Primepcr™Assay Validation Report
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Downloaded the Normalized Gene Expression Profile from the GEO (Package Geoquery (5))
1 SUPPLEMENTARY MATERIALS 2 Supplementary Methods 3 Study design 4 The workflow to identify and validate the TME risk score and TME subtypes in gastric cancer is 5 depicted in Fig. S1. We first estimated the absolute abundance levels of the major stromal and 6 immune cell types in the TME using bulk gene expression data, and assessed the prognostic 7 effect of these cells in a discovery cohort. Next, we constructed a TME risk score and validated it 8 in two independent gene expression validation cohorts and three immunohistochemistry 9 validation cohorts. Finally, we stratified patients into four TME subtypes and examined their 10 genomic and molecular features and relation to established molecular subtypes. 11 Gene expression data 12 To explore the prognostic landscape of the TME, we used four gene expression profile (GEP) 13 datasets of resected gastric cancer patients with publicly available clinical information, namely, 14 ACRG (GSE62254) (1), GSE15459 (2), GSE84437 (3), and TCGA stomach adenocarcinoma 15 (STAD). Specifically, the raw microarray data in the ACRG cohort and GSE15459 were retrieved 16 from the Gene Expression Omnibus (GEO), and normalized by the RMA algorithm (package affy) 17 using custom chip definition files (Brainarray version 23 (4)) that convert Affymetrix probesets to 18 Entrez gene IDs. For GSE84437 dataset, which was measured by the Illumina platform, we 19 downloaded the normalized gene expression profile from the GEO (package GEOquery (5)). The 20 Illumina probes were also mapped to Entrez genes. For multiple probes mapping to the same 21 Entrez gene, we selected the one with the maximum mean expression level as the surrogate for 22 the Entrez gene using the function of collapseRows (6) (package WGCNA). -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

Human Lectins, Their Carbohydrate Affinities and Where to Find Them
biomolecules Review Human Lectins, Their Carbohydrate Affinities and Where to Review HumanFind Them Lectins, Their Carbohydrate Affinities and Where to FindCláudia ThemD. Raposo 1,*, André B. Canelas 2 and M. Teresa Barros 1 1, 2 1 Cláudia D. Raposo * , Andr1 é LAQVB. Canelas‐Requimte,and Department M. Teresa of Chemistry, Barros NOVA School of Science and Technology, Universidade NOVA de Lisboa, 2829‐516 Caparica, Portugal; [email protected] 12 GlanbiaLAQV-Requimte,‐AgriChemWhey, Department Lisheen of Chemistry, Mine, Killoran, NOVA Moyne, School E41 of ScienceR622 Co. and Tipperary, Technology, Ireland; canelas‐ [email protected] NOVA de Lisboa, 2829-516 Caparica, Portugal; [email protected] 2* Correspondence:Glanbia-AgriChemWhey, [email protected]; Lisheen Mine, Tel.: Killoran, +351‐212948550 Moyne, E41 R622 Tipperary, Ireland; [email protected] * Correspondence: [email protected]; Tel.: +351-212948550 Abstract: Lectins are a class of proteins responsible for several biological roles such as cell‐cell in‐ Abstract:teractions,Lectins signaling are pathways, a class of and proteins several responsible innate immune for several responses biological against roles pathogens. such as Since cell-cell lec‐ interactions,tins are able signalingto bind to pathways, carbohydrates, and several they can innate be a immuneviable target responses for targeted against drug pathogens. delivery Since sys‐ lectinstems. In are fact, able several to bind lectins to carbohydrates, were approved they by canFood be and a viable Drug targetAdministration for targeted for drugthat purpose. delivery systems.Information In fact, about several specific lectins carbohydrate were approved recognition by Food by andlectin Drug receptors Administration was gathered for that herein, purpose. plus Informationthe specific organs about specific where those carbohydrate lectins can recognition be found by within lectin the receptors human was body. -
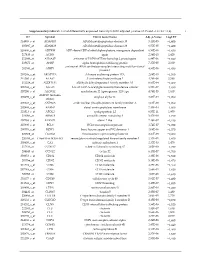
Supplementary Table S1. List of Differentially Expressed
Supplementary table S1. List of differentially expressed transcripts (FDR adjusted p‐value < 0.05 and −1.4 ≤ FC ≥1.4). 1 ID Symbol Entrez Gene Name Adj. p‐Value Log2 FC 214895_s_at ADAM10 ADAM metallopeptidase domain 10 3,11E‐05 −1,400 205997_at ADAM28 ADAM metallopeptidase domain 28 6,57E‐05 −1,400 220606_s_at ADPRM ADP‐ribose/CDP‐alcohol diphosphatase, manganese dependent 6,50E‐06 −1,430 217410_at AGRN agrin 2,34E‐10 1,420 212980_at AHSA2P activator of HSP90 ATPase homolog 2, pseudogene 6,44E‐06 −1,920 219672_at AHSP alpha hemoglobin stabilizing protein 7,27E‐05 2,330 aminoacyl tRNA synthetase complex interacting multifunctional 202541_at AIMP1 4,91E‐06 −1,830 protein 1 210269_s_at AKAP17A A‐kinase anchoring protein 17A 2,64E‐10 −1,560 211560_s_at ALAS2 5ʹ‐aminolevulinate synthase 2 4,28E‐06 3,560 212224_at ALDH1A1 aldehyde dehydrogenase 1 family member A1 8,93E‐04 −1,400 205583_s_at ALG13 ALG13 UDP‐N‐acetylglucosaminyltransferase subunit 9,50E‐07 −1,430 207206_s_at ALOX12 arachidonate 12‐lipoxygenase, 12S type 4,76E‐05 1,630 AMY1C (includes 208498_s_at amylase alpha 1C 3,83E‐05 −1,700 others) 201043_s_at ANP32A acidic nuclear phosphoprotein 32 family member A 5,61E‐09 −1,760 202888_s_at ANPEP alanyl aminopeptidase, membrane 7,40E‐04 −1,600 221013_s_at APOL2 apolipoprotein L2 6,57E‐11 1,600 219094_at ARMC8 armadillo repeat containing 8 3,47E‐08 −1,710 207798_s_at ATXN2L ataxin 2 like 2,16E‐07 −1,410 215990_s_at BCL6 BCL6 transcription repressor 1,74E‐07 −1,700 200776_s_at BZW1 basic leucine zipper and W2 domains 1 1,09E‐06 −1,570 222309_at -

The Centromeric Part of the Human NK Gene Complex: Linkage of LOX-1 and LY49L with the CD94/NKG2 Region
Genes and Immunity (2000) 1, 280–287 2000 Macmillan Publishers Ltd All rights reserved 1466-4879/00 $15.00 www.nature.com/gene The centromeric part of the human NK gene complex: linkage of LOX-1 and LY49L with the CD94/NKG2 region C Bull1, Y Sobanov2,BRo¨hrdanz1, J O’Brien1, H Lehrach1 and E Hofer2 1Max-Planck-Institute for Molecular Genetics, Ihnestrasse 73, D-14195 Berlin, Germany; 2Department of Vascular Biology and Thrombosis Research, Vienna International Research Cooperation Center, University of Vienna, Brunnerstrasse 59, A-1235 Vienna, Austria The natural killer (NK) gene complex is a genomic region containing lectin-type receptor genes. We have established a contig of PAC and BAC clones comprising about 1 Mb of the centromeric part of the NK gene complex. This region extends from the LOX-1 gene, which encodes a receptor for oxidized LDL and was found within 100 kb telomeric of the STS marker D12S77, contains the CD94 and NKG2 NK receptor genes and reaches beyond D12S852 on the proximal side. In this part we have mapped the human LY49L gene, a homologue of the rodent Ly49 genes, which encode important MHC class I receptors for the regulation of NK cell activity in rodents. The LY49L gene is localized 100 to 200 kb centromeric of the NKG2 gene cluster and 300 to 400 kb telomeric of the STS marker D12S841. Genomic sequencing of the complete gene including promoter and intron sequences confirmed that the structure is similar to the mouse Ly49 genes. Screening of several cDNA libraries did not detect any transcripts of putative additional human LY49 genes. -

Gene Section Review
Atlas of Genetics and Cytogenetics in Oncology and Haematology INIST -CNRS OPEN ACCESS JOURNAL Gene Section Review KLRK1 (killer cell lectin-like receptor subfamily K, member 1) Lewis L Lanier UCSF, Department of Microbiology and Immunology, San Francisco, CA 94143-0414, USA (LLL) Published in Atlas Database: June 2014 Online updated version : http://AtlasGeneticsOncology.org/Genes/KLRK1ID41094ch12p13.html DOI: 10.4267/2042/56407 This article is an update of : Lanier LL. KLRK1 (killer cell lectin-like receptor subfamily K, member 1). Atlas Genet Cytogenet Oncol Haematol 2008;12(1):47- 49. This work is licensed under a Creative Commons Attribution-Noncommercial-No Derivative Works 2.0 France Licence. © 2015 Atlas of Genetics and Cytogenetics in Oncology and Haematology Abstract DNA/RNA KLRK1 encodes a type II transmembrane-anchored Note glycoprotein that is expressed as a disulfide-linked KLRK1 is present on chromosome 12 within a homodimer on the surface of Natural Killer (NK) cluster of genes referred to as the "NK complex" cells, gamma/delta TcR+ T cells, CD8+ T cells, and (NKC) because several genes that are preferentially a minor subset of CD4+ T cells. It associates non- expressed by Natural Killer (NK) cells are located covalently with the DAP10 signaling protein and in this region, including on the centromeric side provides activating or costimulatory signals to NK KLRD1 (CD94) and on the telomeric side KLRC4 cells and T cells. NKG2D binds to a family of (NKG2F), KLRC3 (NKG2E), KLRC2 (NKG2C), glycoproteins, in humans the MICA, MICB, and and KLRC1 (NKG2A) (Houchins et al., 1991). ULBP1-6 membrane proteins, which are frequently Description expressed on cells that have become infected with pathogens or undergone transformation. -

WO 2017/055322 Al O
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date WO 2017/055322 Al 6 April 2017 (06.04.2017) W P O PCT (51) International Patent Classification: Umrsl l38, Equipe 13, 15 rue de l'Ecole de Medecine, C12Q 1/68 (2006.01) 75006 Paris (FR). (21) International Application Number: (74) Agent: COLLIN, Matthieu; Inserm Transfert 7 rue Watt, PCT/EP2016/073053 75013 Paris (FR). (22) International Filing Date: (81) Designated States (unless otherwise indicated, for every 28 September 2016 (28.09.201 6) kind of national protection available): AE, AG, AL, AM, AO, AT, AU, AZ, BA, BB, BG, BH, BN, BR, BW, BY, (25) Filing Language: English BZ, CA, CH, CL, CN, CO, CR, CU, CZ, DE, DJ, DK, DM, (26) Publication Language: English DO, DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, HN, HR, HU, ID, IL, IN, IR, IS, JP, KE, KG, KN, KP, KR, (30) Priority Data: KW, KZ, LA, LC, LK, LR, LS, LU, LY, MA, MD, ME, EP15306521.4 29 September 2015 (29.09.2015) EP MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, EP16305886.0 12 July 2016 (12.07.2016) EP OM, PA, PE, PG, PH, PL, PT, QA, RO, RS, RU, RW, SA, (71) Applicants: INSERM (INSTITUT NATIONAL DE LA SC, SD, SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, SANTE ET DE LA RECHERCHE MEDICALE) TN, TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, [FR/FR]; 101, rue de Tolbiac, 75013 Paris (FR). -

Genes NKG2 and CD94 Common Chimpanzee Conservation And
Conservation and Variation in Human and Common Chimpanzee CD94 and NKG2 Genes This information is current as Benny P. Shum, Laura R. Flodin, David G. Muir, Raja of October 2, 2021. Rajalingam, Salim I. Khakoo, Sophia Cleland, Lisbeth A. Guethlein, Markus Uhrberg and Peter Parham J Immunol 2002; 168:240-252; ; doi: 10.4049/jimmunol.168.1.240 http://www.jimmunol.org/content/168/1/240 Downloaded from References This article cites 66 articles, 16 of which you can access for free at: http://www.jimmunol.org/content/168/1/240.full#ref-list-1 http://www.jimmunol.org/ Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication by guest on October 2, 2021 *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2002 by The American Association of Immunologists All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. Conservation and Variation in Human and Common Chimpanzee CD94 and NKG2 Genes1 Benny P. Shum, Laura R. Flodin, David G. Muir, Raja Rajalingam, Salim I. Khakoo,2 Sophia Cleland, Lisbeth A. -

Anti-CD314 / NKG2D Antibody (ARG59400)
Product datasheet [email protected] ARG59400 Package: 100 μl anti-CD314 / NKG2D antibody Store at: -20°C Summary Product Description Rabbit Polyclonal antibody recognizes CD314 / NKG2D Tested Reactivity Hu Tested Application ICC/IF Host Rabbit Clonality Polyclonal Isotype IgG Target Name CD314 / NKG2D Antigen Species Human Immunogen Synthetic peptide within aa. 50-150 of Human CD314 / NKG2D (NP_031386.2). Conjugation Un-conjugated Alternate Names LR; CD314; NKG2D; NKG2-D; D12S2489E Application Instructions Application table Application Dilution ICC/IF 1:50 - 1:200 Application Note * The dilutions indicate recommended starting dilutions and the optimal dilutions or concentrations should be determined by the scientist. Calculated Mw 25 kDa Properties Form Liquid Purification Affinity purified. Buffer PBS (pH 7.3), 0.02% Sodium azide and 50% Glycerol. Preservative 0.02% Sodium azide Stabilizer 50% Glycerol Storage instruction For continuous use, store undiluted antibody at 2-8°C for up to a week. For long-term storage, aliquot and store at -20°C. Storage in frost free freezers is not recommended. Avoid repeated freeze/thaw cycles. Suggest spin the vial prior to opening. The antibody solution should be gently mixed before use. Note For laboratory research only, not for drug, diagnostic or other use. Bioinformation www.arigobio.com 1/2 Gene Symbol KLRK1 Gene Full Name killer cell lectin like receptor K1 Background Natural killer (NK) cells are lymphocytes that can mediate lysis of certain tumor cells and virus-infected cells without previous activation. They can also regulate specific humoral and cell-mediated immunity. NK cells preferentially express several calcium-dependent (C-type) lectins, which have been implicated in the regulation of NK cell function. -

Terminal Effector CD8 T Cells Defined by an IKZF2+IL-7R
Terminal Effector CD8 T Cells Defined by an IKZF2 +IL-7R− Transcriptional Signature Express Fc γRIIIA, Expand in HIV Infection, and Mediate Potent HIV-Specific This information is current as Antibody-Dependent Cellular Cytotoxicity of September 23, 2021. Prossy Naluyima, Kerri G. Lal, Margaret C. Costanzo, Gustavo H. Kijak, Veronica D. Gonzalez, Kim Blom, Leigh Anne Eller, Matthew Creegan, Ting Hong, Dohoon Kim, Thomas C. Quinn, Niklas K. Björkström, Hans-Gustaf Downloaded from Ljunggren, David Serwadda, Elly T. Katabira, Nelson K. Sewankambo, Ronald H. Gray, Jared M. Baeten, Nelson L. Michael, Fred Wabwire-Mangen, Merlin L. Robb, Diane L. Bolton, Johan K. Sandberg and Michael A. Eller J Immunol published online 13 September 2019 http://www.jimmunol.org/ http://www.jimmunol.org/content/early/2019/09/12/jimmun ol.1900422 Supplementary http://www.jimmunol.org/content/suppl/2019/09/12/jimmunol.190042 Material 2.DCSupplemental by guest on September 23, 2021 Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Author Choice Freely available online through The Journal of Immunology Author Choice option Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2019 The Authors. -

KLRC4 (NM 013431) Human Tagged ORF Clone – RG204640 | Origene
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for RG204640 KLRC4 (NM_013431) Human Tagged ORF Clone Product data: Product Type: Expression Plasmids Product Name: KLRC4 (NM_013431) Human Tagged ORF Clone Tag: TurboGFP Symbol: KLRC4 Synonyms: NKG2-F; NKG2F Vector: pCMV6-AC-GFP (PS100010) E. coli Selection: Ampicillin (100 ug/mL) Cell Selection: Neomycin ORF Nucleotide >RG204640 representing NM_013431 Sequence: Red=Cloning site Blue=ORF Green=Tags(s) TTTTGTAATACGACTCACTATAGGGCGGCCGGGAATTCGTCGACTGGATCCGGTACCGAGGAGATCTGCC GCCGCGATCGCC ATGAATAAACAAAGAGGAACCTACTCAGAAGTGAGTCTGGCCCAGGACCCAAAGAGGCAGCAAAGGAAAC TTAAGGGCAATAAAATCTCCATTTCAGGAACCAAACAGGAAATATTCCAAGTAGAATTAAACCTTCAAAA TGCTTCTTCGGATCATCAAGGGAATGACAAGACATATCACTGCAAAGGTTTACTGCCACCTCCAGAGAAG CTCACTGCTGAGGTCCTAGGAATCATTTGCATTGTCCTGATGGCCACTGTGTTAAAAACAATAGTTCTTA TTCCTTGTATTGGAGTACTGGAGCAGAACAATTTTTCCCTGAATAGAAGAATGCAGAAAGCACGTCATTG TGGCCATTGTCCTGAGGAGTGGATTACATATTCCAACAGTTGTTATTACATTGGTAAGGAAAGAAGAACT TGGGAAGAAAGAGTTTGCTGGCCTGTGCTTCGAAGAACTCTGATCTGCTTTCTA ACGCGTACGCGGCCGCTCGAG - GFP Tag - GTTTAA Protein Sequence: >RG204640 representing NM_013431 Red=Cloning site Green=Tags(s) MNKQRGTYSEVSLAQDPKRQQRKLKGNKISISGTKQEIFQVELNLQNASSDHQGNDKTYHCKGLLPPPEK LTAEVLGIICIVLMATVLKTIVLIPCIGVLEQNNFSLNRRMQKARHCGHCPEEWITYSNSCYYIGKERRT WEERVCWPVLRRTLICFL TRTRPLE - GFP Tag - V Restriction Sites: SgfI-MluI This product is to be used for laboratory only. Not -

NKG2D Natural Killer Cell Receptor—A Short Description and Potential Clinical Applications
cells Review NKG2D Natural Killer Cell Receptor—A Short Description and Potential Clinical Applications Jagoda Siemaszko 1, Aleksandra Marzec-Przyszlak 2 and Katarzyna Bogunia-Kubik 1,* 1 Laboratory of Clinical Immunogenetics and Pharmacogenetics, Hirszfeld Institute of Immunology and Experimental Therapy, Polish Academy of Sciences, 53114 Wroclaw, Poland; [email protected] 2 Department of Biosensors and Processing of Biomedical Signals, Faculty of Biomedical Engineering, Silesian University of Technology, 41800 Zabrze, Poland; [email protected] * Correspondence: [email protected] Abstract: Natural Killer (NK) cells are natural cytotoxic, effector cells of the innate immune system. They can recognize transformed or infected cells. NK cells are armed with a set of activating and inhibitory receptors which are able to bind to their ligands on target cells. The right balance between expression and activation of those receptors is fundamental for the proper functionality of NK cells. One of the best known activating receptors is NKG2D, a member of the CD94/NKG2 family. Due to a specific NKG2D binding with its eight different ligands, which are overexpressed in transformed, infected and stressed cells, NK cells are able to recognize and attack their targets. The NKG2D receptor has an enormous significance in various, autoimmune diseases, viral and bacterial infections as well as for transplantation outcomes and complications. This review focuses on the NKG2D receptor, the mechanism of its action, clinical relevance of its gene polymorphisms and a potential application in various clinical settings. Citation: Siemaszko, J.; Marzec-Przyszlak, A.; Keywords: NKG2D receptor; NKG2D ligands; NK cells Bogunia-Kubik, K. NKG2D Natural Killer Cell Receptor—A Short Description and Potential Clinical Applications.