Stromule Extension Along Microtubules Coordinated with Actin
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
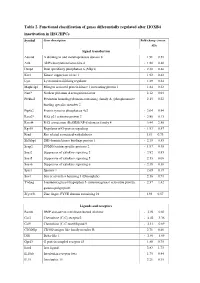
Table 2. Functional Classification of Genes Differentially Regulated After HOXB4 Inactivation in HSC/Hpcs
Table 2. Functional classification of genes differentially regulated after HOXB4 inactivation in HSC/HPCs Symbol Gene description Fold-change (mean ± SD) Signal transduction Adam8 A disintegrin and metalloprotease domain 8 1.91 ± 0.51 Arl4 ADP-ribosylation factor-like 4 - 1.80 ± 0.40 Dusp6 Dual specificity phosphatase 6 (Mkp3) - 2.30 ± 0.46 Ksr1 Kinase suppressor of ras 1 1.92 ± 0.42 Lyst Lysosomal trafficking regulator 1.89 ± 0.34 Mapk1ip1 Mitogen activated protein kinase 1 interacting protein 1 1.84 ± 0.22 Narf* Nuclear prelamin A recognition factor 2.12 ± 0.04 Plekha2 Pleckstrin homology domain-containing. family A. (phosphoinosite 2.15 ± 0.22 binding specific) member 2 Ptp4a2 Protein tyrosine phosphatase 4a2 - 2.04 ± 0.94 Rasa2* RAS p21 activator protein 2 - 2.80 ± 0.13 Rassf4 RAS association (RalGDS/AF-6) domain family 4 3.44 ± 2.56 Rgs18 Regulator of G-protein signaling - 1.93 ± 0.57 Rrad Ras-related associated with diabetes 1.81 ± 0.73 Sh3kbp1 SH3 domain kinase bindings protein 1 - 2.19 ± 0.53 Senp2 SUMO/sentrin specific protease 2 - 1.97 ± 0.49 Socs2 Suppressor of cytokine signaling 2 - 2.82 ± 0.85 Socs5 Suppressor of cytokine signaling 5 2.13 ± 0.08 Socs6 Suppressor of cytokine signaling 6 - 2.18 ± 0.38 Spry1 Sprouty 1 - 2.69 ± 0.19 Sos1 Son of sevenless homolog 1 (Drosophila) 2.16 ± 0.71 Ywhag 3-monooxygenase/tryptophan 5- monooxygenase activation protein. - 2.37 ± 1.42 gamma polypeptide Zfyve21 Zinc finger. FYVE domain containing 21 1.93 ± 0.57 Ligands and receptors Bambi BMP and activin membrane-bound inhibitor - 2.94 ± 0.62 -

Cisplatin and Phenanthriplatin Modulate Long-Noncoding
www.nature.com/scientificreports OPEN Cisplatin and phenanthriplatin modulate long‑noncoding RNA expression in A549 and IMR90 cells revealing regulation of microRNAs, Wnt/β‑catenin and TGF‑β signaling Jerry D. Monroe1,2, Satya A. Moolani2,3, Elvin N. Irihamye2,4, Katheryn E. Lett1, Michael D. Hebert1, Yann Gibert1* & Michael E. Smith2* The monofunctional platinum(II) complex, phenanthriplatin, acts by blocking transcription, but its regulatory efects on long‑noncoding RNAs (lncRNAs) have not been elucidated relative to traditional platinum‑based chemotherapeutics, e.g., cisplatin. Here, we treated A549 non‑small cell lung cancer and IMR90 lung fbroblast cells for 24 h with either cisplatin, phenanthriplatin or a solvent control, and then performed microarray analysis to identify regulated lncRNAs. RNA22 v2 microRNA software was subsequently used to identify microRNAs (miRNAs) that might be suppressed by the most regulated lncRNAs. We found that miR‑25‑5p, ‑30a‑3p, ‑138‑5p, ‑149‑3p, ‑185‑5p, ‑378j, ‑608, ‑650, ‑708‑5p, ‑1253, ‑1254, ‑4458, and ‑4516, were predicted to target the cisplatin upregulated lncRNAs, IMMP2L‑1, CBR3‑1 and ATAD2B‑5, and the phenanthriplatin downregulated lncRNAs, AGO2‑1, COX7A1‑2 and SLC26A3‑1. Then, we used qRT‑PCR to measure the expression of miR‑25‑5p, ‑378j, ‑4516 (A549) and miR‑149‑3p, ‑608, and ‑4458 (IMR90) to identify distinct signaling efects associated with cisplatin and phenanthriplatin. The signaling pathways associated with these miRNAs suggests that phenanthriplatin may modulate Wnt/β‑catenin and TGF‑β signaling through the MAPK/ ERK and PTEN/AKT pathways diferently than cisplatin. Further, as some of these miRNAs may be subject to dissimilar lncRNA targeting in A549 and IMR90 cells, the monofunctional complex may not cause toxicity in normal lung compared to cancer cells by acting through distinct lncRNA and miRNA networks. -

In Acute Myeloid Leukemia
7283 Original Article Comprehensively analyze the expression and prognostic role for ten-eleven translocations (TETs) in acute myeloid leukemia Yan Huang1#, Jie Wei1#, Xunjun Huang1, Weijie Zhou1, Yuling Xu2, Dong-Hong Deng2, Peng Cheng2 1Department of Hematology and Rheumatology, People’s Hospital of Baise, Baise, China; 2Department of Hematology, the First Affiliated Hospital of Guangxi Medical University, Nanning, China Contributions: (I) Conception and design: J Wei, Y Huang; (II) Administrative support: X Huang, Y Xu; (III) Provision of study materials or patients: W Zhou; (IV) Collection and assembly of data: J Wei, Y Huang; (V) Data analysis and interpretation: DH Deng, P Cheng; (VI) Manuscript writing: All authors; (VII) Final approval of manuscript: All authors. #These authors contributed equally to this work. Correspondence to: Yuling Xu. Department of Hematology, The First Affiliated Hospital of Guangxi Medical University, Nanning, China. Email: [email protected]; Dong-Hong Deng. Department of Hematology, The First Affiliated Hospital of Guangxi Medical University, Nanning, China. Email: [email protected]; Peng Cheng. Department of Hematology, The First Affiliated Hospital of Guangxi Medical University, Nanning 530000, China. Email: [email protected]. Background: The ten-eleven translocation (TET) family oxidize 5-methylcytosines (5mCs) and promote the locus-specific reversal of DNA. The role of TETs in acute myeloid leukemia (AML) is mostly unknown. Methods: TETs mRNA expression levels were analyzed via Gene Expression Profiling Interactive Analysis (GEPIA). The association TETs expression levels and methylation with prognosis by UALCAN GenomicScape, and METHsurv. We analyzed TETs’ aberration types, located mutations, and structures via cBioPortal. GeneMANIA performed the functional network. Gene ontology (GO) enrichment was analyzed via LinkedOmics. -

Supplementary Figures and Table Legends
Supplementary Information Supplementary Figure Legends Figure S1. CARM1 is required for estrogen-induced gene transcriptional activation. (A) Box plot showing the expression of CARM1 (FPKM) in a cohort of clinical breast cancer (n=1,102) and normal (n=113) samples from TCGA (The Cancer Genome Atlas). (B, C) Kaplan Meier survival analyses for DMFS (distal metastasis free survival) (n=1,747) and OS (overall survival) (n=1,402) of breast cancer patients using CARM1 as input. (D, E) RNA-seq as described in Figure 1A for specific genes were shown as indicated. (F) Correlation of CARM1’s effects on whole transcriptome in response to estrogen between two RNA-seq biological repeats (n=11,816). (G) Correlation of CARM1 effects on whole transcriptome in response to estrogen based on RNA- seq between two independent siRNAs targeting CARM1 (siCARM1 (1) and siCARM1 (2)) (n=12,234). (H, I) MCF7 cells as described in (G) were subjected to immunoblotting (IB) (H) and RT-qPCR (I) analysis to examine the levels of indicated protein and mRNA, respectively. Actin was served as a loading control. (J) MCF7 cells as described in Figure 1E were subjected to RT-qPCR analysis to examine the expression of genes as indicated. (K) MCF7 cells as described in Figure 1E were subjected to immunoblotting (IB) analysis to examine the protein levels of CARM1. Actin was served as a loading control. (L) Genomic DNA was extracted from CARM1 knockout cells, followed by PCR using specific primer sets spanning gRNA targeting region (boxed in blue). The resultant PCR products were subjected to Sanger sequencing as shown. -

CRISPR-Enhanced Human Adipocyte “Browning” As Cell Therapy for Metabolic Disease
bioRxiv preprint doi: https://doi.org/10.1101/2020.10.13.337923; this version posted October 13, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. CRISPR-enhanced human adipocyte “browning” as cell therapy for metabolic disease Emmanouela Tsagkaraki1,6*, Sarah Nicoloro1*, Tiffany De Souza1, Javier Solivan-Rivera1, Anand Desai1, Yuefei Shen1, Mark Kelly1, Adilson Guilherme1, Felipe Henriques1, Raed Ibraheim2, Nadia Amrani2, Kevin Luk4, Stacy Maitland4, Randall H. Friedline1, Lauren Tauer1, Xiaodi Hu1, Jason K. Kim1,3, Scot A. Wolfe4,5, Erik J. Sontheimer2,5, Silvia Corvera1**, Michael P. Czech1**. 1Program in Molecular Medicine, University of Massachusetts Medical School, Worcester, MA 01605 2RNA Therapeutics Institute, University of Massachusetts Medical School, Worcester MA 01605 3Division of Endocrinology, Metabolism and Diabetes, Department of Medicine, University of Massachusetts Medical School Worcester, MA 01605 4Department of Molecular,Cell and Cancer Biology, University of Massachusetts Medical School, Worcester MA 01605 5Li Weibo Institute for Rare Diseases Research, University of Massachusetts Medical School, Worcester, MA 01605 6University of Crete School of Medicine, Crete 71003 *These authors contributed equally **Co-corresponding authors Michael P. Czech 373 Plantation Street Program in Molecular Medicine University of Massachusetts Medical School Worcester, MA 01605 [email protected] Tel: 1 508 856 2254 Fax: 1 508 856 1617 Silvia Corvera 373 Plantation Street Program in Molecular Medicine University of Massachusetts Medical School Worcester, MA 01605 [email protected] Tel: 1 508 856 6869 Fax: 1 508 856 1617 bioRxiv preprint doi: https://doi.org/10.1101/2020.10.13.337923; this version posted October 13, 2020. -

Human Induced Pluripotent Stem Cell–Derived Podocytes Mature Into Vascularized Glomeruli Upon Experimental Transplantation
BASIC RESEARCH www.jasn.org Human Induced Pluripotent Stem Cell–Derived Podocytes Mature into Vascularized Glomeruli upon Experimental Transplantation † Sazia Sharmin,* Atsuhiro Taguchi,* Yusuke Kaku,* Yasuhiro Yoshimura,* Tomoko Ohmori,* ‡ † ‡ Tetsushi Sakuma, Masashi Mukoyama, Takashi Yamamoto, Hidetake Kurihara,§ and | Ryuichi Nishinakamura* *Department of Kidney Development, Institute of Molecular Embryology and Genetics, and †Department of Nephrology, Faculty of Life Sciences, Kumamoto University, Kumamoto, Japan; ‡Department of Mathematical and Life Sciences, Graduate School of Science, Hiroshima University, Hiroshima, Japan; §Division of Anatomy, Juntendo University School of Medicine, Tokyo, Japan; and |Japan Science and Technology Agency, CREST, Kumamoto, Japan ABSTRACT Glomerular podocytes express proteins, such as nephrin, that constitute the slit diaphragm, thereby contributing to the filtration process in the kidney. Glomerular development has been analyzed mainly in mice, whereas analysis of human kidney development has been minimal because of limited access to embryonic kidneys. We previously reported the induction of three-dimensional primordial glomeruli from human induced pluripotent stem (iPS) cells. Here, using transcription activator–like effector nuclease-mediated homologous recombination, we generated human iPS cell lines that express green fluorescent protein (GFP) in the NPHS1 locus, which encodes nephrin, and we show that GFP expression facilitated accurate visualization of nephrin-positive podocyte formation in -

Supplemental Digital Content (Sdc) Sdc, Materials
SUPPLEMENTAL DIGITAL CONTENT (SDC) SDC, MATERIALS AND METHODS Animals This study used 9-12 week old male C57BL/6 mice (Jackson Laboratory, Bar Harbor, ME). This study conformed to the National Institutes of Health guidelines and was conducted under animal protocols approved by the University of Virginia’s Institutional Animal Care and Use Committee. Murine DCD Lung Procedure Mice were anesthetized by isoflurane inhalation and euthanized by cervical dislocation followed by a 60-minute period of “no-touch” warm ischemia. Mice then underwent extended median sternotomy and midline cervical exposure followed by intubation for the initiation of mechanical ventilation at 120 strokes/minute with room air. The left atrium was vented via an atriotomy followed by infusion of the lungs with 3 mL 4°C Perfadex® solution (Vitrolife Inc., Denver, CO) supplemented with THAM Solution (Vitrolife, Kungsbacka, Sweden), estimating weight-based volume recommendations for pulmonary artery perfusion (140mL/kg) (1). The chest was then packed with ice and the trachea occluded by silk-suture tie at tidal volume (7µL/g body weight) prior to cold static preservation (CSP) for 60 minutes at 4°C. Mice were then randomized into three experimental groups: 1) CSP alone with no EVLP, 2) EVLP with Steen solution and 3) EVLP with Steen solution supplemented with the highly selective A2AR agonist, ATL1223 (30nM, Lewis and Clark Pharmaceuticals, Charlottesville, VA). Mice treated with ATL1223 during EVLP also received ATL1223 treatment (30nM) during the Perfadex flush prior to CSP whereas the EVLP group received vehicle (DMSO) during the flush. CSP lungs, which did not undergo EVLP, underwent immediate functional assessment after re-intubation as described below. -

The Expression of NRIP1 and LCOR in Endometrioid Endometrial Cancer
in vivo 35 : 2631-2640 (2021) doi:10.21873/invivo.12545 The Expression of NRIP1 and LCOR in Endometrioid Endometrial Cancer STEFANOS FLINDRIS 1, NIKOLAOS KATSOULAS 2, ANNA GOUSSIA 3, ANDREAS CHRISTOS LAZARIS 2, IORDANIS NAVROZOGLOU 1, MINAS PASCHOPOULOS 1 and IRENE THYMARA 2 1Department of Obstetrics and Gynecology, University Hospital of Ioannina, Ioannina, Greece; 2First Department of Pathology, School of Medicine, National and Kapodistrian University of Athens, Laiko General Hospital of Athens, Athens, Greece; 3Department of Pathology, University Hospital of Ioannina, Ioannina, Greece Abstract. Background: The aim of the study was to Endometrial cancer is the sixth most common cancer among analyze the expression of nuclear receptor interacting females worldwide and the fourth leading cause of cancer protein 1 (NRIP1) and its partner ligand-dependent nuclear (1). It is considered as the most common gynecological receptor co-repressor (LCOR) in endometrioid endometrial malignancy, accounting for 7% of all female cancer (1). Two cancer and to investigate their association with estrogen broad pathogenetic types of endometrial carcinoma have receptor (ER), progesterone receptor (PR), Ki-67, been recognized: Type I tumors are endometrioid carcinomas clinicopathological parameters and patient survival. and type II are non-endometrioid, including serous and clear- Materials and Methods: Immunohistochemical evaluation cell carcinomas (2). Most cases (90%) of endometrial was carried out to investigate the subcellular expression of carcinomas occur in women older than 50 years of age, with NRIP1 and LCOR in endometrioid endometrial cancer a median age of 62 years at diagnosis, and are detected in samples. Statistical analysis was used to identify the early stages [80% in International Federation of Obstetrics correlations of NRIP1 and LCOR expression with and Gynecology (FIGO) stage I] (2, 3). -

The Human Gene Connectome As a Map of Short Cuts for Morbid Allele Discovery
The human gene connectome as a map of short cuts for morbid allele discovery Yuval Itana,1, Shen-Ying Zhanga,b, Guillaume Vogta,b, Avinash Abhyankara, Melina Hermana, Patrick Nitschkec, Dror Friedd, Lluis Quintana-Murcie, Laurent Abela,b, and Jean-Laurent Casanovaa,b,f aSt. Giles Laboratory of Human Genetics of Infectious Diseases, Rockefeller Branch, The Rockefeller University, New York, NY 10065; bLaboratory of Human Genetics of Infectious Diseases, Necker Branch, Paris Descartes University, Institut National de la Santé et de la Recherche Médicale U980, Necker Medical School, 75015 Paris, France; cPlateforme Bioinformatique, Université Paris Descartes, 75116 Paris, France; dDepartment of Computer Science, Ben-Gurion University of the Negev, Beer-Sheva 84105, Israel; eUnit of Human Evolutionary Genetics, Centre National de la Recherche Scientifique, Unité de Recherche Associée 3012, Institut Pasteur, F-75015 Paris, France; and fPediatric Immunology-Hematology Unit, Necker Hospital for Sick Children, 75015 Paris, France Edited* by Bruce Beutler, University of Texas Southwestern Medical Center, Dallas, TX, and approved February 15, 2013 (received for review October 19, 2012) High-throughput genomic data reveal thousands of gene variants to detect a single mutated gene, with the other polymorphic genes per patient, and it is often difficult to determine which of these being of less interest. This goes some way to explaining why, variants underlies disease in a given individual. However, at the despite the abundance of NGS data, the discovery of disease- population level, there may be some degree of phenotypic homo- causing alleles from such data remains somewhat limited. geneity, with alterations of specific physiological pathways under- We developed the human gene connectome (HGC) to over- come this problem. -

Transcriptional and Epigenetic Control of Brown and Beige Adipose Cell Fate and Function
REVIEWS Transcriptional and epigenetic control of brown and beige adipose cell fate and function Takeshi Inagaki1,2, Juro Sakai1,2 and Shingo Kajimura3 Abstract | White adipocytes store excess energy in the form of triglycerides, whereas brown and beige adipocytes dissipate energy in the form of heat. This thermogenic function relies on the activation of brown and beige adipocyte-specific gene programmes that are coordinately regulated by adipose-selective chromatin architectures and by a set of unique transcriptional and epigenetic regulators. A number of transcriptional and epigenetic regulators are also required for promoting beige adipocyte biogenesis in response to various environmental stimuli. A better understanding of the molecular mechanisms governing the generation and function of brown and beige adipocytes is necessary to allow us to control adipose cell fate and stimulate thermogenesis. This may provide a therapeutic approach for the treatment of obesity and obesity-associated diseases, such as type 2 diabetes. Interscapular BAT Adipose tissue has a central role in whole-body energy subjects who had previously lacked detectable BAT Brown adipose tissue (BAT) is a homeostasis. White adipose tissue (WAT) is the major depots before cold exposure, presumably owing to the specialized organ that adipose organ in mammals. It represents 10% or more emergence of new thermogenic adipocytes. This, then, produces heat. BAT is localized of the body weight of healthy adult humans and is leads to an increase in non-shivering thermogenesis in the interscapular and 6–9 perirenal regions of rodents specialized for the storage of excess energy. Humans and/or an improvement in insulin sensitivity . These and infants. -

Differential Recruitment of Nuclear Receptor Coregulators in Ligand-Dependent Transcriptional Repression by Estrogen Receptor-A
Oncogene (2011) 30, 1608–1614 & 2011 Macmillan Publishers Limited All rights reserved 0950-9232/11 www.nature.com/onc SHORT COMMUNICATION Differential recruitment of nuclear receptor coregulators in ligand-dependent transcriptional repression by estrogen receptor-a KW Merrell, JD Crofts, RL Smith, JH Sin, KE Kmetzsch, A Merrell, RO Miguel, NR Candelaria and C-Y Lin1 Department of Microbiology and Molecular Biology, Brigham Young University, Provo, UT, USA Estrogen receptors (ERs) are normally expressed in breast endocrine-related diseases, including breast cancers tissues and mediate hormonal functions during develop- (Gruber et al., 2002; Yager and Davidson, 2006). Two ment and in female reproductive physiology. In the estrogen receptor (ER) subtypes, ERa and ERb,have majority of breast cancers, ERs are involved in regulating been identified and exhibit differential target tissue tumor cell proliferation and serve as prognostic markers distribution and functions. In the majority of breast and therapeutic targets in the management of hormone- cancers, ERa is expressed in tumor cells and is involved dependent tumors. At the molecular level, ERs function as in promoting tumor cell proliferation and invasiveness ligand-dependent transcription factors and activate target- (Ali and Coombes, 2000). The role of ERb in breast gene expression following hormone stimulation. Recent cancer biology remains to be determined, although some transcriptomic and whole-genome-binding studies suggest, findings suggest that it may attenuate ERa functions however, that ligand-activated ERs can also repress the (Lindberg et al., 2003; Cappelletti et al., 2004; Lin et al., expression of a significant subset of target genes. To 2007a). ER status is an important prognostic marker characterize the molecular mechanisms of transcriptional in breast cancer management and a determinant for repression by ERs, we examined recruitment of nuclear treatment with selective estrogen receptor modulators. -

CREB-Dependent Transcription in Astrocytes: Signalling Pathways, Gene Profiles and Neuroprotective Role in Brain Injury
CREB-dependent transcription in astrocytes: signalling pathways, gene profiles and neuroprotective role in brain injury. Tesis doctoral Luis Pardo Fernández Bellaterra, Septiembre 2015 Instituto de Neurociencias Departamento de Bioquímica i Biologia Molecular Unidad de Bioquímica y Biologia Molecular Facultad de Medicina CREB-dependent transcription in astrocytes: signalling pathways, gene profiles and neuroprotective role in brain injury. Memoria del trabajo experimental para optar al grado de doctor, correspondiente al Programa de Doctorado en Neurociencias del Instituto de Neurociencias de la Universidad Autónoma de Barcelona, llevado a cabo por Luis Pardo Fernández bajo la dirección de la Dra. Elena Galea Rodríguez de Velasco y la Dra. Roser Masgrau Juanola, en el Instituto de Neurociencias de la Universidad Autónoma de Barcelona. Doctorando Directoras de tesis Luis Pardo Fernández Dra. Elena Galea Dra. Roser Masgrau In memoriam María Dolores Álvarez Durán Abuela, eres la culpable de que haya decidido recorrer el camino de la ciencia. Que estas líneas ayuden a conservar tu recuerdo. A mis padres y hermanos, A Meri INDEX I Summary 1 II Introduction 3 1 Astrocytes: physiology and pathology 5 1.1 Anatomical organization 6 1.2 Origins and heterogeneity 6 1.3 Astrocyte functions 8 1.3.1 Developmental functions 8 1.3.2 Neurovascular functions 9 1.3.3 Metabolic support 11 1.3.4 Homeostatic functions 13 1.3.5 Antioxidant functions 15 1.3.6 Signalling functions 15 1.4 Astrocytes in brain pathology 20 1.5 Reactive astrogliosis 22 2 The transcription