated with promoters (2). Supporting this hypothesis, multiple expressed sequences detected by gene-trap clones, including clone D016D06 (3, 4), were co-localized with the cDNA clone AK044220 (Fig. 1A; Fig. S1). Clone D016D06, deposited by the German GeneTrap Consortium (GGTC) (http://tikus.gsf.de) (3, 4), was derived from insertion of a retroviral construct, rFlpROSAβgeo in 129S2 ES cells (Fig. 1A and C). The rFlpROSAβgeo construct carries a promoterless reporter gene, the β−geo cassette - an in-frame fusion of the β-galactosidase and neomycin resistance (Neor) gene (5), with a splicing acceptor (SA) immediately upstream, and a polyA signal downstream of the β−geo cassette (Fig. -

MYC-Containing Amplicons in Acute Myeloid Leukemia: Genomic Structures, Evolution, and Transcriptional Consequences
Leukemia (2018) 32:2152–2166 https://doi.org/10.1038/s41375-018-0033-0 ARTICLE Acute myeloid leukemia Corrected: Correction MYC-containing amplicons in acute myeloid leukemia: genomic structures, evolution, and transcriptional consequences 1 1 2 2 1 Alberto L’Abbate ● Doron Tolomeo ● Ingrid Cifola ● Marco Severgnini ● Antonella Turchiano ● 3 3 1 1 1 Bartolomeo Augello ● Gabriella Squeo ● Pietro D’Addabbo ● Debora Traversa ● Giulia Daniele ● 1 1 3 3 4 Angelo Lonoce ● Mariella Pafundi ● Massimo Carella ● Orazio Palumbo ● Anna Dolnik ● 5 5 6 2 7 Dominique Muehlematter ● Jacqueline Schoumans ● Nadine Van Roy ● Gianluca De Bellis ● Giovanni Martinelli ● 3 4 8 1 Giuseppe Merla ● Lars Bullinger ● Claudia Haferlach ● Clelia Tiziana Storlazzi Received: 4 August 2017 / Revised: 27 October 2017 / Accepted: 13 November 2017 / Published online: 22 February 2018 © The Author(s) 2018. This article is published with open access Abstract Double minutes (dmin), homogeneously staining regions, and ring chromosomes are vehicles of gene amplification in cancer. The underlying mechanism leading to their formation as well as their structure and function in acute myeloid leukemia (AML) remain mysterious. We combined a range of high-resolution genomic methods to investigate the architecture and expression pattern of amplicons involving chromosome band 8q24 in 23 cases of AML (AML-amp). This 1234567890();,: revealed that different MYC-dmin architectures can coexist within the same leukemic cell population, indicating a step-wise evolution rather than a single event origin, such as through chromothripsis. This was supported also by the analysis of the chromothripsis criteria, that poorly matched the model in our samples. Furthermore, we found that dmin could evolve toward ring chromosomes stabilized by neocentromeres. -

Mutational Landscape of Uterine and Ovarian Carcinosarcomas Implicates Histone Genes in Epithelial–Mesenchymal Transition
Mutational landscape of uterine and ovarian carcinosarcomas implicates histone genes in epithelial–mesenchymal transition Siming Zhaoa,b, Stefania Bellonec, Salvatore Lopezc, Durga Thakrala,b, Carlton Schwabc, Diana P. Englishc, Jonathan Blackc, Emiliano Coccoc, Jungmin Choia,b, Luca Zammataroc, Federica Predolinic, Elena Bonazzolic, Mark Bia,b, Natalia Buzad, Pei Huid, Serena Wongd, Maysa Abu-Khalafe, Antonella Ravaggif, Eliana Bignottif, Elisabetta Bandieraf, Chiara Romanif, Paola Todeschinif, Renata Tassif, Laura Zanottif, Franco Odicinof, Sergio Pecorellif, Carla Donzellig, Laura Ardighierig, Fabio Facchettig, Marcella Falchettig, Dan-Arin Silasic, Elena Ratnerc, Masoud Azodic, Peter E. Schwartzc, Shrikant Manea,b, Roberto Angiolih, Corrado Terranovah, Charles Matthew Quicki, Babak Edrakij, Kaya Bilgüvara,b, Moses Leek, Murim Choik, Amy L. Stieglerl, Titus J. Boggonl, Joseph Schlessingerl, Richard P. Liftona,b,m,1, and Alessandro D. Santinc aDepartment of Genetics, Yale University School of Medicine, New Haven, CT 06510; bHoward Hughes Medical Institute, Yale University School of Medicine, New Haven, CT 06510; cDepartment of Obstetrics, Gynecology & Reproductive Sciences, Yale University School of Medicine, New Haven, CT 06510; dDepartment of Pathology, Yale University School of Medicine, New Haven, CT 06510; eInternal Medicine & Oncology, Yale University School of Medicine, New Haven, CT 06510; f“Angelo Nocivelli” Institute of Molecular Medicine, Department of Obstetrics & Gynecology, University of Brescia, 25100 Brescia, Italy; -

Usbiological Datasheet
HIST1H2BG, NT (HIST1H2BC, H2BFL, Histone H2B type 1-C/E/F/G/I, Histone H2B.1 A, Histone H2B.a, Histone H2B.g, Histone H2B.h, Histone H2B.k, Histone H2B.l) (Biotin) Catalog number 036594-Biotin Supplier United States Biological Histones are basic nuclear proteins that are responsible for the nucleosome structure of the chromosomal fiber in eukaryotes. Two molecules of each of the four core histones (H2A, H2B, H3, and H4) form an octamer, around which approximately 146 bp of DNA is wrapped in repeating units, called nucleosomes. The linker histone, H1, interacts with linker DNA between nucleosomes and functions in the compaction of chromatin into higher order structures. This gene is intronless and encodes a member of the histone H2B family. Transcripts from this gene lack polyA tails but instead contain a palindromic termination element. This gene is found in the large histone gene cluster on chromosome 6. [provided by RefSeq]. Applications Suitable for use in Western Blot, ELISA Recommended Dilution ELISA: 1:1,000 Western Blot: 1:100-500 Storage and Stability Store product at 4°C if to be used immediately within two weeks. For long-term storage, aliquot to avoid repeated freezing and thawing and store at -20°C. Aliquots are stable at -20°C for 12 months after receipt. Dilute required amount only prior to immediate use. Further dilutions can be made in assay buffer. For maximum recovery of product, centrifuge the original vial after thawing and prior to removing the cap. Note Applications are based on unconjugated antibody. Immunogen HIST1H2BG antibody is generated from rabbits immunized with a KLH conjugated synthetic peptide between 1-30 amino acids from the N-terminal region of human HIST1H2BG. -

Prediction of Host-Virus Interaction Networks
Prediction of host-virus interaction networks Janet M. Doolittle A dissertation submitted to the faculty of the University of North Carolina at Chapel Hill in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Curriculum of Bioinformatics and Computational Biology. Chapel Hill 2012 Approved by: Tim Elston, PhD Shawn Gomez, Eng.Sc.D. Klaus Hahn, PhD Keith Burridge, PhD Anne Taylor, PhD Jennifer Webster-Cyriaque, PhD ©2012 Janet M. Doolittle ALL RIGHTS RESERVED ii ABSTRACT JANET DOOLITTLE: Prediction of host-virus interaction networks (Under the direction of Shawn Gomez) As with other viral pathogens, HIV-1 and dengue virus (DENV) are dependent on their hosts for the bulk of the functions necessary for viral survival and replication. Thus, successful infection depends on the pathogen's ability to manipulate the biological pathways and processes of the organism it infects, while avoiding elimination by the immune system. Protein-protein interactions are one avenue through which viruses can connect with and exploit these host cellular pathways and processes. We developed a computational approach to predict interactions between HIV and human proteins based on structural similarity of 9 HIV-1 proteins to human proteins having known interactions. Using functional data from RNAi studies as a filter, we generated over 2,000 interaction predictions between HIV proteins and 406 unique human proteins. Additional filtering based on Gene Ontology cellular component annotation reduced the number of predictions to 502 interactions involving 137 human proteins. We find numerous known interactions as well as novel interactions showing significant functional relevance based on supporting Gene Ontology and literature evidence. -

A Yeast Phenomic Model for the Influence of Warburg Metabolism on Genetic Buffering of Doxorubicin Sean M
Santos and Hartman Cancer & Metabolism (2019) 7:9 https://doi.org/10.1186/s40170-019-0201-3 RESEARCH Open Access A yeast phenomic model for the influence of Warburg metabolism on genetic buffering of doxorubicin Sean M. Santos and John L. Hartman IV* Abstract Background: The influence of the Warburg phenomenon on chemotherapy response is unknown. Saccharomyces cerevisiae mimics the Warburg effect, repressing respiration in the presence of adequate glucose. Yeast phenomic experiments were conducted to assess potential influences of Warburg metabolism on gene-drug interaction underlying the cellular response to doxorubicin. Homologous genes from yeast phenomic and cancer pharmacogenomics data were analyzed to infer evolutionary conservation of gene-drug interaction and predict therapeutic relevance. Methods: Cell proliferation phenotypes (CPPs) of the yeast gene knockout/knockdown library were measured by quantitative high-throughput cell array phenotyping (Q-HTCP), treating with escalating doxorubicin concentrations under conditions of respiratory or glycolytic metabolism. Doxorubicin-gene interaction was quantified by departure of CPPs observed for the doxorubicin-treated mutant strain from that expected based on an interaction model. Recursive expectation-maximization clustering (REMc) and Gene Ontology (GO)-based analyses of interactions identified functional biological modules that differentially buffer or promote doxorubicin cytotoxicity with respect to Warburg metabolism. Yeast phenomic and cancer pharmacogenomics data were integrated to predict differential gene expression causally influencing doxorubicin anti-tumor efficacy. Results: Yeast compromised for genes functioning in chromatin organization, and several other cellular processes are more resistant to doxorubicin under glycolytic conditions. Thus, the Warburg transition appears to alleviate requirements for cellular functions that buffer doxorubicin cytotoxicity in a respiratory context. -

Insights Into Functional Connectivity in Mammalian Signal Transduction Pathways by Pairwise
bioRxiv preprint doi: https://doi.org/10.1101/2019.12.30.891200; this version posted December 30, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Insights into Functional Connectivity in Mammalian Signal Transduction Pathways by Pairwise Comparison of Protein Interaction Partners of Critical Signaling Hubs Chilakamarti V. Ramana * Department of Medicine, Dartmouth-Hitchcock Medical Center, Lebanon, NH 03766, USA *Correspondence should be addressed: Chilakamarti V .Ramana, Telephone. (603)-738-2507, E-mail: [email protected] ORCID ID: /0000-0002-5153-8252 1 bioRxiv preprint doi: https://doi.org/10.1101/2019.12.30.891200; this version posted December 30, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Abstract Growth factors and cytokines activate signal transduction pathways and regulate gene expression in eukaryotes. Intracellular domains of activated receptors recruit several protein kinases as well as transcription factors that serve as platforms or hubs for the assembly of multi-protein complexes. The signaling hubs involved in a related biologic function often share common interaction proteins and target genes. This functional connectivity suggests that a pairwise comparison of protein interaction partners of signaling hubs and network analysis of common partners and their expression analysis might lead to the identification of critical nodes in cellular signaling. -

DIPPER, a Spatiotemporal Proteomics Atlas of Human Intervertebral Discs
TOOLS AND RESOURCES DIPPER, a spatiotemporal proteomics atlas of human intervertebral discs for exploring ageing and degeneration dynamics Vivian Tam1,2†, Peikai Chen1†‡, Anita Yee1, Nestor Solis3, Theo Klein3§, Mateusz Kudelko1, Rakesh Sharma4, Wilson CW Chan1,2,5, Christopher M Overall3, Lisbet Haglund6, Pak C Sham7, Kathryn Song Eng Cheah1, Danny Chan1,2* 1School of Biomedical Sciences, , The University of Hong Kong, Hong Kong; 2The University of Hong Kong Shenzhen of Research Institute and Innovation (HKU-SIRI), Shenzhen, China; 3Centre for Blood Research, Faculty of Dentistry, University of British Columbia, Vancouver, Canada; 4Proteomics and Metabolomics Core Facility, The University of Hong Kong, Hong Kong; 5Department of Orthopaedics Surgery and Traumatology, HKU-Shenzhen Hospital, Shenzhen, China; 6Department of Surgery, McGill University, Montreal, Canada; 7Centre for PanorOmic Sciences (CPOS), The University of Hong Kong, Hong Kong Abstract The spatiotemporal proteome of the intervertebral disc (IVD) underpins its integrity *For correspondence: and function. We present DIPPER, a deep and comprehensive IVD proteomic resource comprising [email protected] 94 genome-wide profiles from 17 individuals. To begin with, protein modules defining key †These authors contributed directional trends spanning the lateral and anteroposterior axes were derived from high-resolution equally to this work spatial proteomes of intact young cadaveric lumbar IVDs. They revealed novel region-specific Present address: ‡Department profiles of regulatory activities -

Methylation of Leukocyte DNA and Ovarian Cancer
Fridley et al. BMC Medical Genomics 2014, 7:21 http://www.biomedcentral.com/1755-8794/7/21 RESEARCH ARTICLE Open Access Methylation of leukocyte DNA and ovarian cancer: relationships with disease status and outcome Brooke L Fridley1*, Sebastian M Armasu2, Mine S Cicek2, Melissa C Larson2, Chen Wang2, Stacey J Winham2, Kimberly R Kalli3, Devin C Koestler1,4, David N Rider2, Viji Shridhar5, Janet E Olson2, Julie M Cunningham5 and Ellen L Goode2 Abstract Background: Genome-wide interrogation of DNA methylation (DNAm) in blood-derived leukocytes has become feasible with the advent of CpG genotyping arrays. In epithelial ovarian cancer (EOC), one report found substantial DNAm differences between cases and controls; however, many of these disease-associated CpGs were attributed to differences in white blood cell type distributions. Methods: We examined blood-based DNAm in 336 EOC cases and 398 controls; we included only high-quality CpG loci that did not show evidence of association with white blood cell type distributions to evaluate association with case status and overall survival. Results: Of 13,816 CpGs, no significant associations were observed with survival, although eight CpGs associated with survival at p < 10−3, including methylation within a CpG island located in the promoter region of GABRE (p = 5.38 x 10−5, HR = 0.95). In contrast, 53 CpG methylation sites were significantly associated with EOC risk (p <5 x10−6). The top association was observed for the methylation probe cg04834572 located approximately 315 kb upstream of DUSP13 (p = 1.6 x10−14). Other disease-associated CpGs included those near or within HHIP (cg14580567; p =5.6x10−11), HDAC3 (cg10414058; p = 6.3x10−12), and SCR (cg05498681; p = 4.8x10−7). -
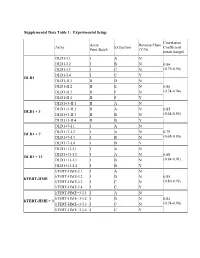
Table 3: Average Gene Expression Profiles by Chromosome
Supplemental Data Table 1: Experimental Setup Correlation Array Reverse Fluor Array Extraction Coefficient Print Batch (Y/N) mean (range) DLD1-I.1 I A N DLD1-I.2 I B N 0.86 DLD1-I.3 I C N (0.79-0.90) DLD1-I.4 I C Y DLD1 DLD1-II.1 II D N DLD1-II.2 II E N 0.86 DLD1-II.3 II F N (0.74-0.94) DLD1-II.4 II F Y DLD1+3-II.1 II A N DLD1+3-II.2 II A N 0.85 DLD1 + 3 DLD1+3-II.3 II B N (0.64-0.95) DLD1+3-II.4 II B Y DLD1+7-I.1 I A N DLD1+7-I.2 I A N 0.79 DLD1 + 7 DLD1+7-I.3 I B N (0.68-0.90) DLD1+7-I.4 I B Y DLD1+13-I.1 I A N DLD1+13-I.2 I A N 0.88 DLD1 + 13 DLD1+13-I.3 I B N (0.84-0.91) DLD1+13-I.4 I B Y hTERT-HME-I.1 I A N hTERT-HME-I.2 I B N 0.85 hTERT-HME hTERT-HME-I.3 I C N (0.80-0.92) hTERT-HME-I.4 I C Y hTERT-HME+3-I.1 I A N hTERT-HME+3-I.2 I B N 0.84 hTERT-HME + 3 hTERT-HME+3-I.3 I C N (0.74-0.90) hTERT-HME+3-I.4 I C Y Supplemental Data Table 2: Average gene expression profiles by chromosome arm DLD1 hTERT-HME Ratio.7 Ratio.1 Ratio.3 Ratio.3 Chrom.