Exploring the Oxidative Stress Mechanism of Buyang Huanwu Decoction in Intervention of Vascular Dementia Based on Systems Biology Strategy
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

CSE642 Final Version
Eindhoven University of Technology MASTER Dimensionality reduction of gene expression data Arts, S. Award date: 2018 Link to publication Disclaimer This document contains a student thesis (bachelor's or master's), as authored by a student at Eindhoven University of Technology. Student theses are made available in the TU/e repository upon obtaining the required degree. The grade received is not published on the document as presented in the repository. The required complexity or quality of research of student theses may vary by program, and the required minimum study period may vary in duration. General rights Copyright and moral rights for the publications made accessible in the public portal are retained by the authors and/or other copyright owners and it is a condition of accessing publications that users recognise and abide by the legal requirements associated with these rights. • Users may download and print one copy of any publication from the public portal for the purpose of private study or research. • You may not further distribute the material or use it for any profit-making activity or commercial gain Eindhoven University of Technology MASTER THESIS Dimensionality Reduction of Gene Expression Data Author: S. (Sako) Arts Daily Supervisor: dr. V. (Vlado) Menkovski Graduation Committee: dr. V. (Vlado) Menkovski dr. D.C. (Decebal) Mocanu dr. N. (Nikolay) Yakovets May 16, 2018 v1.0 Abstract The focus of this thesis is dimensionality reduction of gene expression data. I propose and test a framework that deploys linear prediction algorithms resulting in a reduced set of selected genes relevant to a specified case. Abstract In cancer research there is a large need to automate parts of the process of diagnosis, this is mainly to reduce cost, make it faster and more accurate. -

Caractérisation De Nouveaux Gènes Et Polymorphismes Potentiellement Impliqués Dans Les Interactions Hôtes-Pathogènes
Aix-Marseille Université, Faculté de Médecine de Marseille Ecole Doctorale des Sciences de la Vie et de la Santé THÈSE DE DOCTORAT Présentée par Charbel ABOU-KHATER Date et lieu de naissance: 08-Juilllet-1990, Zahlé, LIBAN En vue de l’obtention du grade de Docteur de l’Université d’Aix-Marseille Mention: Biologie, Spécialité: Microbiologie Caractérisation de nouveaux gènes et polymorphismes potentiellement impliqués dans les interactions hôtes-pathogènes Publiquement soutenue le 5 Juillet 2017 devant le jury composé de : Pr. Daniel OLIVE Directeur de Thèse Pr. Brigitte CROUAU-ROY Rapporteur Dr. Benoît FAVIER Rapporteur Dr. Pierre PONTAROTTI Examinateur Thèse codirigée par Pr. Daniel OLIVE et Dr Laurent ABI-RACHED Laboratoires d’accueil URMITE Research Unit on Emerging Infectious and Tropical Diseases, UMR 6236, Faculty of Medicine, 27, Boulevard Jean Moulin, 13385 Marseille, France CRCM, Centre de Recherche en Cancérologie de Marseille,Inserm 1068, 27 Boulevard Leï Roure, BP 30059, 13273 Marseille Cedex 09, France 2 Acknowledgements First and foremost, praises and thanks to God, Holy Mighty, Holy Immortal, All-Holy Trinity, for His showers of blessings throughout my whole life and to whom I owe my very existence. Glory to the Father, and to the Son, and to the Holy Spirit: now and ever and unto ages of ages. I would like to express my sincere gratitude to my advisors Prof. Daniel Olive and Dr. Laurent Abi-Rached, for the continuous support, for their patience, motivation, and immense knowledge. Someday, I hope to be just like you. A special thanks to my “Godfather” who perfectly fulfilled his role, Dr. -

Unique Adaptations in Neonatal Hepatic Transcriptome, Nutrient
Palombo et al. BMC Genomics (2021) 22:280 https://doi.org/10.1186/s12864-021-07538-w RESEARCH ARTICLE Open Access Unique adaptations in neonatal hepatic transcriptome, nutrient signaling, and one- carbon metabolism in response to feeding ethyl cellulose rumen-protected methionine during late-gestation in Holstein cows Valentino Palombo1,2, Abdulrahman Alharthi2,3, Fernanda Batistel4, Claudia Parys5, Jessie Guyader5, Erminio Trevisi6, Mariasilvia D’Andrea1 and Juan J. Loor2* Abstract Background: Methionine (Met) supply during late-pregnancy enhances fetal development in utero and leads to greater rates of growth during the neonatal period. Due to its central role in coordinating nutrient and one-carbon metabolism along with immune responses of the newborn, the liver could be a key target of the programming effects induced by dietary methyl donors such as Met. To address this hypothesis, liver biopsies from 4-day old calves (n = 6/group) born to Holstein cows fed a control or the control plus ethyl-cellulose rumen-protected Met for the last 28 days prepartum were used for DNA methylation, transcriptome, metabolome, proteome, and one-carbon metabolism enzyme activities. Results: Although greater withers and hip height at birth in Met calves indicated better development in utero, there were no differences in plasma systemic physiological indicators. RNA-seq along with bioinformatics and transcription factor regulator analyses revealed broad alterations in ‘Glucose metabolism’, ‘Lipid metabolism, ‘Glutathione’, and ‘Immune System’ metabolism due to enhanced maternal Met supply. Greater insulin sensitivity assessed via proteomics, and efficiency of transsulfuration pathway activity suggested beneficial effects on nutrient metabolism and metabolic-related stress. Maternal Met supply contributed to greater phosphatidylcholine synthesis in calf liver, with a role in very low density lipoprotein secretion as a mechanism to balance metabolic fates of fatty acids arising from the diet or adipose-depot lipolysis. -

Mouse Ssc4d Knockout Project (CRISPR/Cas9)
https://www.alphaknockout.com Mouse Ssc4d Knockout Project (CRISPR/Cas9) Objective: To create a Ssc4d knockout Mouse model (C57BL/6J) by CRISPR/Cas-mediated genome engineering. Strategy summary: The Ssc4d gene (NCBI Reference Sequence: NM_001160366 ; Ensembl: ENSMUSG00000029699 ) is located on Mouse chromosome 5. 11 exons are identified, with the ATG start codon in exon 2 and the TGA stop codon in exon 11 (Transcript: ENSMUST00000111152). Exon 2~11 will be selected as target site. Cas9 and gRNA will be co-injected into fertilized eggs for KO Mouse production. The pups will be genotyped by PCR followed by sequencing analysis. Note: Exon 2 starts from about 0.06% of the coding region. Exon 2~11 covers 100.0% of the coding region. The size of effective KO region: ~9451 bp. The KO region does not have any other known gene. Page 1 of 9 https://www.alphaknockout.com Overview of the Targeting Strategy Wildtype allele 5' gRNA region gRNA region 3' 1 2 3 4 7 8 9 10 11 Legends Exon of mouse Ssc4d Knockout region Page 2 of 9 https://www.alphaknockout.com Overview of the Dot Plot (up) Window size: 15 bp Forward Reverse Complement Sequence 12 Note: The 2000 bp section upstream of start codon is aligned with itself to determine if there are tandem repeats. Tandem repeats are found in the dot plot matrix. The gRNA site is selected outside of these tandem repeats. Overview of the Dot Plot (down) Window size: 15 bp Forward Reverse Complement Sequence 12 Note: The 2000 bp section downstream of stop codon is aligned with itself to determine if there are tandem repeats. -

Supplementary Information for Table of Contents
Supplementary Information for Tracing the origin of a new organ by inferring the genetic basis of rumen evolution Table of contents Supplementary Notes Supplementary Figures Supplementary Tables References 1 1 Supplementary Notes 2 Part 1. Phylogeny relationship and sample collection 3 1.1 Construction of the phylogenetic tree 4 To present the evolutionary panorama of the multi-chambered stomach evolution in 5 Cetartiodactyla, we constructed a phylogenetic tree using four-fold degenerate sites 6 using single-copy orthologous genes from nine species (human, horse, camel, pig, 7 hippo, killer whale, lesser mouse-deer, roe deer, and sheep) as representatives of the 8 major taxonomy. A final maximum likelihood tree was generated using IQ-TREE 9 multicore (version 1.6.6.a)1 with the parameters “-bb 1000 -m TEST -o Human” (Fig. 10 S17). Note that one of the families in the Suina, the Tayassuidae, has one stomach 11 with three chambers. Although no genomes of species in the Tayassuidae are 12 available, we chose the peccary as the representative of Tayassuidae and present the 13 position of the peccary according to the results of mitochondrial genomes2. The 14 newick format of tree result is as follows: (((((LesserMouse- 15 deer:0.1383065387,(Roedeer:0.0549981637,Sheep:0.0511103239)100:0.0574393138) 16 100:0.0488086408,(Killerwhale:0.0631824323,Hippo:0.0758665489)100:0.00822769 17 45)100:0.0179186732,Pig:0.1394609521)100:0.0101113682,Camel:0.1196798498)10 18 0:0.0411710418,Horse:0.1192130573,Human:0.1795944986); 19 1.2 RNA-seq analysis 20 1.2.1 Collection and sources of samples for RNA-seq 21 We collected and sequenced a total of 323 tissue samples from 11 Texel ♂ × Kazakh 22 ♀ hybrid sheep (Ovis aries) in Yili city (Xinjiang, China) (210 sequenced and 123 23 used in another paper at our laboratory)3. -

The Pdx1 Bound Swi/Snf Chromatin Remodeling Complex Regulates Pancreatic Progenitor Cell Proliferation and Mature Islet Β Cell
Page 1 of 125 Diabetes The Pdx1 bound Swi/Snf chromatin remodeling complex regulates pancreatic progenitor cell proliferation and mature islet β cell function Jason M. Spaeth1,2, Jin-Hua Liu1, Daniel Peters3, Min Guo1, Anna B. Osipovich1, Fardin Mohammadi3, Nilotpal Roy4, Anil Bhushan4, Mark A. Magnuson1, Matthias Hebrok4, Christopher V. E. Wright3, Roland Stein1,5 1 Department of Molecular Physiology and Biophysics, Vanderbilt University, Nashville, TN 2 Present address: Department of Pediatrics, Indiana University School of Medicine, Indianapolis, IN 3 Department of Cell and Developmental Biology, Vanderbilt University, Nashville, TN 4 Diabetes Center, Department of Medicine, UCSF, San Francisco, California 5 Corresponding author: [email protected]; (615)322-7026 1 Diabetes Publish Ahead of Print, published online June 14, 2019 Diabetes Page 2 of 125 Abstract Transcription factors positively and/or negatively impact gene expression by recruiting coregulatory factors, which interact through protein-protein binding. Here we demonstrate that mouse pancreas size and islet β cell function are controlled by the ATP-dependent Swi/Snf chromatin remodeling coregulatory complex that physically associates with Pdx1, a diabetes- linked transcription factor essential to pancreatic morphogenesis and adult islet-cell function and maintenance. Early embryonic deletion of just the Swi/Snf Brg1 ATPase subunit reduced multipotent pancreatic progenitor cell proliferation and resulted in pancreas hypoplasia. In contrast, removal of both Swi/Snf ATPase subunits, Brg1 and Brm, was necessary to compromise adult islet β cell activity, which included whole animal glucose intolerance, hyperglycemia and impaired insulin secretion. Notably, lineage-tracing analysis revealed Swi/Snf-deficient β cells lost the ability to produce the mRNAs for insulin and other key metabolic genes without effecting the expression of many essential islet-enriched transcription factors. -
Primepcr™Assay Validation Report
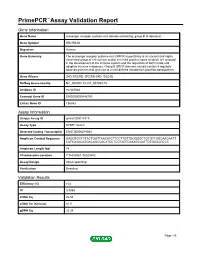
PrimePCR™Assay Validation Report Gene Information Gene Name scavenger receptor cysteine rich domain containing, group B (4 domains) Gene Symbol SRCRB4D Organism Human Gene Summary The scavenger receptor cysteine-rich (SRCR) superfamily is an ancient and highly conserved group of cell surface and/or secreted proteins some of which are involved in the development of the immune system and the regulation of both innate and adaptive immune responses. Group B SRCR domains usually contain 8 regularly spaced cysteines that give rise to a well-defined intradomain disulfide-bond pattern. Gene Aliases S4D-SRCRB, SRCRB-S4D, SSC4D RefSeq Accession No. NC_000007.13, NT_007933.15 UniGene ID Hs.567684 Ensembl Gene ID ENSG00000146700 Entrez Gene ID 136853 Assay Information Unique Assay ID qHsaCID0011418 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000275560 Amplicon Context Sequence GAGGTGCTCTACTGGTTAACATCTTCCTTGTTGGGGGCTGCGTTGGCAAGAATT CATCACACAGGACAGCCACATCCTCGTAGTGAAAGCAATTGTGGACGCCC Amplicon Length (bp) 74 Chromosome Location 7:76028061-76029652 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 103 R2 0.9988 cDNA Cq 26.58 cDNA Tm (Celsius) 81.5 gDNA Cq 33.38 Page 1/5 PrimePCR™Assay Validation Report Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report SRCRB4D, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard -

Supplemental Material
Supplemental Material The Antitumoral Effect of the S- Adenosylhomocysteine Hydrolase Inhibitor, 3-Deazaneplanocin A, is Independent of EZH2 but is Correlated with EGFR Downregulation in Chondrosarcomas Juliette Aury-Landasa Nicolas Girarda Eva Lhuissiera Drifa Adouanea Raphael Delépéeb Karim Boumedienea Catherine Baugéa aBioConnecT EA7451, Normandie Université, UNICAEN, Caen, France, bPRISMM, SF4206 ICORE, Normandie Université, UNICAEN, Comprehensive Cancer Center F. Baclesse, Caen, France Legends of supplementary figures Suppl figure 1: Pretreatment with DZNep reduced CS implantation and growth in xenograft mice model CH2879 cells were treated with DZNep (1 µM) for 5 days before subcutaneous grafting in nude mice. (A) Tumors were measured regularly by a caliper and tumoral volume calculated. (B) 46 days after cell injection, tumors were weighted. Data are expressed as means + SEM. A total of ten mice were used (five pre-treated and five controls). Suppl figure 2: Expression of mRNA of identified genes in DZNep-treated CS and chondrocytes EGFR, MAD1L1 mRNA expression was analyzed by RT-PCR from chondrosarcomas and chondrocytes treated with 1 µM DZNep for 24 h. Data are expressed as means + SEM (N=3-5). *: p-value < 0.05. 1 A 700 B 600 ) pretreatment 3 500 500 * 400 400 * volume (mm volume 300 300 * Tumor 200 200 C 100 100 (mg) weight Tumor 0 0 DZNep 25 30 35 40 45 50 Control Pretreated Time after implantation (days) Suppl figure 1 A 1.6 1.4 EGFR 1.2 Control 1 * DZNep 0.8 * (relative unit) (relative 0.6 * level ** 0.4 mRNA 0.2 0 CH2879 SW1353 JJ012 FS090 HAC 1.41.4 MAD1L1 1.21.2 1 * 0.80.8 (relative unit) (relative 0.6 * level *** 0.40.4 *** mRNA 0.20.2 0 CH2879 SW1353 JJ012 FS090 HAC Suppl figure 2 Table S1: Clinical characteristics of primary chondrosarcomas. -

Escherichia Coli O157:H7 and Lactoferrin Modulate Immunity in Cattle
Escherichia coli O157:H7 and lactoferrin modulate immunity in cattle Evelien Kieckens Laboratory of Immunology Department of Virology, Parasitology and Immunology Thesis is submitted in fulfillment of the requirement for the degree of Doctor in Veterinary Sciences (PhD), Faculty of Veterinary Medicine, Ghent University, 2016 "A model is a lie that helps you see the truth." Howard Skipper The author and promoters give the permission to consult and to copy parts of this work for personal use only. Any other use is subject to the Copyright Laws. Permission to reproduce any material in this work should be obtained from the author. Promoters: Prof. Dr. Eric Cox Department of Virology, Parasitology and Immunology, Faculty of Veterinary Medicine, Ghent University Prof. Dr. Daisy Vanrompay Department of Animal Production, Faculty of Bioscience Engineering, Ghent University Prof. Dr. Lieven De Zutter Department of Veterinary Public Health and Food Safety, Faculty of Veterinary Medicine, Ghent University Members of the Examination Committee: Prof. Dr. Piet Deprez (Chairman of the Examination Committee) Department of Internal medicine and clinical biology of large animals Faculty of Veterinary Medicine, Ghent University, Belgium Prof. Dr. Herman Favoreel (Secretary of the Examination Committee) Department of Virology, Parasitology and Immunology Faculty of Veterinary Medicine, Ghent University, Belgium Dr. ir. Koen De Reu Instituut voor Landbouw en Visserijonderzoek (ILVO), Belgium Prof. Dr. ir. Stefaan De Smet Department of Animal production Faculty of Faculty of Bioscience Engineering, Ghent University, Belgium Prof. Dr. Christian Menge Friedrich-Loeffler-Institut Institute of Molecular Pathogenesis, Jena, Germany Table of Contents Table of Contents List of abbreviations ................................................................................................................ 9 Chapter I: Literature review .................................................................................................. -

Comparison of Selection Signatures Between Korean Native and Commercial Chickens Using 600K SNP Array Data
G C A T T A C G G C A T genes Article Comparison of Selection Signatures between Korean Native and Commercial Chickens Using 600K SNP Array Data Sunghyun Cho 1, Prabuddha Manjula 1, Minjun Kim 1,2, Eunjin Cho 1, Dooho Lee 1, Seung Hwan Lee 1,2, Jun Heon Lee 1,2 and Dongwon Seo 1,2,* 1 Division of Animal and Dairy Science, Chungnam National University, Daejeon 34134, Korea; [email protected] (S.C.); [email protected] (P.M.); [email protected] (M.K.); fl[email protected] (E.C.); [email protected] (D.L.); [email protected] (S.H.L.); [email protected] (J.H.L.) 2 Bio-AI Convergence Research Center, Chungnam National University, Daejeon 34134, Korea * Correspondence: [email protected]; Tel.: +82-42-821-5878 Abstract: Korean native chickens (KNCs) comprise an indigenous chicken breed of South Korea that was restored through a government project in the 1990s. The KNC population has not been developed well and has mostly been used to maintain purebred populations in the government research institution. We investigated the genetic features of the KNC population in a selection signal study for the efficient improvement of this breed. We used 600K single nucleotide polymorphism data sampled from 191 KNCs (NG, 38; NL, 29; NR, 52; NW, 39; and NY, 33) and 54 commercial chickens (Hy-line Brown, 10; Lohmann Brown, 10; Arbor Acres, 10; Cobb, 12; and Ross, 12). Haplotype phasing was performed using EAGLE software as the initial step for the primary data analysis. Pre-processed data were analyzed to detect selection signals using the ‘rehh’ package in R software. -
UC Irvine UC Irvine Electronic Theses and Dissertations
UC Irvine UC Irvine Electronic Theses and Dissertations Title Investigating developmental and functional deficits in neurodegenerative disease using transcriptomic analyses Permalink https://escholarship.org/uc/item/7wp2w13r Author Lim, Ryan Gar-Lok Publication Date 2016 Peer reviewed|Thesis/dissertation eScholarship.org Powered by the California Digital Library University of California UNIVERSITY OF CALIFORNIA, IRVINE Investigating developmental and functional deficits in neurodegenerative disease using transcriptomic analyses DISSERTATION submitted in partial satisfaction of the requirements for the degree of DOCTOR OF PHILOSOPHY in Biomedical Sciences by Ryan Gar-Lok Lim Dissertation Committee: Professor Leslie M. Thompson, Chair Assistant Professor Dritan Agalliu Professor Peter Donovan Professor Suzanne Sandmeyer 2016 Introduction, Figure 1.1 © 2014 Macmillan Publishers Limited. Appendix 1 © 2016 Elsevier Ltd. All other materials © 2016 Ryan Gar-Lok Lim DEDICATION This dissertation is dedicated to my parents, sister, and my wife. I love you all very much and could not have accomplished any of this without your love and support. Please take the time to reflect back on all of the moments we’ve shared, and know, that it is because of those moments I have been able to succeed. This accomplishment is as much yours as it is mine. ii TABLE OF CONTENTS Page LIST OF FIGURES vi LIST OF TABLES ix ACKNOWLEDGMENTS x CURRICULUM VITAE xiii ABSTRACT OF THE DISSERTATION xv Introduction Huntington’s disease, the neurovascular unit and the blood-brain -

Genomic Scans for Selective Sweeps Through Haplotype Homozygosity
www.nature.com/scientificreports OPEN Genomic scans for selective sweeps through haplotype homozygosity and allelic fxation in 14 indigenous sheep breeds from Middle East and South Asia Sirous Eydivandi1,2*, Mahmoud Amiri Roudbar3, Mohammad Osman Karimi4 & Goutam Sahana2 The performance and productivity of livestock have consistently improved by natural and artifcial selection over the centuries. Both these selections are expected to leave patterns on the genome and lead to changes in allele frequencies, but natural selection has played the major role among indigenous populations. Detecting selective sweeps in livestock may assist in understanding the processes involved in domestication, genome evolution and discovery of genomic regions associated with economically important traits. We investigated population genetic diversity and selection signals in this study using SNP genotype data of 14 indigenous sheep breeds from Middle East and South Asia, including six breeds from Iran, namely Iranian Balochi, Afshari, Moghani, Qezel, Zel, and Lori- Bakhtiari, three breeds from Afghanistan, namely Afghan Balochi, Arabi, and Gadik, three breeds from India, namely Indian Garole, Changthangi, and Deccani, and two breeds from Bangladesh, namely Bangladeshi Garole and Bangladesh East. The SNP genotype data were generated by the Illumina OvineSNP50 Genotyping BeadChip array. To detect genetic diversity and population structure, we used principal component analysis (PCA), admixture, phylogenetic analyses, and Runs of homozygosity. We applied four complementary statistical tests, FST (fxation index), xp-EHH (cross-population extended haplotype homozygosity), Rsb (extended haplotype homozygosity between-populations), and FLK (the extension of the Lewontin and Krakauer) to detect selective sweeps. Our results not only confrm the previous studies but also provide a suite of novel candidate genes involved in diferent traits in sheep.