Molecular Phylogeography of the Tench
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

5Th Indo-Pacific Fish Conference
)tn Judo - Pacifi~ Fish Conference oun a - e II denia ( vernb ~ 3 - t 1997 A ST ACTS Organized by Under the aegis of L'Institut français Société de recherche scientifique Française pour le développement d'Ichtyologie en coopération ' FI Fish Conference Nouméa - New Caledonia November 3 - 8 th, 1997 ABSTRACTS LATE ARRIVAL ZOOLOGICAL CATALOG OF AUSTRALIAN FISHES HOESE D.F., PAXTON J. & G. ALLEN Australian Museum, Sydney, Australia Currently over 4000 species of fishes are known from Australia. An analysis ofdistribution patterns of 3800 species is presented. Over 20% of the species are endemic to Australia, with endemic species occuiring primarily in southern Australia. There is also a small component of the fauna which is found only in the southwestern Pacific (New Caledonia, Lord Howe Island, Norfolk Island and New Zealand). The majority of the other species are widely distributed in the western Pacific Ocean. AGE AND GROWTH OF TROPICAL TUNAS FROM THE WESTERN CENTRAL PACIFIC OCEAN, AS INDICATED BY DAILY GROWm INCREMENTS AND TAGGING DATA. LEROY B. South Pacific Commission, Nouméa, New Caledonia The Oceanic Fisheries Programme of the South Pacific Commission is currently pursuing a research project on age and growth of two tropical tuna species, yellowfm tuna (Thunnus albacares) and bigeye tuna (Thunnus obesus). The daily periodicity of microincrements forrned with the sagittal otoliths of these two spceies has been validated by oxytetracycline marking in previous studies. These validation studies have come from fishes within three regions of the Pacific (eastem, central and western tropical Pacific). Otolith microincrements are counted along transverse section with a light microscope. -

Genome Size Covaries More Positively with Propagule Size Than Adult Size: New Insights Into an Old Problem
biology Article Genome Size Covaries More Positively with Propagule Size than Adult Size: New Insights into an Old Problem Douglas S. Glazier Department of Biology, Juniata College, Huntingdon, PA 16652, USA; [email protected] Simple Summary: The amount of hereditary information (DNA) contained in the cell nuclei of larger or more complex organisms is often no greater than that of smaller or simpler organisms. Why this is so is an evolutionary mystery. Here, I show that the amount of DNA per cell nucleus (‘genome size’) relates more positively to egg size than body size in crustaceans (including shrimp, lobsters and crabs). Genome size also seems to relate more to the size of eggs or other gametes and reproductive propagules (e.g., sperm, spores, pollen and seeds) than to adult size in other animals and plants. I explain these patterns as being the result of genome size relating more to cell size (including that of single-celled eggs) than the number of cells in a body. Since most organisms begin life as single cells or propagules with relatively few cells, propagule size may importantly affect or be affected by genome size regardless of body size. Relationships between genome size and body size should thus become weaker as body size (and the amount of cell multiplication required during development) increases, as observed in crustaceans and other kinds of organisms. Abstract: The body size and (or) complexity of organisms is not uniformly related to the amount of genetic material (DNA) contained in each of their cell nuclei (‘genome size’). This surprising mismatch between the physical structure of organisms and their underlying genetic information appears to relate to variable accumulation of repetitive DNA sequences, but why this variation has Citation: Glazier, D.S. -

Aphanomyces Invadans R NUMBER: R6979 PROGRAMME: Aquaculture Research Programme (ARP) PROGRAMME MANAGER (INSTITUTION): Prof J.F
DFID Applied studies on epizootic ulcerative syndrome Final Report February 2001 Project R6979 of the Aquaculture Research Programme of the Department for International Development of the United Kingdom This document is an output from a project funded by the UK Department for International Development (DFID) for the benefit of developing countries. The views expressed are not necessarily those of DFID. Hard copies of this report can be obtained from: ARP Manager Institute of Aquaculture University of Stirling Stirling FK9 4LA Scotland, UK Institute of Aquaculture University of Stirling, Scotland Aquatic Animal Health Research Institute Department of Fisheries, Thailand PROJECT COMPLETION SUMMARY SHEET Sheet Completed 28 February 2001 TITLE OF PROJECT: Applied studies on EUS - The ecology, immunogenicity and treatment of Aphanomyces invadans R NUMBER: R6979 PROGRAMME: Aquaculture Research Programme (ARP) PROGRAMME MANAGER (INSTITUTION): Prof J.F. Muir PROGRAMME PURPOSE: Productive benefits of aquatic resources for poor people generated through improved knowledge of aquaculture processes and their management PRODUCTION SYSTEM: Land/Water Interface BENEFICIARIES: Asian freshwater fish farmers TARGET INSTITUTIONS: AAHRI, FRI, BAU, CARE-LIFE, CIFA, BFAR, FDD, FRTI, RIA1 GEOGRAPHIC FOCUS: Southeast and South Asia Planned Actual START DATE: 1 July 1997 1 July 1997 FINISH DATE: 31 December 2000 28 February 2001 TOTAL COST: £250,613 £248,613 1. Project purpose: To generate the information needed for the formulation of strategies to contain EUS; and to develop and introduce improved prophylactic and therapeutic treatments to provide fish farmers with a means of reducing losses due to EUS. 2. Outputs: Some modifications were made to the original logical framework in January 1999. -
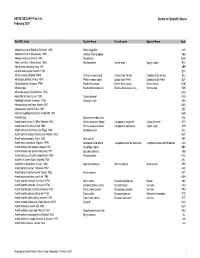
ASFIS ISSCAAP Fish List February 2007 Sorted on Scientific Name
ASFIS ISSCAAP Fish List Sorted on Scientific Name February 2007 Scientific name English Name French name Spanish Name Code Abalistes stellaris (Bloch & Schneider 1801) Starry triggerfish AJS Abbottina rivularis (Basilewsky 1855) Chinese false gudgeon ABB Ablabys binotatus (Peters 1855) Redskinfish ABW Ablennes hians (Valenciennes 1846) Flat needlefish Orphie plate Agujón sable BAF Aborichthys elongatus Hora 1921 ABE Abralia andamanika Goodrich 1898 BLK Abralia veranyi (Rüppell 1844) Verany's enope squid Encornet de Verany Enoploluria de Verany BLJ Abraliopsis pfefferi (Verany 1837) Pfeffer's enope squid Encornet de Pfeffer Enoploluria de Pfeffer BJF Abramis brama (Linnaeus 1758) Freshwater bream Brème d'eau douce Brema común FBM Abramis spp Freshwater breams nei Brèmes d'eau douce nca Bremas nep FBR Abramites eques (Steindachner 1878) ABQ Abudefduf luridus (Cuvier 1830) Canary damsel AUU Abudefduf saxatilis (Linnaeus 1758) Sergeant-major ABU Abyssobrotula galatheae Nielsen 1977 OAG Abyssocottus elochini Taliev 1955 AEZ Abythites lepidogenys (Smith & Radcliffe 1913) AHD Acanella spp Branched bamboo coral KQL Acanthacaris caeca (A. Milne Edwards 1881) Atlantic deep-sea lobster Langoustine arganelle Cigala de fondo NTK Acanthacaris tenuimana Bate 1888 Prickly deep-sea lobster Langoustine spinuleuse Cigala raspa NHI Acanthalburnus microlepis (De Filippi 1861) Blackbrow bleak AHL Acanthaphritis barbata (Okamura & Kishida 1963) NHT Acantharchus pomotis (Baird 1855) Mud sunfish AKP Acanthaxius caespitosa (Squires 1979) Deepwater mud lobster Langouste -

Viet Nam Ramsar Information Sheet Published on 16 October 2018
RIS for Site no. 2360, Van Long Wetland Nature Reserve, Viet Nam Ramsar Information Sheet Published on 16 October 2018 Viet Nam Van Long Wetland Nature Reserve Designation date 10 February 2017 Site number 2360 Coordinates 20°23'35"N 105°51'10"E Area 2 736,00 ha https://rsis.ramsar.org/ris/2360 Created by RSIS V.1.6 on - 16 October 2018 RIS for Site no. 2360, Van Long Wetland Nature Reserve, Viet Nam Color codes Fields back-shaded in light blue relate to data and information required only for RIS updates. Note that some fields concerning aspects of Part 3, the Ecological Character Description of the RIS (tinted in purple), are not expected to be completed as part of a standard RIS, but are included for completeness so as to provide the requested consistency between the RIS and the format of a ‘full’ Ecological Character Description, as adopted in Resolution X.15 (2008). If a Contracting Party does have information available that is relevant to these fields (for example from a national format Ecological Character Description) it may, if it wishes to, include information in these additional fields. 1 - Summary Summary Van Long Wetland Nature Reserve is a wetland comprised of rivers and a shallow lake with large amounts of submerged vegetation. The wetland area is centred on a block of limestone karst that rises abruptly from the flat coastal plain of the northern Vietnam. It is located within the Gia Vien district of Ninh Binh Province. The wetland is one of the rarest intact lowland inland wetlands remaining in the Red River Delta, Vietnam. -

Squalius Alburnoides
UNIVERSIDADE DE LISBOA FACULDADE DE CIÊNCIAS Departamento de Biologia Animal Reproductive behaviour and the evolutionary history of the hybridogenetic complex Squalius alburnoides (Pisces, Cyprinidae) CARLA PATRÍCIA CÂNDIDO DE SOUSA SANTOS Doutoramento em Biologia (Biologia Evolutiva) 2007 UNIVERSIDADE DE LISBOA FACULDADE DE CIÊNCIAS Departamento de Biologia Animal Reproductive behaviour and the evolutionary history of the hybridogenetic complex Squalius alburnoides (Pisces, Cyprinidae) CARLA PATRÍCIA CÂNDIDO DE SOUSA SANTOS Tese co-orientada por: Professor Doutor Vítor Almada (ISPA) Professora Doutora Maria João Collares-Pereira (FCUL) Doutoramento em Biologia (Biologia Evolutiva) 2007 Os trabalhos de investigação realizados durante esta tese tiveram o apoio financeiro da Fundação para a Ciência e a Tecnologia (Bolsa de Doutoramento SFRH/BD/8320/2002 e Programas Plurianuais U I&D 331/94 e U I&D 329/94). The research conducted in this thesis was funded by the FCT Pluriannual Program U I&D 331/94 and U I&D 329/94 (FEDER participation) and by a PhD grant from FCT (SFRH/BD/8320/2002). Nota prévia Na elaboração da presente dissertação foram integralmente utilizados artigos já publicados em revistas internacionais indexadas, à excepção de apenas um artigo que se encontra submetido para publicação (ver “List of Papers”). Uma vez que os referidos trabalhos foram realizados em colaboração com outros investigadores, e nos termos do Artigo 15º, Capítulo I, do Regulamento de Doutoramento da Universidade de Lisboa publicado no Diário da República – II série Nº 194 de 19-08-1993, a autora esclarece que participou em todas as fases de elaboração dos artigos: planeamento, obtenção e análise de dados, interpretação dos resultados e redacção do manuscrito final. -

Introduction of Alien Fish Species to Waterbodies of Uzbekistan
International Journal of Science and Research (IJSR) ISSN (Online): 2319-7064 Index Copernicus Value (2016): 79.57 | Impact Factor (2015): 6.391 Introduction of Alien Fish Species to Waterbodies of Uzbekistan M. A. Yuldashov Institute of Fish Culture, 2, Universitet St., Kibray district, Tashkent region, Uzbekistan Abstract: Totally 47 alien fish species were introduced to Uzbekistan including 23 as target and 24 as accidental. One time introduction was carried out for health care; all other were done for fisheries purposes: (i) to the Aral Sea, (ii) to newly created irrigation water bodies, (iii) to aquaculture. Secondary settlement was marked for 27 species, for 20 species settlement was not found. Natural reproduction was marked for 31 species. At present 22 introduced species vanished in region; 7 species are not numerous, 20 species are numerous in water bodies. List of high value marketable fish includes 6 species, commercial – 18, trash fishes – 23 species. A number of introduced species did not find conditions for reproduction or could not survive and vanished. Several species survived and began to reproduce in the Aral Sea but vanished as salinity of water increased. Some species survived, but did not spread to another waterbodies. Several alien species have widely spread all over the Aral Sea basin; that’s happened after introduction of far eastern fishes in early 1960s. Keywords: introduction of alien fishes to Uzbekistan 1. Introduction stellatus vanished in the Aral Sea because of increasing of water salinity. Water bodies of Uzbekistan are part of inland basin of the 3) Acipenser baerii (siberian sturgeon) is native to Siberian Aral Sea, Central Asia. -

André Phillips Phd Thesis
THE MECHANISMS AND CONSEQUENCES OF OVIPOSITION DECISIONS IN THE EUROPEAN BITTERLING André Phillips A Thesis Submitted for the Degree of PhD at the University of St Andrews 2018 Full metadata for this item is available in St Andrews Research Repository at: http://research-repository.st-andrews.ac.uk/ Please use this identifier to cite or link to this item: http://hdl.handle.net/10023/15524 This item is protected by original copyright The mechanisms and consequences of oviposition decisions in the European bitterling André Phillips 1. Candidate’s declarations: I, André Phillips hereby certify that this thesis, which is approximately 38,000 words in length, has been written by me, and that it is the record of work carried out by me, or principally by myself in collaboration with others as acknowledged, and that it has not been submitted in any previous application for a higher degree. I was admitted as a research student in October, 2014 and as a candidate for the degree of PhD in October, 2014; the higher study for which this is a record was carried out in the University of St Andrews between 2014 and 2017. I, André Phillips, received assistance in the writing of this thesis in respect of grammar & spelling, which was provided by Helen Spence-Jones Date …… signature of candidate ……… 2. Supervisor’s declaration: I hereby certify that the candidate has fulfilled the conditions of the Resolution and Regulations appropriate for the degree of ……… in the University of St Andrews and that the candidate is qualified to submit this thesis in application for that degree. -

Energetic Costs in the Relationship Between Bitterling and Mussels in East Asia
1 Energetic costs in the relationship between bitterling and mussels in East Asia 2 3 Caroline Methling1, Karel Douda2, Huanzhang Liu3, Romain Rouchet1, Veronika Bartáková1, 4 Dan Yu3, Carl Smith1,4,5, Martin Reichard1* 5 6 1The Czech Academy of Sciences, Institute of Vertebrate Biology, Brno, Czech Republic 7 2Department of Zoology and Fisheries, Czech University of Life Sciences Prague, Prague, Czech 8 Republic 9 3The Key Lab of Aquatic Biodiversity and Conservation, Institute of Hydrobiology, Chinese 10 Academy of Sciences, Wuhan, China 11 4Department of Ecology and Vertebrate Zoology, University of Łódź, Łódź, Poland 12 5School of Biology and Bell-Pettigrew Museum of Natural History, University of St Andrews, St 13 Andrews, UK 14 15 *Corresponding author: Martin Reichard, Institute of Vertebrate Biology, Academy of Sciences 16 of the Czech Republic, Květná 8, 603 65 Brno, Czech Republic; E-mail: [email protected] 17 18 Running title: Energetic costs in bitterling-mussel relationship 1 19 Abstract 20 Bitterling fishes and unionid mussels are involved in a two-sided coevolutionary association. On 21 one side, bitterling exploit unionids by ovipositing in their gills. On the other side, unionids develop 22 via a larval stage (glochidium) that attach to fish gills. Both interactions are parasitic and expected 23 to have negative consequences for the host. Here we examined the effects of this association on the 24 metabolic rates of mussel and fish hosts by measuring oxygen uptake rates (MO2). Measurements 25 were performed on two widespread and broadly coexisting species; the rose bitterling Rhodeus 26 ocellatus and Chinese pond mussel Sinanodonta woodiana. -

Of the BC Wildlife Act *Note: This List Is to Be Used As a Guidance Document and Therefore May Not Include All Species
Species Lister Under Schedule 3 of the Controlled Alien Species Regulation of the BC Wildlife Act *Note: This list is to be used as a guidance document and therefore may not include all species. Group Common name Family Genus Species (1) Channidae Chel Snakehead Channidae Channa amphibeus Northern Snakehead Channidae Channa argus Chinese Snakehead, Chinafish Channidae Channa asiatica Orangespotted Snakehead Channidae Channa aurantimaculata Bangka Snakehead Channidae Channa bankanensis Baram Snakehead Channidae Channa baramensis Barca Snakehead Channidae Channa barca Rainbow Snakehead Channidae Channa bleheri Burmese Snakehead Channidae Channa burmanica Bluespotted Snakehead Channidae Channa cyanospilos Malabar Snakehead Channidae Channa diplogramma Dwarf Snakehead Channidae Channa gachua Inle Snakehead Channidae Channa harcourtbutleri NA Channidae Channa hoaluensis NA Channidae Channa longistomata Splendid Snakehead Channidae Channa lucius Blotched Snakehead, Snakehead Mullet Channidae Channa maculata Emperor Snakehead Channidae Channa marulioides Bullseye Snakehead Channidae Channa marulius Blackfinned Snakehead Channidae Channa melanoptera NA Channidae Channa melanostigma Black Snakehead, Manu Channidae Channa melasoma Giant Snakehead, Red Snakehead Channidae Channa micropeltes NA Channidae Channa ninhbinhensis Night Snakehead Channidae Channa nox Ceylon Snakehead, Walking Snakehead, Smooth-breasted Snakefish Channidae Channa orientalis NA Channidae Channa ornatipinnis Panaw Snakehead Channidae Channa panaw Ocellated Snakehead Channidae -

Reprodukční Chování a Pohlavní Výběr U Hořavek (Acheilognathinae) S Důrazem Na Hořavku Duhovou (Rhodeus Amarus)
MASARYKOVA UNIVERZITA V BRNĚ PŘÍRODOVĚDECKÁ FAKULTA ÚSTAV BOTANIKY A ZOOLOGIE Reprodukční chování a pohlavní výběr u hořavek (Acheilognathinae) s důrazem na hořavku duhovou (Rhodeus amarus) Bakalářská práce Brno 2008 Autor: Radomil Řežucha MASARYKOVA UNIVERZITA V BRNĚ PŘÍRODOVĚDECKÁ FAKULTA ÚSTAV BOTANIKY A ZOOLOGIE Reprodukční chování a pohlavní výběr u hořavek (Acheilognathinae) s důrazem na hořavku duhovou (Rhodeus amarus) Bakalářská práce Autor: Radomil Řežucha Brno 2008 Vedoucí BP: RNDr. Martin Reichard, PhD. Prohlášení Souhlasím s uložením této bakalářské práce v knihovně Ústavu botaniky a zoologie PřF MU v Brně, případně v jiné knihovně MU, s jejím veřejným půjčováním a využitím pro vě- decké, vzdělávací nebo jiné veřejně prospěšné účely, a to za předpokladu, že převzaté infor- mace budou řádně citovány a nebudou využívány komerčně. Datum …………………….. ……………………………. Radomil Řežucha Poděkování Chtěl bych poděkovat RNDr. Martinu Reichardovi, Phd. za pomoc a podporu při zpraco- vání této práce. Abstrakt Cílem této práce je shrnout poznatky o reprodukčním chování hořavek (Acheilognathi- nae), hlavně pak hořavky duhové (Rhodeus amarus), a charakterizovat mechanizmy pohlav- ního výběru, na něj navazující párovací systémy a alternativní párovací taktiky. Pohlavní výběr, tedy ústřední téma této práce, zvýhodňuje určitého jedince před ostatní- mi jedinci stejného pohlaví a dělí se na intrasexuální a intersexuální. Intersexuální pohlavní výběr podléhá samičím preferencím, které se dají vysvětlit Fisherovou hypotézou, hypotézou dobrých genů či využitím smyslových vjemů. Z evolučního hlediska je pro samičky nezbytné maximalizovat svůj přímý a nepřímý zisk. Přímý zisk ovlivňuje samičku samotnou a může být ve formě rodičovské péče, svatebních darů, většího počtu oplodněných vajíček či kvalit- ního místa pro rozmnožování. Nepřímý zisk ovlivňuje potomstvo samičky prostřednictvím kvalitních genů partnera a samička se pro jeho maximalizaci může řídit množstvím signálů samečkovy kvality, např. -

Energetic Costs in the Relationship Between Bitterling and Mussels in East Asia
applyparastyle "body/p[1]" parastyle "Text_First" Biological Journal of the Linnean Society, 2018, XX, 1–10. With 3 figures. Downloaded from https://academic.oup.com/biolinnean/advance-article-abstract/doi/10.1093/biolinnean/bly178/5164100 by Ustav biologie obratlovcu AV CR, v.v.i. user on 08 November 2018 Energetic costs in the relationship between bitterling and mussels in East Asia CAROLINE METHLING1, KAREL DOUDA2, HUANZHANG LIU3, ROMAIN ROUCHET1, VERONIKA BARTÁKOVÁ1, DAN YU3, CARL SMITH1,4,5,6 and MARTIN REICHARD1,* 1The Czech Academy of Sciences, Institute of Vertebrate Biology, Brno, Czech Republic 2Department of Zoology and Fisheries, Czech University of Life Sciences Prague, Prague, Czech Republic 3The Key Laboratory of Aquatic Biodiversity and Conservation, Institute of Hydrobiology, Chinese Academy of Sciences, Wuhan, China 4Department of Ecology and Vertebrate Zoology, University of Łódź, Łódź, Poland 5School of Biology, University of St Andrews, St Andrews, UK 6Bell Pettigrew Museum of Natural History, University of St Andrews, St Andrews, UK Received 31 July 2018; revised 11 October 2018; accepted for publication 11 October 2018 Bitterling fishes and unionid mussels are involved in a two-sided co-evolutionary association. On the one side, bitterling exploit unionids by ovipositing in their gills. On the other side, unionids develop via a larval stage (glo- chidium) that attaches to fish gills. Both interactions are parasitic and expected to have negative consequences for the host. Here, we examine the effects of this association on the metabolic rates of mussel and fish hosts by measur- ing oxygen uptake rates (MO2). Measurements were performed on two widespread and broadly coexisting species, namely the rose bitterling Rhodeus ocellatus and Chinese pond mussel Sinanodonta woodiana.