In Acute Myeloid Leukemia
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
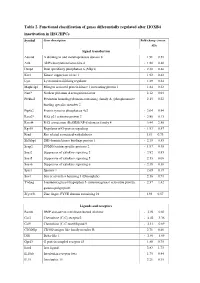
Table 2. Functional Classification of Genes Differentially Regulated After HOXB4 Inactivation in HSC/Hpcs
Table 2. Functional classification of genes differentially regulated after HOXB4 inactivation in HSC/HPCs Symbol Gene description Fold-change (mean ± SD) Signal transduction Adam8 A disintegrin and metalloprotease domain 8 1.91 ± 0.51 Arl4 ADP-ribosylation factor-like 4 - 1.80 ± 0.40 Dusp6 Dual specificity phosphatase 6 (Mkp3) - 2.30 ± 0.46 Ksr1 Kinase suppressor of ras 1 1.92 ± 0.42 Lyst Lysosomal trafficking regulator 1.89 ± 0.34 Mapk1ip1 Mitogen activated protein kinase 1 interacting protein 1 1.84 ± 0.22 Narf* Nuclear prelamin A recognition factor 2.12 ± 0.04 Plekha2 Pleckstrin homology domain-containing. family A. (phosphoinosite 2.15 ± 0.22 binding specific) member 2 Ptp4a2 Protein tyrosine phosphatase 4a2 - 2.04 ± 0.94 Rasa2* RAS p21 activator protein 2 - 2.80 ± 0.13 Rassf4 RAS association (RalGDS/AF-6) domain family 4 3.44 ± 2.56 Rgs18 Regulator of G-protein signaling - 1.93 ± 0.57 Rrad Ras-related associated with diabetes 1.81 ± 0.73 Sh3kbp1 SH3 domain kinase bindings protein 1 - 2.19 ± 0.53 Senp2 SUMO/sentrin specific protease 2 - 1.97 ± 0.49 Socs2 Suppressor of cytokine signaling 2 - 2.82 ± 0.85 Socs5 Suppressor of cytokine signaling 5 2.13 ± 0.08 Socs6 Suppressor of cytokine signaling 6 - 2.18 ± 0.38 Spry1 Sprouty 1 - 2.69 ± 0.19 Sos1 Son of sevenless homolog 1 (Drosophila) 2.16 ± 0.71 Ywhag 3-monooxygenase/tryptophan 5- monooxygenase activation protein. - 2.37 ± 1.42 gamma polypeptide Zfyve21 Zinc finger. FYVE domain containing 21 1.93 ± 0.57 Ligands and receptors Bambi BMP and activin membrane-bound inhibitor - 2.94 ± 0.62 -

Cisplatin and Phenanthriplatin Modulate Long-Noncoding
www.nature.com/scientificreports OPEN Cisplatin and phenanthriplatin modulate long‑noncoding RNA expression in A549 and IMR90 cells revealing regulation of microRNAs, Wnt/β‑catenin and TGF‑β signaling Jerry D. Monroe1,2, Satya A. Moolani2,3, Elvin N. Irihamye2,4, Katheryn E. Lett1, Michael D. Hebert1, Yann Gibert1* & Michael E. Smith2* The monofunctional platinum(II) complex, phenanthriplatin, acts by blocking transcription, but its regulatory efects on long‑noncoding RNAs (lncRNAs) have not been elucidated relative to traditional platinum‑based chemotherapeutics, e.g., cisplatin. Here, we treated A549 non‑small cell lung cancer and IMR90 lung fbroblast cells for 24 h with either cisplatin, phenanthriplatin or a solvent control, and then performed microarray analysis to identify regulated lncRNAs. RNA22 v2 microRNA software was subsequently used to identify microRNAs (miRNAs) that might be suppressed by the most regulated lncRNAs. We found that miR‑25‑5p, ‑30a‑3p, ‑138‑5p, ‑149‑3p, ‑185‑5p, ‑378j, ‑608, ‑650, ‑708‑5p, ‑1253, ‑1254, ‑4458, and ‑4516, were predicted to target the cisplatin upregulated lncRNAs, IMMP2L‑1, CBR3‑1 and ATAD2B‑5, and the phenanthriplatin downregulated lncRNAs, AGO2‑1, COX7A1‑2 and SLC26A3‑1. Then, we used qRT‑PCR to measure the expression of miR‑25‑5p, ‑378j, ‑4516 (A549) and miR‑149‑3p, ‑608, and ‑4458 (IMR90) to identify distinct signaling efects associated with cisplatin and phenanthriplatin. The signaling pathways associated with these miRNAs suggests that phenanthriplatin may modulate Wnt/β‑catenin and TGF‑β signaling through the MAPK/ ERK and PTEN/AKT pathways diferently than cisplatin. Further, as some of these miRNAs may be subject to dissimilar lncRNA targeting in A549 and IMR90 cells, the monofunctional complex may not cause toxicity in normal lung compared to cancer cells by acting through distinct lncRNA and miRNA networks. -

Transcriptome Analyses of Rhesus Monkey Pre-Implantation Embryos Reveal A
Downloaded from genome.cshlp.org on September 23, 2021 - Published by Cold Spring Harbor Laboratory Press Transcriptome analyses of rhesus monkey pre-implantation embryos reveal a reduced capacity for DNA double strand break (DSB) repair in primate oocytes and early embryos Xinyi Wang 1,3,4,5*, Denghui Liu 2,4*, Dajian He 1,3,4,5, Shengbao Suo 2,4, Xian Xia 2,4, Xiechao He1,3,6, Jing-Dong J. Han2#, Ping Zheng1,3,6# Running title: reduced DNA DSB repair in monkey early embryos Affiliations: 1 State Key Laboratory of Genetic Resources and Evolution, Kunming Institute of Zoology, Chinese Academy of Sciences, Kunming, Yunnan 650223, China 2 Key Laboratory of Computational Biology, CAS Center for Excellence in Molecular Cell Science, Collaborative Innovation Center for Genetics and Developmental Biology, Chinese Academy of Sciences-Max Planck Partner Institute for Computational Biology, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences, Shanghai 200031, China 3 Yunnan Key Laboratory of Animal Reproduction, Kunming Institute of Zoology, Chinese Academy of Sciences, Kunming, Yunnan 650223, China 4 University of Chinese Academy of Sciences, Beijing, China 5 Kunming College of Life Science, University of Chinese Academy of Sciences, Kunming, Yunnan 650204, China 6 Primate Research Center, Kunming Institute of Zoology, Chinese Academy of Sciences, Kunming, 650223, China * Xinyi Wang and Denghui Liu contributed equally to this work 1 Downloaded from genome.cshlp.org on September 23, 2021 - Published by Cold Spring Harbor Laboratory Press # Correspondence: Jing-Dong J. Han, Email: [email protected]; Ping Zheng, Email: [email protected] Key words: rhesus monkey, pre-implantation embryo, DNA damage 2 Downloaded from genome.cshlp.org on September 23, 2021 - Published by Cold Spring Harbor Laboratory Press ABSTRACT Pre-implantation embryogenesis encompasses several critical events including genome reprogramming, zygotic genome activation (ZGA) and cell fate commitment. -

The Structure-Function Relationship of Angular Estrogens and Estrogen Receptor Alpha to Initiate Estrogen-Induced Apoptosis in Breast Cancer Cells S
Supplemental material to this article can be found at: http://molpharm.aspetjournals.org/content/suppl/2020/05/03/mol.120.119776.DC1 1521-0111/98/1/24–37$35.00 https://doi.org/10.1124/mol.120.119776 MOLECULAR PHARMACOLOGY Mol Pharmacol 98:24–37, July 2020 Copyright ª 2020 The Author(s) This is an open access article distributed under the CC BY Attribution 4.0 International license. The Structure-Function Relationship of Angular Estrogens and Estrogen Receptor Alpha to Initiate Estrogen-Induced Apoptosis in Breast Cancer Cells s Philipp Y. Maximov, Balkees Abderrahman, Yousef M. Hawsawi, Yue Chen, Charles E. Foulds, Antrix Jain, Anna Malovannaya, Ping Fan, Ramona F. Curpan, Ross Han, Sean W. Fanning, Bradley M. Broom, Daniela M. Quintana Rincon, Jeffery A. Greenland, Geoffrey L. Greene, and V. Craig Jordan Downloaded from Departments of Breast Medical Oncology (P.Y.M., B.A., P.F., D.M.Q.R., J.A.G., V.C.J.) and Computational Biology and Bioinformatics (B.M.B.), University of Texas, MD Anderson Cancer Center, Houston, Texas; King Faisal Specialist Hospital and Research (Gen.Org.), Research Center, Jeddah, Kingdom of Saudi Arabia (Y.M.H.); The Ben May Department for Cancer Research, University of Chicago, Chicago, Illinois (R.H., S.W.F., G.L.G.); Center for Precision Environmental Health and Department of Molecular and Cellular Biology (C.E.F.), Mass Spectrometry Proteomics Core (A.J., A.M.), Verna and Marrs McLean Department of Biochemistry and Molecular Biology, Mass Spectrometry Proteomics Core (A.M.), and Dan L. Duncan molpharm.aspetjournals.org -

TAZ-CAMTA1 and YAP-TFE3 Alter the TAZ/YAP Transcriptome By
RESEARCH ARTICLE TAZ-CAMTA1 and YAP-TFE3 alter the TAZ/YAP transcriptome by recruiting the ATAC histone acetyltransferase complex Nicole Merritt1†, Keith Garcia1,2†, Dushyandi Rajendran3, Zhen-Yuan Lin3, Xiaomeng Zhang4, Katrina A Mitchell4,5, Nicholas Borcherding6, Colleen Fullenkamp1, Michael S Chimenti7, Anne-Claude Gingras3, Kieran F Harvey4,5,8, Munir R Tanas1,2,9,10* 1Department of Pathology, University of Iowa, Iowa City, United States; 2Cancer Biology Graduate Program, University of Iowa, Iowa City, United States; 3Lunenfeld- Tanenbaum Research Institute, Mount Sinai Hospital, Toronto, United States; 4Peter MacCallum Cancer Centre, Melbourne, Australia; 5Sir Peter MacCallum Department of Oncology, The University of Melbourne, Parkville, Australia; 6Department of Pathology and Immunology, Washington University, St. Louis, United States; 7Iowa Institute of Human Genetics, Carver College of Medicine, University of Iowa, Iowa City, United States; 8Department of Anatomy and Developmental Biology and Biomedicine Discovery Institute, Monash University, Clayton, Australia; 9Holden Comprehensive Cancer Center, University of Iowa, Iowa City, United States; 10Pathology and Laboratory Medicine, Veterans Affairs Medical Center, Iowa City, United States *For correspondence: Abstract Epithelioid hemangioendothelioma (EHE) is a vascular sarcoma that metastasizes early [email protected] in its clinical course and lacks an effective medical therapy. The TAZ-CAMTA1 and YAP-TFE3 fusion proteins are chimeric transcription factors and initiating oncogenic drivers of EHE. A combined †These authors contributed proteomic/genetic screen in human cell lines identified YEATS2 and ZZZ3, components of the equally to this work Ada2a-containing histone acetyltransferase (ATAC) complex, as key interactors of both fusion Competing interests: The proteins despite the dissimilarity of the C terminal fusion partners CAMTA1 and TFE3. -

Supplementary Figures and Table Legends
Supplementary Information Supplementary Figure Legends Figure S1. CARM1 is required for estrogen-induced gene transcriptional activation. (A) Box plot showing the expression of CARM1 (FPKM) in a cohort of clinical breast cancer (n=1,102) and normal (n=113) samples from TCGA (The Cancer Genome Atlas). (B, C) Kaplan Meier survival analyses for DMFS (distal metastasis free survival) (n=1,747) and OS (overall survival) (n=1,402) of breast cancer patients using CARM1 as input. (D, E) RNA-seq as described in Figure 1A for specific genes were shown as indicated. (F) Correlation of CARM1’s effects on whole transcriptome in response to estrogen between two RNA-seq biological repeats (n=11,816). (G) Correlation of CARM1 effects on whole transcriptome in response to estrogen based on RNA- seq between two independent siRNAs targeting CARM1 (siCARM1 (1) and siCARM1 (2)) (n=12,234). (H, I) MCF7 cells as described in (G) were subjected to immunoblotting (IB) (H) and RT-qPCR (I) analysis to examine the levels of indicated protein and mRNA, respectively. Actin was served as a loading control. (J) MCF7 cells as described in Figure 1E were subjected to RT-qPCR analysis to examine the expression of genes as indicated. (K) MCF7 cells as described in Figure 1E were subjected to immunoblotting (IB) analysis to examine the protein levels of CARM1. Actin was served as a loading control. (L) Genomic DNA was extracted from CARM1 knockout cells, followed by PCR using specific primer sets spanning gRNA targeting region (boxed in blue). The resultant PCR products were subjected to Sanger sequencing as shown. -

Histone Demethylases at the Center of Cellular Differentiation and Disease
Downloaded from genesdev.cshlp.org on September 30, 2021 - Published by Cold Spring Harbor Laboratory Press REVIEW Erasing the methyl mark: histone demethylases at the center of cellular differentiation and disease Paul A.C. Cloos,2 Jesper Christensen, Karl Agger, and Kristian Helin1 Biotech Research and Innovation Centre (BRIC) and Centre for Epigenetics, University of Copenhagen, DK-2200 Copenhagen, Denmark The enzymes catalyzing lysine and arginine methylation impacts on the transcriptional activity of the underlying of histones are essential for maintaining transcriptional DNA by acting as a recognition template for effector programs and determining cell fate and identity. Until proteins modifying the chromatin environment and lead- recently, histone methylation was regarded irreversible. ing to either repression or activation. Thus, histone However, within the last few years, several families of methylation can be associated with either activation or histone demethylases erasing methyl marks associated repression of transcription depending on which effector with gene repression or activation have been identified, protein is being recruited. It should be noted that the underscoring the plasticity and dynamic nature of his- unmodified residues can also serve as a binding template tone methylation. Recent discoveries have revealed that for effector proteins leading to specific chromatin states histone demethylases take part in large multiprotein (Lan et al. 2007b). complexes synergizing with histone deacetylases, histone Arginine residues can be modified by one or two meth- methyltransferases, and nuclear receptors to control de- yl groups; the latter form in either a symmetric or asym- velopmental and transcriptional programs. Here we re- metric conformation (Rme1, Rme2s, and Rme2a), per- view the emerging biochemical and biological functions mitting a total of four states: one unmethylated and of the histone demethylases and discuss their potential three methylated forms. -

Stromule Extension Along Microtubules Coordinated with Actin
RESEARCH ARTICLE Stromule extension along microtubules coordinated with actin-mediated anchoring guides perinuclear chloroplast movement during innate immunity Amutha Sampath Kumar1†, Eunsook Park2,3†‡, Alexander Nedo1,4, Ali Alqarni1,4,5, Li Ren5, Kyle Hoban1,4, Shannon Modla1, John H McDonald4, Chandra Kambhamettu5,6, Savithramma P Dinesh-Kumar2,3*, Jeffrey Lewis Caplan1,4,5* 1Delaware Biotechnology Institute, University of Delaware, Newark, United States; 2Department of Plant Biology, College of Biological Sciences, University of California, Davis, Davis, United States; 3The Genome Center, College of Biological Sciences, University of California, Davis, Davis, United States; 4Department of Biological Sciences, College of Arts and Sciences, University of Delaware, Newark, United States; 5Department of Plant and Soil Sciences, College of Agriculture and Natural Resources, University of Delaware, Newark, United States; 6Department of Computer and Information Sciences, College of Engineering, University of *For correspondence: Delaware, Newark, United States [email protected] (SPD-K); [email protected] (JLC) † These authors contributed Abstract Dynamic tubular extensions from chloroplasts called stromules have recently been equally to this work shown to connect with nuclei and function during innate immunity. We demonstrate that stromules Present address: ‡Department extend along microtubules (MTs) and MT organization directly affects stromule dynamics since of Plant Sciences, College of stabilization of MTs chemically or genetically -

CRISPR-Enhanced Human Adipocyte “Browning” As Cell Therapy for Metabolic Disease
bioRxiv preprint doi: https://doi.org/10.1101/2020.10.13.337923; this version posted October 13, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. CRISPR-enhanced human adipocyte “browning” as cell therapy for metabolic disease Emmanouela Tsagkaraki1,6*, Sarah Nicoloro1*, Tiffany De Souza1, Javier Solivan-Rivera1, Anand Desai1, Yuefei Shen1, Mark Kelly1, Adilson Guilherme1, Felipe Henriques1, Raed Ibraheim2, Nadia Amrani2, Kevin Luk4, Stacy Maitland4, Randall H. Friedline1, Lauren Tauer1, Xiaodi Hu1, Jason K. Kim1,3, Scot A. Wolfe4,5, Erik J. Sontheimer2,5, Silvia Corvera1**, Michael P. Czech1**. 1Program in Molecular Medicine, University of Massachusetts Medical School, Worcester, MA 01605 2RNA Therapeutics Institute, University of Massachusetts Medical School, Worcester MA 01605 3Division of Endocrinology, Metabolism and Diabetes, Department of Medicine, University of Massachusetts Medical School Worcester, MA 01605 4Department of Molecular,Cell and Cancer Biology, University of Massachusetts Medical School, Worcester MA 01605 5Li Weibo Institute for Rare Diseases Research, University of Massachusetts Medical School, Worcester, MA 01605 6University of Crete School of Medicine, Crete 71003 *These authors contributed equally **Co-corresponding authors Michael P. Czech 373 Plantation Street Program in Molecular Medicine University of Massachusetts Medical School Worcester, MA 01605 [email protected] Tel: 1 508 856 2254 Fax: 1 508 856 1617 Silvia Corvera 373 Plantation Street Program in Molecular Medicine University of Massachusetts Medical School Worcester, MA 01605 [email protected] Tel: 1 508 856 6869 Fax: 1 508 856 1617 bioRxiv preprint doi: https://doi.org/10.1101/2020.10.13.337923; this version posted October 13, 2020. -

Human Induced Pluripotent Stem Cell–Derived Podocytes Mature Into Vascularized Glomeruli Upon Experimental Transplantation
BASIC RESEARCH www.jasn.org Human Induced Pluripotent Stem Cell–Derived Podocytes Mature into Vascularized Glomeruli upon Experimental Transplantation † Sazia Sharmin,* Atsuhiro Taguchi,* Yusuke Kaku,* Yasuhiro Yoshimura,* Tomoko Ohmori,* ‡ † ‡ Tetsushi Sakuma, Masashi Mukoyama, Takashi Yamamoto, Hidetake Kurihara,§ and | Ryuichi Nishinakamura* *Department of Kidney Development, Institute of Molecular Embryology and Genetics, and †Department of Nephrology, Faculty of Life Sciences, Kumamoto University, Kumamoto, Japan; ‡Department of Mathematical and Life Sciences, Graduate School of Science, Hiroshima University, Hiroshima, Japan; §Division of Anatomy, Juntendo University School of Medicine, Tokyo, Japan; and |Japan Science and Technology Agency, CREST, Kumamoto, Japan ABSTRACT Glomerular podocytes express proteins, such as nephrin, that constitute the slit diaphragm, thereby contributing to the filtration process in the kidney. Glomerular development has been analyzed mainly in mice, whereas analysis of human kidney development has been minimal because of limited access to embryonic kidneys. We previously reported the induction of three-dimensional primordial glomeruli from human induced pluripotent stem (iPS) cells. Here, using transcription activator–like effector nuclease-mediated homologous recombination, we generated human iPS cell lines that express green fluorescent protein (GFP) in the NPHS1 locus, which encodes nephrin, and we show that GFP expression facilitated accurate visualization of nephrin-positive podocyte formation in -

Supplemental Digital Content (Sdc) Sdc, Materials
SUPPLEMENTAL DIGITAL CONTENT (SDC) SDC, MATERIALS AND METHODS Animals This study used 9-12 week old male C57BL/6 mice (Jackson Laboratory, Bar Harbor, ME). This study conformed to the National Institutes of Health guidelines and was conducted under animal protocols approved by the University of Virginia’s Institutional Animal Care and Use Committee. Murine DCD Lung Procedure Mice were anesthetized by isoflurane inhalation and euthanized by cervical dislocation followed by a 60-minute period of “no-touch” warm ischemia. Mice then underwent extended median sternotomy and midline cervical exposure followed by intubation for the initiation of mechanical ventilation at 120 strokes/minute with room air. The left atrium was vented via an atriotomy followed by infusion of the lungs with 3 mL 4°C Perfadex® solution (Vitrolife Inc., Denver, CO) supplemented with THAM Solution (Vitrolife, Kungsbacka, Sweden), estimating weight-based volume recommendations for pulmonary artery perfusion (140mL/kg) (1). The chest was then packed with ice and the trachea occluded by silk-suture tie at tidal volume (7µL/g body weight) prior to cold static preservation (CSP) for 60 minutes at 4°C. Mice were then randomized into three experimental groups: 1) CSP alone with no EVLP, 2) EVLP with Steen solution and 3) EVLP with Steen solution supplemented with the highly selective A2AR agonist, ATL1223 (30nM, Lewis and Clark Pharmaceuticals, Charlottesville, VA). Mice treated with ATL1223 during EVLP also received ATL1223 treatment (30nM) during the Perfadex flush prior to CSP whereas the EVLP group received vehicle (DMSO) during the flush. CSP lungs, which did not undergo EVLP, underwent immediate functional assessment after re-intubation as described below. -

The Expression of NRIP1 and LCOR in Endometrioid Endometrial Cancer
in vivo 35 : 2631-2640 (2021) doi:10.21873/invivo.12545 The Expression of NRIP1 and LCOR in Endometrioid Endometrial Cancer STEFANOS FLINDRIS 1, NIKOLAOS KATSOULAS 2, ANNA GOUSSIA 3, ANDREAS CHRISTOS LAZARIS 2, IORDANIS NAVROZOGLOU 1, MINAS PASCHOPOULOS 1 and IRENE THYMARA 2 1Department of Obstetrics and Gynecology, University Hospital of Ioannina, Ioannina, Greece; 2First Department of Pathology, School of Medicine, National and Kapodistrian University of Athens, Laiko General Hospital of Athens, Athens, Greece; 3Department of Pathology, University Hospital of Ioannina, Ioannina, Greece Abstract. Background: The aim of the study was to Endometrial cancer is the sixth most common cancer among analyze the expression of nuclear receptor interacting females worldwide and the fourth leading cause of cancer protein 1 (NRIP1) and its partner ligand-dependent nuclear (1). It is considered as the most common gynecological receptor co-repressor (LCOR) in endometrioid endometrial malignancy, accounting for 7% of all female cancer (1). Two cancer and to investigate their association with estrogen broad pathogenetic types of endometrial carcinoma have receptor (ER), progesterone receptor (PR), Ki-67, been recognized: Type I tumors are endometrioid carcinomas clinicopathological parameters and patient survival. and type II are non-endometrioid, including serous and clear- Materials and Methods: Immunohistochemical evaluation cell carcinomas (2). Most cases (90%) of endometrial was carried out to investigate the subcellular expression of carcinomas occur in women older than 50 years of age, with NRIP1 and LCOR in endometrioid endometrial cancer a median age of 62 years at diagnosis, and are detected in samples. Statistical analysis was used to identify the early stages [80% in International Federation of Obstetrics correlations of NRIP1 and LCOR expression with and Gynecology (FIGO) stage I] (2, 3).