Small Bristles, the Drosophila Ortholog of NXF-1, Is Essential for Mrna Export Throughout Development
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Non-Synonymous Mutations Mapped to Chromosome X Associated With
de Camargo et al. BMC Genomics (2015) 16:384 DOI 10.1186/s12864-015-1595-0 RESEARCH ARTICLE Open Access Non-synonymous mutations mapped to chromosome X associated with andrological and growth traits in beef cattle Gregório Miguel Ferreira de Camargo1,2,3, Laercio R Porto-Neto2, Matthew J Kelly3, Rowan J Bunch2, Sean M McWilliam2, Humberto Tonhati1, Sigrid A Lehnert2, Marina R S Fortes3* and Stephen S Moore4 Abstract Background: Previous genome-wide association analyses identified QTL regions in the X chromosome for percentage of normal sperm and scrotal circumference in Brahman and Tropical Composite cattle. These traits are important to be studied because they are indicators of male fertility and are correlated with female sexual precocity and reproductive longevity. The aim was to investigate candidate genes in these regions and to identify putative causative mutations that influence these traits. In addition, we tested the identified mutations for female fertility and growth traits. Results: Using a combination of bioinformatics and molecular assay technology, twelve non-synonymous SNPs in eleven genes were genotyped in a cattle population. Three and nine SNPs explained more than 1% of the additive genetic variance for percentage of normal sperm and scrotal circumference, respectively. The SNPs that had a major influence in percentage of normal sperm were mapped to LOC100138021 and TAF7L genes; and in TEX11 and AR genes for scrotal circumference. One SNP in TEX11 was explained ~13% of the additive genetic variance for scrotal circumference at 12 months. The tested SNP were also associated with weight measurements, but not with female fertility traits. -

Implication of PIWI Interacting Rnas Karine Casier, Antoine Boivin, Clément Carré, Laure Teysset
Environmentally-Induced Transgenerational Epigenetic Inheritance: Implication of PIWI Interacting RNAs Karine Casier, Antoine Boivin, Clément Carré, Laure Teysset To cite this version: Karine Casier, Antoine Boivin, Clément Carré, Laure Teysset. Environmentally-Induced Transgen- erational Epigenetic Inheritance: Implication of PIWI Interacting RNAs. Cells, MDPI, 2019, 8 (9), pp.1108. 10.3390/cells8091108. hal-02344602 HAL Id: hal-02344602 https://hal.sorbonne-universite.fr/hal-02344602 Submitted on 4 Nov 2019 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. cells Review Environmentally-Induced Transgenerational Epigenetic Inheritance: Implication of PIWI Interacting RNAs Karine Casier , Antoine Boivin , Clément Carré and Laure Teysset * Transgenerational Epigenetics & small RNA Biology, Sorbonne Université, CNRS, Laboratoire Biologie du Développement, Institut de Biologie Paris-Seine, UMR7622, 75005 Paris, France; [email protected] (K.C.); [email protected] (A.B.); [email protected] (C.C.) * Correspondence: [email protected] Received: 16 August 2019; Accepted: 17 September 2019; Published: 19 September 2019 Abstract: Environmentally-induced transgenerational epigenetic inheritance is an emerging field. The understanding of associated epigenetic mechanisms is currently in progress with open questions still remaining. In this review, we present an overview of the knowledge of environmentally-induced transgenerational inheritance and associated epigenetic mechanisms, mainly in animals. -

Genome-Wide Mapping in a House Mouse Hybrid Zone Reveals Hybrid Sterility Loci and Dobzhansky-Muller Interactions Leslie M Turner1,2, Bettina Harr1*
RESEARCH ARTICLE elifesciences.org Genome-wide mapping in a house mouse hybrid zone reveals hybrid sterility loci and Dobzhansky-Muller interactions Leslie M Turner1,2, Bettina Harr1* 1Department of Evolutionary Genetics, Max Planck Institute for Evolutionary Biology, Plön, Germany; 2Laboratory of Genetics, University of Wisconsin, Madison, United States Abstract Mapping hybrid defects in contact zones between incipient species can identify genomic regions contributing to reproductive isolation and reveal genetic mechanisms of speciation. The house mouse features a rare combination of sophisticated genetic tools and natural hybrid zones between subspecies. Male hybrids often show reduced fertility, a common reproductive barrier between incipient species. Laboratory crosses have identified sterility loci, but each encompasses hundreds of genes. We map genetic determinants of testis weight and testis gene expression using offspring of mice captured in a hybrid zone between M. musculus musculus and M. m. domesticus. Many generations of admixture enables high-resolution mapping of loci contributing to these sterility-related phenotypes. We identify complex interactions among sterility loci, suggesting multiple, non-independent genetic incompatibilities contribute to barriers to gene flow in the hybrid zone. DOI: 10.7554/eLife.02504.001 Introduction *For correspondence: harr@ New species arise when reproductive barriers form, preventing gene flow between populations evolbio.mpg.de (Coyne and Orr, 2004). Recently, two approaches have substantially -

The Role of Lissencephaly-1 Protein in Male Germ Cell Differentiation
THE ROLE OF LISSENCEPHALY-1 PROTEIN IN MALE GERM CELL DIFFERENTIATION Dissertation zur Erlangung des Doktorgrades der Mathematisch-Naturwissenschaftlichen Fakultäten der Georg-August-Universität zu Göttingen vorgelegt von Nadja Drusenheimer aus Remscheid Göttingen 2009 D 7 Referent: Prof. Dr. med Dr. h.c. W. Engel Korreferentin: Prof. Dr. S. Hoyer-Fender Tag der mündlichen Prüfung: Table of Contents I TABLE OF CONTENTS TABLE OF CONTENTS ----------------------------------------------------------------------- I ABBREVIATIONS ----------------------------------------------------------------------------- V 1. INTRODUCTION ----------------------------------------------------------------------------1 1.1 Pafah1b1/Lis1 is an evolutionary conserved gene---------------------------------------------------------- 1 1.2 Mutations in LIS1 gene cause classical lissencephaly ----------------------------------------------------- 2 1.3 Expression and function of Lis1 ------------------------------------------------------------------------------- 3 1.4 Generation of the gene trap line L39GT/GT ------------------------------------------------------------------- 6 1.5 Analysis of the gene trap line L39GT/GT----------------------------------------------------------------------- 7 1.6 Objectives of this study------------------------------------------------------------------------------------------ 9 2. MATERIALS AND METHODS --------------------------------------------------------- 10 2.1 Materials ----------------------------------------------------------------------------------------------------------10 -

Comprehensive Analysis Reveals Novel Gene Signature in Head and Neck Squamous Cell Carcinoma: Predicting Is Associated with Poor Prognosis in Patients
5892 Original Article Comprehensive analysis reveals novel gene signature in head and neck squamous cell carcinoma: predicting is associated with poor prognosis in patients Yixin Sun1,2#, Quan Zhang1,2#, Lanlin Yao2#, Shuai Wang3, Zhiming Zhang1,2 1Department of Breast Surgery, The First Affiliated Hospital of Xiamen University, School of Medicine, Xiamen University, Xiamen, China; 2School of Medicine, Xiamen University, Xiamen, China; 3State Key Laboratory of Cellular Stress Biology, School of Life Sciences, Xiamen University, Xiamen, China Contributions: (I) Conception and design: Y Sun, Q Zhang; (II) Administrative support: Z Zhang; (III) Provision of study materials or patients: Y Sun, Q Zhang; (IV) Collection and assembly of data: Y Sun, L Yao; (V) Data analysis and interpretation: Y Sun, S Wang; (VI) Manuscript writing: All authors; (VII) Final approval of manuscript: All authors. #These authors contributed equally to this work. Correspondence to: Zhiming Zhang. Department of Surgery, The First Affiliated Hospital of Xiamen University, Xiamen, China. Email: [email protected]. Background: Head and neck squamous cell carcinoma (HNSC) remains an important public health problem, with classic risk factors being smoking and excessive alcohol consumption and usually has a poor prognosis. Therefore, it is important to explore the underlying mechanisms of tumorigenesis and screen the genes and pathways identified from such studies and their role in pathogenesis. The purpose of this study was to identify genes or signal pathways associated with the development of HNSC. Methods: In this study, we downloaded gene expression profiles of GSE53819 from the Gene Expression Omnibus (GEO) database, including 18 HNSC tissues and 18 normal tissues. -

FMR1 Antibody (N-Term) Affinity Purified Rabbit Polyclonal Antibody (Pab) Catalog # AP6879A
10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 FMR1 Antibody (N-term) Affinity Purified Rabbit Polyclonal Antibody (Pab) Catalog # AP6879A Specification FMR1 Antibody (N-term) - Product Information Application WB, IHC-P, FC,E Primary Accession Q06787 Reactivity Human, Mouse Host Rabbit Clonality Polyclonal Isotype Rabbit Ig Antigen Region 20-48 FMR1 Antibody (N-term) - Additional Information Gene ID 2332 Other Names Western blot analysis of FMR1 Antibody Fragile X mental retardation protein 1, (N-term) (Cat. #AP6879a) in mouse lung FMRP, Protein FMR-1, FMR1 tissue lysates (35ug/lane). FMR1 (arrow) was detected using the purified Pab. Target/Specificity This FMR1 antibody is generated from rabbits immunized with a KLH conjugated synthetic peptide between 20-48 amino acids from the N-terminal region of human FMR1. Dilution WB~~1:1000 IHC-P~~1:50~100 FC~~1:10~50 Format Purified polyclonal antibody supplied in PBS with 0.09% (W/V) sodium azide. This antibody is purified through a protein A Formalin-fixed and paraffin-embedded column, followed by peptide affinity human brain tissue reacted with FMR1 purification. Antibody (N-term), which was peroxidase-conjugated to the secondary Storage antibody, followed by DAB staining. This data Maintain refrigerated at 2-8°C for up to 2 demonstrates the use of this antibody for weeks. For long term storage store at -20°C immunohistochemistry; clinical relevance has in small aliquots to prevent freeze-thaw not been evaluated. cycles. Precautions FMR1 Antibody (N-term) is for research use only and not for use in diagnostic or therapeutic procedures. -

The Pdx1 Bound Swi/Snf Chromatin Remodeling Complex Regulates Pancreatic Progenitor Cell Proliferation and Mature Islet Β Cell
Page 1 of 125 Diabetes The Pdx1 bound Swi/Snf chromatin remodeling complex regulates pancreatic progenitor cell proliferation and mature islet β cell function Jason M. Spaeth1,2, Jin-Hua Liu1, Daniel Peters3, Min Guo1, Anna B. Osipovich1, Fardin Mohammadi3, Nilotpal Roy4, Anil Bhushan4, Mark A. Magnuson1, Matthias Hebrok4, Christopher V. E. Wright3, Roland Stein1,5 1 Department of Molecular Physiology and Biophysics, Vanderbilt University, Nashville, TN 2 Present address: Department of Pediatrics, Indiana University School of Medicine, Indianapolis, IN 3 Department of Cell and Developmental Biology, Vanderbilt University, Nashville, TN 4 Diabetes Center, Department of Medicine, UCSF, San Francisco, California 5 Corresponding author: [email protected]; (615)322-7026 1 Diabetes Publish Ahead of Print, published online June 14, 2019 Diabetes Page 2 of 125 Abstract Transcription factors positively and/or negatively impact gene expression by recruiting coregulatory factors, which interact through protein-protein binding. Here we demonstrate that mouse pancreas size and islet β cell function are controlled by the ATP-dependent Swi/Snf chromatin remodeling coregulatory complex that physically associates with Pdx1, a diabetes- linked transcription factor essential to pancreatic morphogenesis and adult islet-cell function and maintenance. Early embryonic deletion of just the Swi/Snf Brg1 ATPase subunit reduced multipotent pancreatic progenitor cell proliferation and resulted in pancreas hypoplasia. In contrast, removal of both Swi/Snf ATPase subunits, Brg1 and Brm, was necessary to compromise adult islet β cell activity, which included whole animal glucose intolerance, hyperglycemia and impaired insulin secretion. Notably, lineage-tracing analysis revealed Swi/Snf-deficient β cells lost the ability to produce the mRNAs for insulin and other key metabolic genes without effecting the expression of many essential islet-enriched transcription factors. -

Telomere-To-Telomere Assembly of a Complete Human X Chromosome
Article Telomere-to-telomere assembly of a complete human X chromosome https://doi.org/10.1038/s41586-020-2547-7 Karen H. Miga1,24 ✉, Sergey Koren2,24, Arang Rhie2, Mitchell R. Vollger3, Ariel Gershman4, Andrey Bzikadze5, Shelise Brooks6, Edmund Howe7, David Porubsky3, Glennis A. Logsdon3, Received: 30 July 2019 Valerie A. Schneider8, Tamara Potapova7, Jonathan Wood9, William Chow9, Joel Armstrong1, Accepted: 29 May 2020 Jeanne Fredrickson10, Evgenia Pak11, Kristof Tigyi1, Milinn Kremitzki12, Christopher Markovic12, Valerie Maduro13, Amalia Dutra11, Gerard G. Bouffard6, Alexander M. Chang2, Published online: 14 July 2020 Nancy F. Hansen14, Amy B. Wilfert3, Françoise Thibaud-Nissen8, Anthony D. Schmitt15, Open access Jon-Matthew Belton15, Siddarth Selvaraj15, Megan Y. Dennis16, Daniela C. Soto16, Ruta Sahasrabudhe17, Gulhan Kaya16, Josh Quick18, Nicholas J. Loman18, Nadine Holmes19, Check for updates Matthew Loose19, Urvashi Surti20, Rosa ana Risques10, Tina A. Graves Lindsay12, Robert Fulton12, Ira Hall12, Benedict Paten1, Kerstin Howe9, Winston Timp4, Alice Young6, James C. Mullikin6, Pavel A. Pevzner21, Jennifer L. Gerton7, Beth A. Sullivan22, Evan E. Eichler3,23 & Adam M. Phillippy2 ✉ After two decades of improvements, the current human reference genome (GRCh38) is the most accurate and complete vertebrate genome ever produced. However, no single chromosome has been fnished end to end, and hundreds of unresolved gaps persist1,2. Here we present a human genome assembly that surpasses the continuity of GRCh382, along with a gapless, telomere-to-telomere assembly of a human chromosome. This was enabled by high-coverage, ultra-long-read nanopore sequencing of the complete hydatidiform mole CHM13 genome, combined with complementary technologies for quality improvement and validation. -
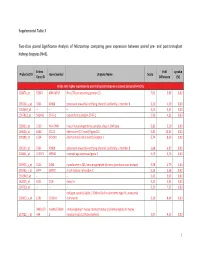
Supplemental Table 3 Two-Class Paired Significance Analysis of Microarrays Comparing Gene Expression Between Paired
Supplemental Table 3 Two‐class paired Significance Analysis of Microarrays comparing gene expression between paired pre‐ and post‐transplant kidneys biopsies (N=8). Entrez Fold q‐value Probe Set ID Gene Symbol Unigene Name Score Gene ID Difference (%) Probe sets higher expressed in post‐transplant biopsies in paired analysis (N=1871) 218870_at 55843 ARHGAP15 Rho GTPase activating protein 15 7,01 3,99 0,00 205304_s_at 3764 KCNJ8 potassium inwardly‐rectifying channel, subfamily J, member 8 6,30 4,50 0,00 1563649_at ‐‐ ‐‐ ‐‐ 6,24 3,51 0,00 1567913_at 541466 CT45‐1 cancer/testis antigen CT45‐1 5,90 4,21 0,00 203932_at 3109 HLA‐DMB major histocompatibility complex, class II, DM beta 5,83 3,20 0,00 204606_at 6366 CCL21 chemokine (C‐C motif) ligand 21 5,82 10,42 0,00 205898_at 1524 CX3CR1 chemokine (C‐X3‐C motif) receptor 1 5,74 8,50 0,00 205303_at 3764 KCNJ8 potassium inwardly‐rectifying channel, subfamily J, member 8 5,68 6,87 0,00 226841_at 219972 MPEG1 macrophage expressed gene 1 5,59 3,76 0,00 203923_s_at 1536 CYBB cytochrome b‐245, beta polypeptide (chronic granulomatous disease) 5,58 4,70 0,00 210135_s_at 6474 SHOX2 short stature homeobox 2 5,53 5,58 0,00 1562642_at ‐‐ ‐‐ ‐‐ 5,42 5,03 0,00 242605_at 1634 DCN decorin 5,23 3,92 0,00 228750_at ‐‐ ‐‐ ‐‐ 5,21 7,22 0,00 collagen, type III, alpha 1 (Ehlers‐Danlos syndrome type IV, autosomal 201852_x_at 1281 COL3A1 dominant) 5,10 8,46 0,00 3493///3 IGHA1///IGHA immunoglobulin heavy constant alpha 1///immunoglobulin heavy 217022_s_at 494 2 constant alpha 2 (A2m marker) 5,07 9,53 0,00 1 202311_s_at -

Human Leucine-Rich Repeat Proteins: a Genome-Wide Bioinformatic Categorization and Functional Analysis in Innate Immunity
Human leucine-rich repeat proteins: a genome-wide bioinformatic categorization and functional analysis in innate immunity Aylwin C. Y. Nga,b,1, Jason M. Eisenberga,b,1, Robert J. W. Heatha, Alan Huetta, Cory M. Robinsonc, Gerard J. Nauc, and Ramnik J. Xaviera,b,2 aCenter for Computational and Integrative Biology, and Gastrointestinal Unit, Massachusetts General Hospital and Harvard Medical School, Boston, MA 02114; bThe Broad Institute of Massachusetts Institute of Technology and Harvard, Cambridge, MA 02142; and cMicrobiology and Molecular Genetics, University of Pittsburgh School of Medicine, Pittsburgh, PA 15261 Edited by Jeffrey I. Gordon, Washington University School of Medicine, St. Louis, MO, and approved June 11, 2010 (received for review February 17, 2010) In innate immune sensing, the detection of pathogen-associated proteins have been implicated in human diseases to date, notably molecular patterns by recognition receptors typically involve polymorphisms in NOD2 in Crohn disease (8, 9), CIITA in leucine-rich repeats (LRRs). We provide a categorization of 375 rheumatoid arthritis and multiple sclerosis (10), and TLR5 in human LRR-containing proteins, almost half of which lack other Legionnaire disease (11). identifiable functional domains. We clustered human LRR proteins Most LRR domains consist of a chain of between 2 and 45 by first assigning LRRs to LRR classes and then grouping the proteins LRRs (12). Each repeat in turn is typically 20 to 30 residues long based on these class assignments, revealing several of the resulting and can be divided into a highly conserved segment (HCS) fol- protein groups containing a large number of proteins with certain lowed by a variable segment (VS). -

The Drosophila TART Transposon Manipulates the Pirna Pathway As a Counter-Defense 2 Strategy to Limit Host Silencing 3 4 Christopher E
bioRxiv preprint doi: https://doi.org/10.1101/2020.02.20.957324; this version posted February 20, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 1 The Drosophila TART transposon manipulates the piRNA pathway as a counter-defense 2 strategy to limit host silencing 3 4 Christopher E. Ellison, Meenakshi S. Kagda*, Weihuan Cao 5 Department of Genetics, Human Genetics Institute of New Jersey, Rutgers, The State University of 6 New Jersey, Piscataway, New Jersey 7 8 *Current Affiliation: Department of Genetics, Stanford University, Stanford, California 9 10 Corresponding author: 11 Christopher E. Ellison 12 [email protected] 13 14 15 Short title: Drosophila transposon counter-defense 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.02.20.957324; this version posted February 20, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 1 Abstract 2 Co-evolution between transposable elements (TEs) and their hosts can be antagonistic, where TEs 3 evolve to avoid silencing and the host responds by reestablishing TE suppression, or mutualistic, where 4 TEs are co-opted to benefit their host. -
Identification of RNA-Binding Proteins As Targetable Putative Oncogenes
International Journal of Molecular Sciences Article Identification of RNA-Binding Proteins as Targetable Putative Oncogenes in Neuroblastoma Jessica L. Bell 1,2,*, Sven Hagemann 1, Jessica K. Holien 3,4 , Tao Liu 2, Zsuzsanna Nagy 2,5, Johannes H. Schulte 6,7, Danny Misiak 1 and Stefan Hüttelmaier 1,* 1 Institute of Molecular Medicine, Sect. Molecular Cell Biology, Martin Luther University Halle-Wittenberg, Charles Tanford Protein Center, 06120 Halle Saale, Germany; [email protected] (S.H.); [email protected] (D.M.) 2 Children’s Cancer Institute Australia, Randwick, NSW 2031, Australia; [email protected] (T.L.); [email protected] (Z.N.) 3 St. Vincent’s Institute of Medical Research, Fitzroy, Victoria 3065, Australia; [email protected] 4 Biosciences and Food Technology, School of Science, College of Science, Engineering and Health, RMIT University, Melbourne, Victoria 3053, Australia 5 School of Women’s & Children’s Health, UNSW Sydney, Randwick, NSW 2031, Australia 6 Department of Pediatric Oncology/Hematology, Charité-Universitätsmedizin Berlin, 10117 Berlin, Germany; [email protected] 7 German Consortium for Translational Cancer Research (DKTK), Partner Site Charité Berlin, 10117 Berlin, Germany * Correspondence: [email protected] (J.L.B.); [email protected] (S.H.) Received: 23 April 2020; Accepted: 14 July 2020; Published: 19 July 2020 Abstract: Neuroblastoma is a common childhood cancer with almost a third of those affected still dying, thus new therapeutic strategies need to be explored. Current experimental therapies focus mostly on inhibiting oncogenic transcription factor signalling. Although LIN28B, DICER and other RNA-binding proteins (RBPs) have reported roles in neuroblastoma development and patient outcome, the role of RBPs in neuroblastoma is relatively unstudied.