A Pan-Cancer Analysis of CD161, a Potential New Immune Checkpoint
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Downloaded the Normalized Gene Expression Profile from the GEO (Package Geoquery (5))
1 SUPPLEMENTARY MATERIALS 2 Supplementary Methods 3 Study design 4 The workflow to identify and validate the TME risk score and TME subtypes in gastric cancer is 5 depicted in Fig. S1. We first estimated the absolute abundance levels of the major stromal and 6 immune cell types in the TME using bulk gene expression data, and assessed the prognostic 7 effect of these cells in a discovery cohort. Next, we constructed a TME risk score and validated it 8 in two independent gene expression validation cohorts and three immunohistochemistry 9 validation cohorts. Finally, we stratified patients into four TME subtypes and examined their 10 genomic and molecular features and relation to established molecular subtypes. 11 Gene expression data 12 To explore the prognostic landscape of the TME, we used four gene expression profile (GEP) 13 datasets of resected gastric cancer patients with publicly available clinical information, namely, 14 ACRG (GSE62254) (1), GSE15459 (2), GSE84437 (3), and TCGA stomach adenocarcinoma 15 (STAD). Specifically, the raw microarray data in the ACRG cohort and GSE15459 were retrieved 16 from the Gene Expression Omnibus (GEO), and normalized by the RMA algorithm (package affy) 17 using custom chip definition files (Brainarray version 23 (4)) that convert Affymetrix probesets to 18 Entrez gene IDs. For GSE84437 dataset, which was measured by the Illumina platform, we 19 downloaded the normalized gene expression profile from the GEO (package GEOquery (5)). The 20 Illumina probes were also mapped to Entrez genes. For multiple probes mapping to the same 21 Entrez gene, we selected the one with the maximum mean expression level as the surrogate for 22 the Entrez gene using the function of collapseRows (6) (package WGCNA). -

The Inhibitory NKR-P1B:Clr-B Recognition Axis Facilitates Detection of Oncogenic Transformation and Cancer Immunosurveillance Miho Tanaka1,2, Jason H
Published OnlineFirst April 24, 2018; DOI: 10.1158/0008-5472.CAN-17-1688 Cancer Tumor Biology and Immunology Research The Inhibitory NKR-P1B:Clr-b Recognition Axis Facilitates Detection of Oncogenic Transformation and Cancer Immunosurveillance Miho Tanaka1,2, Jason H. Fine1,2, Christina L. Kirkham1,2, Oscar A. Aguilar1,2, Antoaneta Belcheva1, Alberto Martin1, Troy Ketela3,4, Jason Moffat3,4, David S.J. Allan1,2, and James R. Carlyle1,2 Abstract Natural killer (NK) cells express receptors specific for MHC class in a primary lymphoma model despite preferential rejection of – – I (MHC-I) molecules involved in "missing-self" recognition of Clr-b / hematopoietic cells previously observed following adop- cancer and virus-infected cells. Here we elucidate the role of MHC- tive transfer into na€ve wild-type mice in vivo. Collectively, these I-independent NKR-P1B:Clr-b interactions in the detection of findings suggest that the inhibitory NKR-P1B:Clr-b axis plays a oncogenic transformation by NK cells. Ras oncogene overexpres- beneficial role in innate detection of oncogenic transformation sion was found to promote a real-time loss of Clr-b on mouse via NK-cell–mediated cancer immune surveillance, in addition fibroblasts and leukemia cells, mediated in part via the Raf/MEK/ to a pathologic role in the immune escape of primary lympho- ERK and PI3K pathways. Ras-driven Clr-b downregulation ma cells in Em-cMyc mice in vivo. These results provide a model occurred at the level of the Clrb (Clec2d) promoter, nascent Clr-b for the human NKR-P1A:LLT1 system in cancer immunosur- transcripts, and cell surface Clr-b protein, in turn promoting veillance in patients with lymphoma and suggest it may rep- missing-self recognition via the NKR-P1B inhibitory receptor. -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

Human Lectins, Their Carbohydrate Affinities and Where to Find Them
biomolecules Review Human Lectins, Their Carbohydrate Affinities and Where to Review HumanFind Them Lectins, Their Carbohydrate Affinities and Where to FindCláudia ThemD. Raposo 1,*, André B. Canelas 2 and M. Teresa Barros 1 1, 2 1 Cláudia D. Raposo * , Andr1 é LAQVB. Canelas‐Requimte,and Department M. Teresa of Chemistry, Barros NOVA School of Science and Technology, Universidade NOVA de Lisboa, 2829‐516 Caparica, Portugal; [email protected] 12 GlanbiaLAQV-Requimte,‐AgriChemWhey, Department Lisheen of Chemistry, Mine, Killoran, NOVA Moyne, School E41 of ScienceR622 Co. and Tipperary, Technology, Ireland; canelas‐ [email protected] NOVA de Lisboa, 2829-516 Caparica, Portugal; [email protected] 2* Correspondence:Glanbia-AgriChemWhey, [email protected]; Lisheen Mine, Tel.: Killoran, +351‐212948550 Moyne, E41 R622 Tipperary, Ireland; [email protected] * Correspondence: [email protected]; Tel.: +351-212948550 Abstract: Lectins are a class of proteins responsible for several biological roles such as cell‐cell in‐ Abstract:teractions,Lectins signaling are pathways, a class of and proteins several responsible innate immune for several responses biological against roles pathogens. such as Since cell-cell lec‐ interactions,tins are able signalingto bind to pathways, carbohydrates, and several they can innate be a immuneviable target responses for targeted against drug pathogens. delivery Since sys‐ lectinstems. In are fact, able several to bind lectins to carbohydrates, were approved they by canFood be and a viable Drug targetAdministration for targeted for drugthat purpose. delivery systems.Information In fact, about several specific lectins carbohydrate were approved recognition by Food by andlectin Drug receptors Administration was gathered for that herein, purpose. plus Informationthe specific organs about specific where those carbohydrate lectins can recognition be found by within lectin the receptors human was body. -

Literature Mining Sustains and Enhances Knowledge Discovery from Omic Studies
LITERATURE MINING SUSTAINS AND ENHANCES KNOWLEDGE DISCOVERY FROM OMIC STUDIES by Rick Matthew Jordan B.S. Biology, University of Pittsburgh, 1996 M.S. Molecular Biology/Biotechnology, East Carolina University, 2001 M.S. Biomedical Informatics, University of Pittsburgh, 2005 Submitted to the Graduate Faculty of School of Medicine in partial fulfillment of the requirements for the degree of Doctor of Philosophy University of Pittsburgh 2016 UNIVERSITY OF PITTSBURGH SCHOOL OF MEDICINE This dissertation was presented by Rick Matthew Jordan It was defended on December 2, 2015 and approved by Shyam Visweswaran, M.D., Ph.D., Associate Professor Rebecca Jacobson, M.D., M.S., Professor Songjian Lu, Ph.D., Assistant Professor Dissertation Advisor: Vanathi Gopalakrishnan, Ph.D., Associate Professor ii Copyright © by Rick Matthew Jordan 2016 iii LITERATURE MINING SUSTAINS AND ENHANCES KNOWLEDGE DISCOVERY FROM OMIC STUDIES Rick Matthew Jordan, M.S. University of Pittsburgh, 2016 Genomic, proteomic and other experimentally generated data from studies of biological systems aiming to discover disease biomarkers are currently analyzed without sufficient supporting evidence from the literature due to complexities associated with automated processing. Extracting prior knowledge about markers associated with biological sample types and disease states from the literature is tedious, and little research has been performed to understand how to use this knowledge to inform the generation of classification models from ‘omic’ data. Using pathway analysis methods to better understand the underlying biology of complex diseases such as breast and lung cancers is state-of-the-art. However, the problem of how to combine literature- mining evidence with pathway analysis evidence is an open problem in biomedical informatics research. -

CLEC2D (NM 001197317) Human Tagged ORF Clone – RG230997 | Origene
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for RG230997 CLEC2D (NM_001197317) Human Tagged ORF Clone Product data: Product Type: Expression Plasmids Product Name: CLEC2D (NM_001197317) Human Tagged ORF Clone Tag: TurboGFP Symbol: CLEC2D Synonyms: CLAX; LLT1; OCIL Vector: pCMV6-AC-GFP (PS100010) E. coli Selection: Ampicillin (100 ug/mL) Cell Selection: Neomycin ORF Nucleotide >RG230997 representing NM_001197317 Sequence: Red=Cloning site Blue=ORF Green=Tags(s) TTTTGTAATACGACTCACTATAGGGCGGCCGGGAATTCGTCGACTGGATCCGGTACCGAGGAGATCTGCC GCCGCGATCGCC ATGCATGACAGTAACAATGTGGAGAAAGACATTACACCATCTGAATTGCCTGCAAACCCAGCAATAAGAG CTAACTGCCATCAAGAGCCATCAGTATGTCTTCAAGCTGCATGCCCAGAAAGCTGGATTGGTTTTCAAAG AAAGTGTTTCTATTTTTCTGATGACACCAAGAACTGGACATCAAGTCAGAGGTTTTGTGACTCACAAGAT GCTGATCTTGCTCAGGTTGAAAGCTTCCAGGAACTGAATTTCCTGTTGAGATATAAAGGCCCATCTGATC ACTGGATTGGGCTGAGCAGAGAACAAGGCCAACCATGGAAATGGATAAATGGTACTGAATGGACAAGACA GTTTCCTATCCTGGGAGCAGGAGAGTGTGCCTATTTGAATGACAAAGGTGCCAGTAGTGCCAGGCACTAC ACAGAGAGGAAGTGGATTTGTTCCAAATCAGATATACATGTC ACGCGTACGCGGCCGCTCGAG - GFP Tag - GTTTAA Protein Sequence: >RG230997 representing NM_001197317 Red=Cloning site Green=Tags(s) MHDSNNVEKDITPSELPANPAIRANCHQEPSVCLQAACPESWIGFQRKCFYFSDDTKNWTSSQRFCDSQD ADLAQVESFQELNFLLRYKGPSDHWIGLSREQGQPWKWINGTEWTRQFPILGAGECAYLNDKGASSARHY TERKWICSKSDIHV TRTRPLE - GFP Tag - V Restriction Sites: SgfI-MluI This product is to be used for laboratory only. Not -

C07k - 2021.08
CPC - C07K - 2021.08 C07K PEPTIDES (peptides in foodstuffs A23; obtaining protein compositions for foodstuffs, working-up proteins for foodstuffs A23J; preparations for medicinal purposes A61K; peptides containing beta-lactam rings C07D; cyclic dipeptides not having in their molecule any other peptide link than those which form their ring, e.g. piperazine-2,5-diones, C07D; ergot alkaloids of the cyclic peptide type C07D 519/02; macromolecular compounds having statistically distributed amino acid units in their molecules, i.e. when the preparation does not provide for a specific; but for a random sequence of the amino acid units, homopolyamides and block copolyamides derived from amino acids C08G 69/00; macromolecular products derived from proteins C08H 1/00; preparation of glue or gelatine C09H; single cell proteins, enzymes C12N; genetic engineering processes for obtaining peptides C12N 15/00; compositions for measuring or testing processes involving enzymes C12Q; investigation or analysis of biological material G01N 33/00) Relationships with other classification places An amino acid per se is classified in C07D while peptides (starting from dipeptides) are classified in C07K. Subclass C07K is a function oriented entry for the compounds themselves and does not cover the application or use of the compounds under the subclass definition. For classifying such information other entries exist, for example: preservation of bodies of humans or animals or plants or parts thereof; Biocides, e.g. as disinfectants, as pesticides, as herbicides; pest repellants or attractants; plant growth regulators are classified in A01N. Preparations for medical, dental, or toilet purposes are classified in A61K. Amino acids or derivatives thereof are classified in C07C or C07D. -

UC San Francisco Previously Published Works
UCSF UC San Francisco Previously Published Works Title FitSNPs: highly differentially expressed genes are more likely to have variants associated with disease. Permalink https://escholarship.org/uc/item/91k8p2km Journal Genome biology, 9(12) ISSN 1474-7596 Authors Chen, Rong Morgan, Alex A Dudley, Joel et al. Publication Date 2008 DOI 10.1186/gb-2008-9-12-r170 Peer reviewed eScholarship.org Powered by the California Digital Library University of California Open Access Research2008ChenetVolume al. 9, Issue 12, Article R170 FitSNPs: highly differentially expressed genes are more likely to have variants associated with disease Rong Chen*†‡, Alex A Morgan*†‡, Joel Dudley*†‡, Tarangini Deshpande§, Li Li†, Keiichi Kodama*†‡, Annie P Chiang*†‡ and Atul J Butte*†‡ Addresses: *Stanford Center for Biomedical Informatics Research, 251 Cmpus Drive, Stanford, CA 94305, USA. †Department of Pediatrics, Stanford University School of Medicine, Stanford, CA 94305, USA. ‡Lucile Packard Children's Hospital, 725 Welch Road, Palo Alto, CA 94304, USA. §NuMedii Inc., Menlo Park, CA 94025, USA. Correspondence: Atul J Butte. Email: [email protected] Published: 5 December 2008 Received: 17 June 2008 Revised: 26 September 2008 Genome Biology 2008, 9:R170 (doi:10.1186/gb-2008-9-12-r170) Accepted: 5 December 2008 The electronic version of this article is the complete one and can be found online at http://genomebiology.com/2008/9/12/R170 © 2008 Chen et al.; licensee BioMed Central Ltd. This is an open access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. -

Single-Cell Transcriptomes Reveal a Complex Cellular Landscape in the Middle Ear and Differential Capacities for Acute Response to Infection
fgene-11-00358 April 9, 2020 Time: 15:55 # 1 ORIGINAL RESEARCH published: 15 April 2020 doi: 10.3389/fgene.2020.00358 Single-Cell Transcriptomes Reveal a Complex Cellular Landscape in the Middle Ear and Differential Capacities for Acute Response to Infection Allen F. Ryan1*, Chanond A. Nasamran2, Kwang Pak1, Clara Draf1, Kathleen M. Fisch2, Nicholas Webster3 and Arwa Kurabi1 1 Departments of Surgery/Otolaryngology, UC San Diego School of Medicine, VA Medical Center, La Jolla, CA, United States, 2 Medicine/Center for Computational Biology & Bioinformatics, UC San Diego School of Medicine, VA Medical Center, La Jolla, CA, United States, 3 Medicine/Endocrinology, UC San Diego School of Medicine, VA Medical Center, La Jolla, CA, United States Single-cell transcriptomics was used to profile cells of the normal murine middle ear. Clustering analysis of 6770 transcriptomes identified 17 cell clusters corresponding to distinct cell types: five epithelial, three stromal, three lymphocyte, two monocyte, Edited by: two endothelial, one pericyte and one melanocyte cluster. Within some clusters, Amélie Bonnefond, Institut National de la Santé et de la cell subtypes were identified. While many corresponded to those cell types known Recherche Médicale (INSERM), from prior studies, several novel types or subtypes were noted. The results indicate France unexpected cellular diversity within the resting middle ear mucosa. The resolution of Reviewed by: Fabien Delahaye, uncomplicated, acute, otitis media is too rapid for cognate immunity to play a major Institut Pasteur de Lille, France role. Thus innate immunity is likely responsible for normal recovery from middle ear Nelson L. S. Tang, infection. The need for rapid response to pathogens suggests that innate immune The Chinese University of Hong Kong, China genes may be constitutively expressed by middle ear cells. -

Genetic Mapping of the Major Histocompatibility Complex in the Zebra Finch (Taeniopygia Guttata)
This is a repository copy of Genetic mapping of the major histocompatibility complex in the zebra finch (Taeniopygia guttata). White Rose Research Online URL for this paper: http://eprints.whiterose.ac.uk/152446/ Version: Accepted Version Article: Ekblom, R., Stapley, J., Ball, A.D. et al. (3 more authors) (2011) Genetic mapping of the major histocompatibility complex in the zebra finch (Taeniopygia guttata). Immunogenetics, 63 (8). pp. 523-530. ISSN 0093-7711 https://doi.org/10.1007/s00251-011-0525-9 This is a post-peer-review, pre-copyedit version of an article published in Immunogenetics. The final authenticated version is available online at: http://dx.doi.org/10.1007/s00251-011-0525-9. Reuse Items deposited in White Rose Research Online are protected by copyright, with all rights reserved unless indicated otherwise. They may be downloaded and/or printed for private study, or other acts as permitted by national copyright laws. The publisher or other rights holders may allow further reproduction and re-use of the full text version. This is indicated by the licence information on the White Rose Research Online record for the item. Takedown If you consider content in White Rose Research Online to be in breach of UK law, please notify us by emailing [email protected] including the URL of the record and the reason for the withdrawal request. [email protected] https://eprints.whiterose.ac.uk/ Manuscript Click here to download Manuscript: zfMHCmapp_manuscript_resubm2.doc Click here to view linked References 1 Genetic mapping of the major histocompatibility complex in the 1 2 3 4 2 zebra finch (Taeniopygia guttata) 5 6 3 7 8 1,2* 2 2, 3 2 2 9 4 Robert Ekblom , Jessica Stapley , Alex D. -
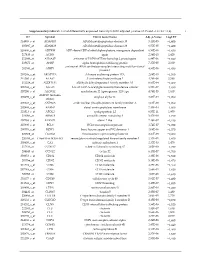
Supplementary Table S1. List of Differentially Expressed
Supplementary table S1. List of differentially expressed transcripts (FDR adjusted p‐value < 0.05 and −1.4 ≤ FC ≥1.4). 1 ID Symbol Entrez Gene Name Adj. p‐Value Log2 FC 214895_s_at ADAM10 ADAM metallopeptidase domain 10 3,11E‐05 −1,400 205997_at ADAM28 ADAM metallopeptidase domain 28 6,57E‐05 −1,400 220606_s_at ADPRM ADP‐ribose/CDP‐alcohol diphosphatase, manganese dependent 6,50E‐06 −1,430 217410_at AGRN agrin 2,34E‐10 1,420 212980_at AHSA2P activator of HSP90 ATPase homolog 2, pseudogene 6,44E‐06 −1,920 219672_at AHSP alpha hemoglobin stabilizing protein 7,27E‐05 2,330 aminoacyl tRNA synthetase complex interacting multifunctional 202541_at AIMP1 4,91E‐06 −1,830 protein 1 210269_s_at AKAP17A A‐kinase anchoring protein 17A 2,64E‐10 −1,560 211560_s_at ALAS2 5ʹ‐aminolevulinate synthase 2 4,28E‐06 3,560 212224_at ALDH1A1 aldehyde dehydrogenase 1 family member A1 8,93E‐04 −1,400 205583_s_at ALG13 ALG13 UDP‐N‐acetylglucosaminyltransferase subunit 9,50E‐07 −1,430 207206_s_at ALOX12 arachidonate 12‐lipoxygenase, 12S type 4,76E‐05 1,630 AMY1C (includes 208498_s_at amylase alpha 1C 3,83E‐05 −1,700 others) 201043_s_at ANP32A acidic nuclear phosphoprotein 32 family member A 5,61E‐09 −1,760 202888_s_at ANPEP alanyl aminopeptidase, membrane 7,40E‐04 −1,600 221013_s_at APOL2 apolipoprotein L2 6,57E‐11 1,600 219094_at ARMC8 armadillo repeat containing 8 3,47E‐08 −1,710 207798_s_at ATXN2L ataxin 2 like 2,16E‐07 −1,410 215990_s_at BCL6 BCL6 transcription repressor 1,74E‐07 −1,700 200776_s_at BZW1 basic leucine zipper and W2 domains 1 1,09E‐06 −1,570 222309_at -

LLT1) Checkpoint: a Novel Independent Prognostic Factor in HPV-Negative Oropharyngeal Squamous Cell Carcinoma
biomedicines Article Lectin-Like Transcript 1 (LLT1) Checkpoint: A Novel Independent Prognostic Factor in HPV-Negative Oropharyngeal Squamous Cell Carcinoma 1,2, 1,2,3, 2,4 Mario Sanchez-Canteli y, Francisco Hermida-Prado y , Christian Sordo-Bahamonde , Irene Montoro-Jiménez 1,2, Esperanza Pozo-Agundo 1,2, Eva Allonca 1,2 , Aitana Vallina-Álvarez 2,5,César Álvarez-Marcos 1,2,3, Segundo Gonzalez 2,4 , Juana M. García-Pedrero 1,2,3,* and Juan P. Rodrigo 1,2,3,* 1 Department of Otolaryngology, Hospital Universitario Central de Asturias, Instituto de Investigación Sanitaria del Principado de Asturias, 33011 Oviedo, Spain; [email protected] (M.S.-C.); [email protected] (F.H.-P.); [email protected] (I.M.-J.); [email protected] (E.P.-A.); [email protected] (E.A.); [email protected] (C.Á.-M.) 2 Instituto Universitario de Oncología del Principado de Asturias, University of Oviedo, 33006 Oviedo, Spain; [email protected] (C.S.-B.); [email protected] (A.V.-Á.); [email protected] (S.G.) 3 CIBERONC, Instituto de Salud Carlos III, 28029 Madrid, Spain 4 Department of Functional Biology, Instituto de Investigación Sanitaria del Principado de Asturias, University of Oviedo, 33006 Oviedo, Spain 5 Department of Pathology, Hospital Universitario Central de Asturias, ISPA, 33011 Oviedo, Spain * Correspondence: juanagp.fi[email protected] (J.M.G.-P.); [email protected] (J.P.R.) These authors contributed equally to this work. y Received: 30 October 2020; Accepted: 23 November 2020; Published: 25 November 2020 Abstract: Lectin-like transcript 1 (LLT1) expression by tumor cells contributes to immune evasion, thereby emerging as a natural killer (NK) cell-mediated immunotherapeutic target.