The Functional Analysis of Granulins
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Genetic Analysis of Retinopathy in Type 1 Diabetes
Genetic Analysis of Retinopathy in Type 1 Diabetes by Sayed Mohsen Hosseini A thesis submitted in conformity with the requirements for the degree of Doctor of Philosophy Institute of Medical Science University of Toronto © Copyright by S. Mohsen Hosseini 2014 Genetic Analysis of Retinopathy in Type 1 Diabetes Sayed Mohsen Hosseini Doctor of Philosophy Institute of Medical Science University of Toronto 2014 Abstract Diabetic retinopathy (DR) is a leading cause of blindness worldwide. Several lines of evidence suggest a genetic contribution to the risk of DR; however, no genetic variant has shown convincing association with DR in genome-wide association studies (GWAS). To identify common polymorphisms associated with DR, meta-GWAS were performed in three type 1 diabetes cohorts of White subjects: Diabetes Complications and Control Trial (DCCT, n=1304), Wisconsin Epidemiologic Study of Diabetic Retinopathy (WESDR, n=603) and Renin-Angiotensin System Study (RASS, n=239). Severe (SDR) and mild (MDR) retinopathy outcomes were defined based on repeated fundus photographs in each study graded for retinopathy severity on the Early Treatment Diabetic Retinopathy Study (ETDRS) scale. Multivariable models accounted for glycemia (measured by A1C), diabetes duration and other relevant covariates in the association analyses of additive genotypes with SDR and MDR. Fixed-effects meta- analysis was used to combine the results of GWAS performed separately in WESDR, ii RASS and subgroups of DCCT, defined by cohort and treatment group. Top association signals were prioritized for replication, based on previous supporting knowledge from the literature, followed by replication in three independent white T1D studies: Genesis-GeneDiab (n=502), Steno (n=936) and FinnDiane (n=2194). -

Variation in Genomic Landscape of Clear Cell Renal Cell Carcinoma
ARTICLE Received 30 May 2014 | Accepted 3 Sep 2014 | Published 29 Oct 2014 DOI: 10.1038/ncomms6135 Variation in genomic landscape of clear cell renal cell carcinoma across Europe Ghislaine Scelo1,*, Yasser Riazalhosseini2,3,*, Liliana Greger4,*, Louis Letourneau3,*, Mar Gonza`lez-Porta4,*, Magdalena B. Wozniak1, Mathieu Bourgey3, Patricia Harnden5, Lars Egevad6, Sharon M. Jackson5, Mehran Karimzadeh2,3, Madeleine Arseneault2,3, Pierre Lepage3, Alexandre How-Kit7, Antoine Daunay7, Victor Renault7,He´le`ne Blanche´7, Emmanuel Tubacher7, Jeremy Sehmoun7, Juris Viksna8, Edgars Celms8, Martins Opmanis8, Andris Zarins8, Naveen S. Vasudev5, Morag Seywright9, Behnoush Abedi-Ardekani1, Christine Carreira1, Peter J. Selby5, Jon J. Cartledge10, Graham Byrnes1, Jiri Zavadil1, Jing Su4, Ivana Holcatova11, Antonin Brisuda12, David Zaridze13, Anush Moukeria13, Lenka Foretova14, Marie Navratilova14, Dana Mates15, Viorel Jinga16, Artem Artemov17, Artem Nedoluzhko18, Alexander Mazur17, Sergey Rastorguev18, Eugenia Boulygina18, Simon Heath19, Marta Gut19, Marie-Therese Bihoreau20, Doris Lechner20, Mario Foglio20, Ivo G. Gut19, Konstantin Skryabin17,18, Egor Prokhortchouk17,18, Anne Cambon-Thomsen21, Johan Rung4, Guillaume Bourque2,3, Paul Brennan1,Jo¨rg Tost20, Rosamonde E. Banks5,AlvisBrazma4 & G. Mark Lathrop2,3,7,20,w The incidence of renal cell carcinoma (RCC) is increasing worldwide, and its prevalence is particularly high in some parts of Central Europe. Here we undertake whole-genome and transcriptome sequencing of clear cell RCC (ccRCC), the most common form of the disease, in patients from four different European countries with contrasting disease incidence to explore the underlying genomic architecture of RCC. Our findings support previous reports on frequent aberrations in the epigenetic machinery and PI3K/mTOR signalling, and uncover novel pathways and genes affected by recurrent mutations and abnormal transcriptome patterns including focal adhesion, components of extracellular matrix (ECM) and genes encoding FAT cadherins. -

Supplementary Data
Supplemental figures Supplemental figure 1: Tumor sample selection. A total of 98 thymic tumor specimens were stored in Memorial Sloan-Kettering Cancer Center tumor banks during the study period. 64 cases corresponded to previously untreated tumors, which were resected upfront after diagnosis. Adjuvant treatment was delivered in 7 patients (radiotherapy in 4 cases, cyclophosphamide- doxorubicin-vincristine (CAV) chemotherapy in 3 cases). 34 tumors were resected after induction treatment, consisting of chemotherapy in 16 patients (cyclophosphamide-doxorubicin- cisplatin (CAP) in 11 cases, cisplatin-etoposide (PE) in 3 cases, cisplatin-etoposide-ifosfamide (VIP) in 1 case, and cisplatin-docetaxel in 1 case), in radiotherapy (45 Gy) in 1 patient, and in sequential chemoradiation (CAP followed by a 45 Gy-radiotherapy) in 1 patient. Among these 34 patients, 6 received adjuvant radiotherapy. 1 Supplemental Figure 2: Amino acid alignments of KIT H697 in the human protein and related orthologs, using (A) the Homologene database (exons 14 and 15), and (B) the UCSC Genome Browser database (exon 14). Residue H697 is highlighted with red boxes. Both alignments indicate that residue H697 is highly conserved. 2 Supplemental Figure 3: Direct comparison of the genomic profiles of thymic squamous cell carcinomas (n=7) and lung primary squamous cell carcinomas (n=6). (A) Unsupervised clustering analysis. Gains are indicated in red, and losses in green, by genomic position along the 22 chromosomes. (B) Genomic profiles and recurrent copy number alterations in thymic carcinomas and lung squamous cell carcinomas. Gains are indicated in red, and losses in blue. 3 Supplemental Methods Mutational profiling The exonic regions of interest (NCBI Human Genome Build 36.1) were broken into amplicons of 500 bp or less, and specific primers were designed using Primer 3 (on the World Wide Web for general users and for biologist programmers (see Supplemental Table 2) [1]. -
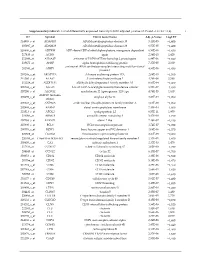
Supplementary Table S1. List of Differentially Expressed
Supplementary table S1. List of differentially expressed transcripts (FDR adjusted p‐value < 0.05 and −1.4 ≤ FC ≥1.4). 1 ID Symbol Entrez Gene Name Adj. p‐Value Log2 FC 214895_s_at ADAM10 ADAM metallopeptidase domain 10 3,11E‐05 −1,400 205997_at ADAM28 ADAM metallopeptidase domain 28 6,57E‐05 −1,400 220606_s_at ADPRM ADP‐ribose/CDP‐alcohol diphosphatase, manganese dependent 6,50E‐06 −1,430 217410_at AGRN agrin 2,34E‐10 1,420 212980_at AHSA2P activator of HSP90 ATPase homolog 2, pseudogene 6,44E‐06 −1,920 219672_at AHSP alpha hemoglobin stabilizing protein 7,27E‐05 2,330 aminoacyl tRNA synthetase complex interacting multifunctional 202541_at AIMP1 4,91E‐06 −1,830 protein 1 210269_s_at AKAP17A A‐kinase anchoring protein 17A 2,64E‐10 −1,560 211560_s_at ALAS2 5ʹ‐aminolevulinate synthase 2 4,28E‐06 3,560 212224_at ALDH1A1 aldehyde dehydrogenase 1 family member A1 8,93E‐04 −1,400 205583_s_at ALG13 ALG13 UDP‐N‐acetylglucosaminyltransferase subunit 9,50E‐07 −1,430 207206_s_at ALOX12 arachidonate 12‐lipoxygenase, 12S type 4,76E‐05 1,630 AMY1C (includes 208498_s_at amylase alpha 1C 3,83E‐05 −1,700 others) 201043_s_at ANP32A acidic nuclear phosphoprotein 32 family member A 5,61E‐09 −1,760 202888_s_at ANPEP alanyl aminopeptidase, membrane 7,40E‐04 −1,600 221013_s_at APOL2 apolipoprotein L2 6,57E‐11 1,600 219094_at ARMC8 armadillo repeat containing 8 3,47E‐08 −1,710 207798_s_at ATXN2L ataxin 2 like 2,16E‐07 −1,410 215990_s_at BCL6 BCL6 transcription repressor 1,74E‐07 −1,700 200776_s_at BZW1 basic leucine zipper and W2 domains 1 1,09E‐06 −1,570 222309_at -

4766.Full.Pdf
Identification of Novel Th2-Associated Genes in T Memory Responses to Allergens Anthony Bosco, Kathy L. McKenna, Catherine J. Devitt, Martin J. Firth, Peter D. Sly and Patrick G. Holt This information is current as of October 1, 2021. J Immunol 2006; 176:4766-4777; ; doi: 10.4049/jimmunol.176.8.4766 http://www.jimmunol.org/content/176/8/4766 Downloaded from References This article cites 119 articles, 43 of which you can access for free at: http://www.jimmunol.org/content/176/8/4766.full#ref-list-1 Why The JI? Submit online. http://www.jimmunol.org/ • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average by guest on October 1, 2021 Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2006 by The American Association of Immunologists All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Identification of Novel Th2-Associated Genes in T Memory Responses to Allergens1 Anthony Bosco, Kathy L. McKenna, Catherine J. Devitt, Martin J. Firth, Peter D. Sly, and Patrick G. Holt2 Atopic diseases are associated with hyperexpression of Th2 cytokines by allergen-specific T memory cells. -

Functional Genomic Annotation of Genetic Risk Loci Highlights Inflammation and Epithelial Biology Networks in CKD
BASIC RESEARCH www.jasn.org Functional Genomic Annotation of Genetic Risk Loci Highlights Inflammation and Epithelial Biology Networks in CKD Nora Ledo, Yi-An Ko, Ae-Seo Deok Park, Hyun-Mi Kang, Sang-Youb Han, Peter Choi, and Katalin Susztak Renal Electrolyte and Hypertension Division, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania ABSTRACT Genome-wide association studies (GWASs) have identified multiple loci associated with the risk of CKD. Almost all risk variants are localized to the noncoding region of the genome; therefore, the role of these variants in CKD development is largely unknown. We hypothesized that polymorphisms alter transcription factor binding, thereby influencing the expression of nearby genes. Here, we examined the regulation of transcripts in the vicinity of CKD-associated polymorphisms in control and diseased human kidney samples and used systems biology approaches to identify potentially causal genes for prioritization. We interro- gated the expression and regulation of 226 transcripts in the vicinity of 44 single nucleotide polymorphisms using RNA sequencing and gene expression arrays from 95 microdissected control and diseased tubule samples and 51 glomerular samples. Gene expression analysis from 41 tubule samples served for external validation. 92 transcripts in the tubule compartment and 34 transcripts in glomeruli showed statistically significant correlation with eGFR. Many novel genes, including ACSM2A/2B, FAM47E, and PLXDC1, were identified. We observed that the expression of multiple genes in the vicinity of any single CKD risk allele correlated with renal function, potentially indicating that genetic variants influence multiple transcripts. Network analysis of GFR-correlating transcripts highlighted two major clusters; a positive correlation with epithelial and vascular functions and an inverse correlation with inflammatory gene cluster. -

Quantitative Trait Loci Mapping of Macrophage Atherogenic Phenotypes
QUANTITATIVE TRAIT LOCI MAPPING OF MACROPHAGE ATHEROGENIC PHENOTYPES BRIAN RITCHEY Bachelor of Science Biochemistry John Carroll University May 2009 submitted in partial fulfillment of requirements for the degree DOCTOR OF PHILOSOPHY IN CLINICAL AND BIOANALYTICAL CHEMISTRY at the CLEVELAND STATE UNIVERSITY December 2017 We hereby approve this thesis/dissertation for Brian Ritchey Candidate for the Doctor of Philosophy in Clinical-Bioanalytical Chemistry degree for the Department of Chemistry and the CLEVELAND STATE UNIVERSITY College of Graduate Studies by ______________________________ Date: _________ Dissertation Chairperson, Johnathan D. Smith, PhD Department of Cellular and Molecular Medicine, Cleveland Clinic ______________________________ Date: _________ Dissertation Committee member, David J. Anderson, PhD Department of Chemistry, Cleveland State University ______________________________ Date: _________ Dissertation Committee member, Baochuan Guo, PhD Department of Chemistry, Cleveland State University ______________________________ Date: _________ Dissertation Committee member, Stanley L. Hazen, MD PhD Department of Cellular and Molecular Medicine, Cleveland Clinic ______________________________ Date: _________ Dissertation Committee member, Renliang Zhang, MD PhD Department of Cellular and Molecular Medicine, Cleveland Clinic ______________________________ Date: _________ Dissertation Committee member, Aimin Zhou, PhD Department of Chemistry, Cleveland State University Date of Defense: October 23, 2017 DEDICATION I dedicate this work to my entire family. In particular, my brother Greg Ritchey, and most especially my father Dr. Michael Ritchey, without whose support none of this work would be possible. I am forever grateful to you for your devotion to me and our family. You are an eternal inspiration that will fuel me for the remainder of my life. I am extraordinarily lucky to have grown up in the family I did, which I will never forget. -

Multiscale Genomic Analysis of The
University of Tennessee Health Science Center UTHSC Digital Commons Theses and Dissertations (ETD) College of Graduate Health Sciences 5-2009 Multiscale Genomic Analysis of the Corticolimbic System: Uncovering the Molecular and Anatomic Substrates of Anxiety-Related Behavior Khyobeni Mozhui University of Tennessee Health Science Center Follow this and additional works at: https://dc.uthsc.edu/dissertations Part of the Mental and Social Health Commons, Nervous System Commons, and the Neurosciences Commons Recommended Citation Mozhui, Khyobeni , "Multiscale Genomic Analysis of the Corticolimbic System: Uncovering the Molecular and Anatomic Substrates of Anxiety-Related Behavior" (2009). Theses and Dissertations (ETD). Paper 180. http://dx.doi.org/10.21007/etd.cghs.2009.0219. This Dissertation is brought to you for free and open access by the College of Graduate Health Sciences at UTHSC Digital Commons. It has been accepted for inclusion in Theses and Dissertations (ETD) by an authorized administrator of UTHSC Digital Commons. For more information, please contact [email protected]. Multiscale Genomic Analysis of the Corticolimbic System: Uncovering the Molecular and Anatomic Substrates of Anxiety-Related Behavior Document Type Dissertation Degree Name Doctor of Philosophy (PhD) Program Anatomy and Neurobiology Research Advisor Robert W. Williams, Ph.D. Committee John D. Boughter, Ph.D. Eldon E. Geisert, Ph.D. Kristin M. Hamre, Ph.D. Jeffery D. Steketee, Ph.D. DOI 10.21007/etd.cghs.2009.0219 This dissertation is available at UTHSC Digital -

Tepzz 8Z6z54a T
(19) TZZ ZZ_T (11) EP 2 806 054 A1 (12) EUROPEAN PATENT APPLICATION (43) Date of publication: (51) Int Cl.: 26.11.2014 Bulletin 2014/48 C40B 40/06 (2006.01) C12Q 1/68 (2006.01) C40B 30/04 (2006.01) C07H 21/00 (2006.01) (21) Application number: 14175049.7 (22) Date of filing: 28.05.2009 (84) Designated Contracting States: (74) Representative: Irvine, Jonquil Claire AT BE BG CH CY CZ DE DK EE ES FI FR GB GR HGF Limited HR HU IE IS IT LI LT LU LV MC MK MT NL NO PL 140 London Wall PT RO SE SI SK TR London EC2Y 5DN (GB) (30) Priority: 28.05.2008 US 56827 P Remarks: •Thecomplete document including Reference Tables (62) Document number(s) of the earlier application(s) in and the Sequence Listing can be downloaded from accordance with Art. 76 EPC: the EPO website 09753364.0 / 2 291 553 •This application was filed on 30-06-2014 as a divisional application to the application mentioned (71) Applicant: Genomedx Biosciences Inc. under INID code 62. Vancouver, British Columbia V6J 1J8 (CA) •Claims filed after the date of filing of the application/ after the date of receipt of the divisional application (72) Inventor: Davicioni, Elai R.68(4) EPC). Vancouver British Columbia V6J 1J8 (CA) (54) Systems and methods for expression- based discrimination of distinct clinical disease states in prostate cancer (57) A system for expression-based discrimination of distinct clinical disease states in prostate cancer is provided that is based on the identification of sets of gene transcripts, which are characterized in that changes in expression of each gene transcript within a set of gene transcripts can be correlated with recurrent or non- recur- rent prostate cancer. -

View Correction with a P Value,0.05
BASIC RESEARCH www.jasn.org Functional Genomic Annotation of Genetic Risk Loci Highlights Inflammation and Epithelial Biology Networks in CKD Nora Ledo, Yi-An Ko, Ae-Seo Deok Park, Hyun-Mi Kang, Sang-Youb Han, Peter Choi, and Katalin Susztak Renal Electrolyte and Hypertension Division, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania ABSTRACT Genome-wide association studies (GWASs) have identified multiple loci associated with the risk of CKD. Almost all risk variants are localized to the noncoding region of the genome; therefore, the role of these variants in CKD development is largely unknown. We hypothesized that polymorphisms alter transcription factor binding, thereby influencing the expression of nearby genes. Here, we examined the regulation of transcripts in the vicinity of CKD-associated polymorphisms in control and diseased human kidney samples and used systems biology approaches to identify potentially causal genes for prioritization. We interro- gated the expression and regulation of 226 transcripts in the vicinity of 44 single nucleotide polymorphisms using RNA sequencing and gene expression arrays from 95 microdissected control and diseased tubule samples and 51 glomerular samples. Gene expression analysis from 41 tubule samples served for external validation. 92 transcripts in the tubule compartment and 34 transcripts in glomeruli showed statistically significant correlation with eGFR. Many novel genes, including ACSM2A/2B, FAM47E, and PLXDC1, were identified. We observed that the expression of multiple genes in the vicinity of any single CKD risk allele correlated with renal function, potentially indicating that genetic variants influence multiple transcripts. Network analysis of GFR-correlating transcripts highlighted two major clusters; a positive correlation with epithelial and vascular functions and an inverse correlation with inflammatory gene cluster. -

Novel Estrogen Receptor-A Binding Sites and Estradiol Target Genes Identified by Chromatin Immunoprecipitation Cloning in Breast Cancer
Research Article Novel Estrogen Receptor-A Binding Sites and Estradiol Target Genes Identified by Chromatin Immunoprecipitation Cloning in Breast Cancer Zhihong Lin,1 Scott Reierstad,1 Chiang-Ching Huang,2 and Serdar E. Bulun1 Departments of 1Obstetrics and Gynecology and 2Preventive Medicine, Feinberg School of Medicine, Northwestern University, Chicago, Illinois Abstract by association with other transcription factors on promoter targets A A (1, 4, 5). Until recently, however, little had been known about Estrogen receptor- (ER ) and its ligand estradiol play a critical roles in breast cancer growth and are important the distribution of ER -binding sites within the genome and the therapeutic targets for this disease. Using chromatin immu- identity of genes regulated by these cis-acting elements. A Two recent pioneering publications by Carroll et al. (6, 7) noprecipitation (ChIP)-on-chip, ligand-bound ER was re- a cently found to function as a master transcriptional regulator revolutionized our understanding of ER action. Using chromatin cis immunoprecipitation (ChIP)-on-chip, this group mapped a large via binding to many -acting sites genome-wide. Here, we a used an alternative technology (ChIP cloning) and identified number of ER -binding sites on a chromosome and genome-wide scale, identifying novel cis-regulatory sites and target genes in 94 ERA target loci in breast cancer cells. The ERA-binding sites contained both classic estrogen response elements and MCF-7 breast cancer cells (6, 7). The majority of these binding sites nonclassic binding sequences, showed specific transcriptional were distant from the transcription start sites of regulated genes activity in reporter gene assay, and interacted with the key (6, 7). -

MAFB Determines Human Macrophage Anti-Inflammatory
MAFB Determines Human Macrophage Anti-Inflammatory Polarization: Relevance for the Pathogenic Mechanisms Operating in Multicentric Carpotarsal Osteolysis This information is current as of September 25, 2021. Víctor D. Cuevas, Laura Anta, Rafael Samaniego, Emmanuel Orta-Zavalza, Juan Vladimir de la Rosa, Geneviève Baujat, Ángeles Domínguez-Soto, Paloma Sánchez-Mateos, María M. Escribese, Antonio Castrillo, Valérie Cormier-Daire, Miguel A. Vega and Ángel L. Corbí Downloaded from J Immunol published online 16 January 2017 http://www.jimmunol.org/content/early/2017/01/15/jimmun ol.1601667 http://www.jimmunol.org/ Supplementary http://www.jimmunol.org/content/suppl/2017/01/15/jimmunol.160166 Material 7.DCSupplemental Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision by guest on September 25, 2021 • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2017 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. Published January 16, 2017, doi:10.4049/jimmunol.1601667 The Journal of Immunology MAFB Determines Human Macrophage Anti-Inflammatory Polarization: Relevance for the Pathogenic Mechanisms Operating in Multicentric Carpotarsal Osteolysis Vı´ctor D.