The Expression of NRIP1 and LCOR in Endometrioid Endometrial Cancer
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Investigation of RIP140 and Lcor As Independent Markers for Poor Prognosis in Cervical Cancer
www.impactjournals.com/oncotarget/ Oncotarget, 2017, Vol. 8, (No. 62), pp: 105356-105371 Research Paper Investigation of RIP140 and LCoR as independent markers for poor prognosis in cervical cancer Aurelia Vattai1, Vincent Cavailles2, Sophie Sixou3, Susanne Beyer1, Christina Kuhn1, Mina Peryanova1, Helene Heidegger1, Kerstin Hermelink1, Doris Mayr4, Sven Mahner1, Christian Dannecker1, Udo Jeschke1 and Bernd Kost1 1Department of Gynaecology and Obstetrics, Ludwig-Maximilians University of Munich, 80337 Munich, Germany 2Institut de Recherche en Cancérologie de Montpellier (IRCM), INSERM U1194, Université Montpellier, F-34298 Montpellier, France 3Université Toulouse III - Paul Sabatier, F-31062 Toulouse, France 4Department of Pathology, Ludwig-Maximilians University of Munich, 81337 Munich, Germany Correspondence to: Udo Jeschke, email: [email protected] Keywords: cervical carcinoma; squamous cell carcinoma; adenocarcinoma; RIP140/NRIP1; LCoR Received: May 18, 2017 Accepted: July 25, 2017 Published: October 31, 2017 Copyright: Vattai et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License 3.0 (CC BY 3.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. ABSTRACT Introduction: RIP140 (Receptor Interacting Protein) is involved in the regulation of oncogenic signaling pathways and in the development of breast and colon cancers. The aim of the study was to analyze the expression of RIP140 and its partner LCoR in cervical cancers, to decipher their relationship with histone protein modifications and to identify a potential link with patient survival. Methods: Immunohistochemical analyses were carried out to quantify RIP140 and LCoR expression in formalin-fixed paraffin-embedded tissue sections cervical cancer samples. -
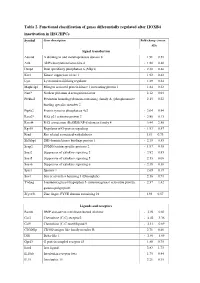
Table 2. Functional Classification of Genes Differentially Regulated After HOXB4 Inactivation in HSC/Hpcs
Table 2. Functional classification of genes differentially regulated after HOXB4 inactivation in HSC/HPCs Symbol Gene description Fold-change (mean ± SD) Signal transduction Adam8 A disintegrin and metalloprotease domain 8 1.91 ± 0.51 Arl4 ADP-ribosylation factor-like 4 - 1.80 ± 0.40 Dusp6 Dual specificity phosphatase 6 (Mkp3) - 2.30 ± 0.46 Ksr1 Kinase suppressor of ras 1 1.92 ± 0.42 Lyst Lysosomal trafficking regulator 1.89 ± 0.34 Mapk1ip1 Mitogen activated protein kinase 1 interacting protein 1 1.84 ± 0.22 Narf* Nuclear prelamin A recognition factor 2.12 ± 0.04 Plekha2 Pleckstrin homology domain-containing. family A. (phosphoinosite 2.15 ± 0.22 binding specific) member 2 Ptp4a2 Protein tyrosine phosphatase 4a2 - 2.04 ± 0.94 Rasa2* RAS p21 activator protein 2 - 2.80 ± 0.13 Rassf4 RAS association (RalGDS/AF-6) domain family 4 3.44 ± 2.56 Rgs18 Regulator of G-protein signaling - 1.93 ± 0.57 Rrad Ras-related associated with diabetes 1.81 ± 0.73 Sh3kbp1 SH3 domain kinase bindings protein 1 - 2.19 ± 0.53 Senp2 SUMO/sentrin specific protease 2 - 1.97 ± 0.49 Socs2 Suppressor of cytokine signaling 2 - 2.82 ± 0.85 Socs5 Suppressor of cytokine signaling 5 2.13 ± 0.08 Socs6 Suppressor of cytokine signaling 6 - 2.18 ± 0.38 Spry1 Sprouty 1 - 2.69 ± 0.19 Sos1 Son of sevenless homolog 1 (Drosophila) 2.16 ± 0.71 Ywhag 3-monooxygenase/tryptophan 5- monooxygenase activation protein. - 2.37 ± 1.42 gamma polypeptide Zfyve21 Zinc finger. FYVE domain containing 21 1.93 ± 0.57 Ligands and receptors Bambi BMP and activin membrane-bound inhibitor - 2.94 ± 0.62 -

Cisplatin and Phenanthriplatin Modulate Long-Noncoding
www.nature.com/scientificreports OPEN Cisplatin and phenanthriplatin modulate long‑noncoding RNA expression in A549 and IMR90 cells revealing regulation of microRNAs, Wnt/β‑catenin and TGF‑β signaling Jerry D. Monroe1,2, Satya A. Moolani2,3, Elvin N. Irihamye2,4, Katheryn E. Lett1, Michael D. Hebert1, Yann Gibert1* & Michael E. Smith2* The monofunctional platinum(II) complex, phenanthriplatin, acts by blocking transcription, but its regulatory efects on long‑noncoding RNAs (lncRNAs) have not been elucidated relative to traditional platinum‑based chemotherapeutics, e.g., cisplatin. Here, we treated A549 non‑small cell lung cancer and IMR90 lung fbroblast cells for 24 h with either cisplatin, phenanthriplatin or a solvent control, and then performed microarray analysis to identify regulated lncRNAs. RNA22 v2 microRNA software was subsequently used to identify microRNAs (miRNAs) that might be suppressed by the most regulated lncRNAs. We found that miR‑25‑5p, ‑30a‑3p, ‑138‑5p, ‑149‑3p, ‑185‑5p, ‑378j, ‑608, ‑650, ‑708‑5p, ‑1253, ‑1254, ‑4458, and ‑4516, were predicted to target the cisplatin upregulated lncRNAs, IMMP2L‑1, CBR3‑1 and ATAD2B‑5, and the phenanthriplatin downregulated lncRNAs, AGO2‑1, COX7A1‑2 and SLC26A3‑1. Then, we used qRT‑PCR to measure the expression of miR‑25‑5p, ‑378j, ‑4516 (A549) and miR‑149‑3p, ‑608, and ‑4458 (IMR90) to identify distinct signaling efects associated with cisplatin and phenanthriplatin. The signaling pathways associated with these miRNAs suggests that phenanthriplatin may modulate Wnt/β‑catenin and TGF‑β signaling through the MAPK/ ERK and PTEN/AKT pathways diferently than cisplatin. Further, as some of these miRNAs may be subject to dissimilar lncRNA targeting in A549 and IMR90 cells, the monofunctional complex may not cause toxicity in normal lung compared to cancer cells by acting through distinct lncRNA and miRNA networks. -

In Acute Myeloid Leukemia
7283 Original Article Comprehensively analyze the expression and prognostic role for ten-eleven translocations (TETs) in acute myeloid leukemia Yan Huang1#, Jie Wei1#, Xunjun Huang1, Weijie Zhou1, Yuling Xu2, Dong-Hong Deng2, Peng Cheng2 1Department of Hematology and Rheumatology, People’s Hospital of Baise, Baise, China; 2Department of Hematology, the First Affiliated Hospital of Guangxi Medical University, Nanning, China Contributions: (I) Conception and design: J Wei, Y Huang; (II) Administrative support: X Huang, Y Xu; (III) Provision of study materials or patients: W Zhou; (IV) Collection and assembly of data: J Wei, Y Huang; (V) Data analysis and interpretation: DH Deng, P Cheng; (VI) Manuscript writing: All authors; (VII) Final approval of manuscript: All authors. #These authors contributed equally to this work. Correspondence to: Yuling Xu. Department of Hematology, The First Affiliated Hospital of Guangxi Medical University, Nanning, China. Email: [email protected]; Dong-Hong Deng. Department of Hematology, The First Affiliated Hospital of Guangxi Medical University, Nanning, China. Email: [email protected]; Peng Cheng. Department of Hematology, The First Affiliated Hospital of Guangxi Medical University, Nanning 530000, China. Email: [email protected]. Background: The ten-eleven translocation (TET) family oxidize 5-methylcytosines (5mCs) and promote the locus-specific reversal of DNA. The role of TETs in acute myeloid leukemia (AML) is mostly unknown. Methods: TETs mRNA expression levels were analyzed via Gene Expression Profiling Interactive Analysis (GEPIA). The association TETs expression levels and methylation with prognosis by UALCAN GenomicScape, and METHsurv. We analyzed TETs’ aberration types, located mutations, and structures via cBioPortal. GeneMANIA performed the functional network. Gene ontology (GO) enrichment was analyzed via LinkedOmics. -

PALI1 Facilitates DNA and Nucleosome Binding by PRC2 and Triggers an Allosteric Activation of Catalysis
ARTICLE https://doi.org/10.1038/s41467-021-24866-3 OPEN PALI1 facilitates DNA and nucleosome binding by PRC2 and triggers an allosteric activation of catalysis Qi Zhang1,3, Samuel C. Agius1,3, Sarena F. Flanigan1, Michael Uckelmann1, Vitalina Levina1, Brady M. Owen1 & ✉ Chen Davidovich 1,2 1234567890():,; The polycomb repressive complex 2 (PRC2) is a histone methyltransferase that maintains cell identities. JARID2 is the only accessory subunit of PRC2 that known to trigger an allosteric activation of methyltransferase. Yet, this mechanism cannot be generalised to all PRC2 variants as, in vertebrates, JARID2 is mutually exclusive with most of the accessory subunits of PRC2. Here we provide functional and structural evidence that the vertebrate- specific PRC2 accessory subunit PALI1 emerged through a convergent evolution to mimic JARID2 at the molecular level. Mechanistically, PRC2 methylates PALI1 K1241, which then binds to the PRC2-regulatory subunit EED to allosterically activate PRC2. PALI1 K1241 is methylated in mouse and human cell lines and is essential for PALI1-induced allosteric activation of PRC2. High-resolution crystal structures revealed that PALI1 mimics the reg- ulatory interactions formed between JARID2 and EED. Independently, PALI1 also facilitates DNA and nucleosome binding by PRC2. In acute myelogenous leukemia cells, overexpression of PALI1 leads to cell differentiation, with the phenotype altered by a separation-of-function PALI1 mutation, defective in allosteric activation and active in DNA binding. Collectively, we show that PALI1 facilitates catalysis and substrate binding by PRC2 and provide evidence that subunit-induced allosteric activation is a general property of holo-PRC2 complexes. 1 Department of Biochemistry and Molecular Biology, Biomedicine Discovery Institute, Faculty of Medicine, Nursing and Health Sciences, Monash University, Clayton, VIC, Australia. -

Regulation of Lcor and RIP140 Expression in Cervical Intraepithelial Neoplasia and Correlation with CIN Progression and Dediffer
Journal of Cancer Research and Clinical Oncology (2020) 146:1847–1855 https://doi.org/10.1007/s00432-020-03178-x ORIGINAL ARTICLE – CLINICAL ONCOLOGY Regulation of LCoR and RIP140 expression in cervical intraepithelial neoplasia and correlation with CIN progression and dediferentiation Tilman L. R. Vogelsang1 · Elisa Schmoeckel3 · Christina Kuhn1 · Thomas Blankenstein1 · Mina Temelkov1 · Helene Heidegger1 · Theresa Maria Kolben1 · Thomas Kolben1 · Sven Mahner1 · Doris Mayr3 · Udo Jeschke1,2 · Aurelia Vattai1 Received: 10 February 2020 / Accepted: 3 March 2020 / Published online: 10 March 2020 © The Author(s) 2020 Abstract Purpose Ligand-dependent corepressor (LCoR) and receptor-interacting protein 140 (RIP140/NRIP1) play an important role in the regulation of multiple oncogenic signaling pathways and the development of cancer. LCoR and RIP140 form a nuclear complex in breast cancer cells and are of prognostic value in further prostate and cervical cancer. The purpose of this study was to analyze the regulation of these proteins in the development of cervical intraepithelial neoplasia (CIN I–III). Methods Immunohistochemical analysis was obtained to quantify RIP140 and LCoR expression in formalin-fxed parafn embedded tissue sections of cervical intraepithelial neoplasia samples. Tissue (n = 94) was collected from patients treated in the Department of Gynecology and Obstetrics, Ludwig-Maximilians-University of Munich, Germany, between 2002 and 2014. Correlations of expression levels with clinical outcome were carried out to assess for prognostic relevance in patients with CIN2 progression. Kruskal–Wallis test and Mann–Whitney U test were used for data analysis. Results Nuclear LCoR overexpression correlates signifcantly with CIN II progression. Nuclear RIP140 expression signif- cantly increases and nuclear LCoR expression decreases with higher grading of cervical intraepithelial neoplasia. -

Supplementary Figures and Table Legends
Supplementary Information Supplementary Figure Legends Figure S1. CARM1 is required for estrogen-induced gene transcriptional activation. (A) Box plot showing the expression of CARM1 (FPKM) in a cohort of clinical breast cancer (n=1,102) and normal (n=113) samples from TCGA (The Cancer Genome Atlas). (B, C) Kaplan Meier survival analyses for DMFS (distal metastasis free survival) (n=1,747) and OS (overall survival) (n=1,402) of breast cancer patients using CARM1 as input. (D, E) RNA-seq as described in Figure 1A for specific genes were shown as indicated. (F) Correlation of CARM1’s effects on whole transcriptome in response to estrogen between two RNA-seq biological repeats (n=11,816). (G) Correlation of CARM1 effects on whole transcriptome in response to estrogen based on RNA- seq between two independent siRNAs targeting CARM1 (siCARM1 (1) and siCARM1 (2)) (n=12,234). (H, I) MCF7 cells as described in (G) were subjected to immunoblotting (IB) (H) and RT-qPCR (I) analysis to examine the levels of indicated protein and mRNA, respectively. Actin was served as a loading control. (J) MCF7 cells as described in Figure 1E were subjected to RT-qPCR analysis to examine the expression of genes as indicated. (K) MCF7 cells as described in Figure 1E were subjected to immunoblotting (IB) analysis to examine the protein levels of CARM1. Actin was served as a loading control. (L) Genomic DNA was extracted from CARM1 knockout cells, followed by PCR using specific primer sets spanning gRNA targeting region (boxed in blue). The resultant PCR products were subjected to Sanger sequencing as shown. -

HER2 Gene Signatures: (I) Novel and (Ii) Established by Desmedt Et Al 2008 (31)
Table S1: HER2 gene signatures: (i) Novel and (ii) Established by Desmedt et al 2008 (31). Pearson R [neratinib] is the correlation with neratinib response using a pharmacogenomic model of breast cancer cell lines (accessed online via CellMinerCDB). *indicates significantly correlated genes. n/a = data not available in CellMinerCDB Genesig N Gene ID Pearson R [neratinib] p-value (i) Novel 20 ERBB2 0.77 1.90E-08* SPDEF 0.45 4.20E-03* TFAP2B 0.2 0.24 CD24 0.38 0.019* SERHL2 0.41 0.0097* CNTNAP2 0.12 0.47 RPL19 0.29 0.073 CAPN13 0.51 1.00E-03* RPL23 0.22 0.18 LRRC26 n/a n/a PRODH 0.42 9.00E-03* GPRC5C 0.44 0.0056* GGCT 0.38 1.90E-02* CLCA2 0.31 5.70E-02 KDM5B 0.33 4.20E-02* SPP1 -0.25 1.30E-01 PHLDA1 -0.54 5.30E-04* C15orf48 0.06 7.10E-01 SUSD3 -0.09 5.90E-01 SERPINA1 0.14 4.10E-01 (ii) Established 24 ERBB2 0.77 1.90E-08* PERLD1 0.77 1.20E-08* PSMD3 0.33 0.04* PNMT 0.33 4.20E-02* GSDML 0.4 1.40E-02* CASC3 0.26 0.11 LASP1 0.32 0.049* WIPF2 0.27 9.70E-02 EPN3 0.42 8.50E-03* PHB 0.38 0.019* CLCA2 0.31 5.70E-02 ORMDL2 0.06 0.74 RAP1GAP 0.53 0.00059* CUEDC1 0.09 0.61 HOXC11 0.2 0.23 CYP2J2 0.45 0.0044* HGD 0.14 0.39 ABCA12 0.07 0.67 ATP2C2 0.42 0.0096* ITGA3 0 0.98 CEACAM5 0.4 0.012* TMEM16K 0.15 0.37 NR1D1 n/a n/a SNX7 -0.28 0.092 FJX1 -0.26 0.12 KCTD9 -0.11 0.53 PCTK3 -0.04 0.83 CREG1 0.17 0.3 Table S2: Up-regulated genes from the top 500 DEGs for each comparison by WAD score METABRIC METABRIC METABRIC TCGA ERBB2amp ERBB2mut oncERBB2mut HER2+ ERBB2 PIP ANKRD30A ERBB2 GRB7 CYP4Z1 CYP4Z1 SCGB2A2 PGAP3 PROM1 LRRC26 SPDEF GSDMB CD24 PPP1R1B FOXA1 -

Expression and Role of Nuclear Receptor Coregulators in Colorectal Cancer Mouna Triki, Marion Lapierre, Vincent Cavaillès, Raja Mokdad-Gargouri
Expression and role of nuclear receptor coregulators in colorectal cancer Mouna Triki, Marion Lapierre, Vincent Cavaillès, Raja Mokdad-Gargouri To cite this version: Mouna Triki, Marion Lapierre, Vincent Cavaillès, Raja Mokdad-Gargouri. Expression and role of nuclear receptor coregulators in colorectal cancer. World Journal of Gastroenterology, Baishideng Publishing Group Co. Limited, 2017, 23 (25), pp.4480 - 4490. 10.3748/wjg.v23.i25.4480. inserm- 02438832 HAL Id: inserm-02438832 https://www.hal.inserm.fr/inserm-02438832 Submitted on 14 Jan 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Submit a Manuscript: http://www.f6publishing.com World J Gastroenterol 2017 July 7; 23(25): 4480-4490 DOI: 10.3748/wjg.v23.i25.4480 ISSN 1007-9327 (print) ISSN 2219-2840 (online) REVIEW Expression and role of nuclear receptor coregulators in colorectal cancer Mouna Triki, Marion Lapierre, Vincent Cavailles, Raja Mokdad-Gargouri Mouna Triki, Marion Lapierre, Vincent Cavailles, IRCM, Peer-review started: August 3, 2016 Institut de Recherche en Cancérologie de -

Stromule Extension Along Microtubules Coordinated with Actin
RESEARCH ARTICLE Stromule extension along microtubules coordinated with actin-mediated anchoring guides perinuclear chloroplast movement during innate immunity Amutha Sampath Kumar1†, Eunsook Park2,3†‡, Alexander Nedo1,4, Ali Alqarni1,4,5, Li Ren5, Kyle Hoban1,4, Shannon Modla1, John H McDonald4, Chandra Kambhamettu5,6, Savithramma P Dinesh-Kumar2,3*, Jeffrey Lewis Caplan1,4,5* 1Delaware Biotechnology Institute, University of Delaware, Newark, United States; 2Department of Plant Biology, College of Biological Sciences, University of California, Davis, Davis, United States; 3The Genome Center, College of Biological Sciences, University of California, Davis, Davis, United States; 4Department of Biological Sciences, College of Arts and Sciences, University of Delaware, Newark, United States; 5Department of Plant and Soil Sciences, College of Agriculture and Natural Resources, University of Delaware, Newark, United States; 6Department of Computer and Information Sciences, College of Engineering, University of *For correspondence: Delaware, Newark, United States [email protected] (SPD-K); [email protected] (JLC) † These authors contributed Abstract Dynamic tubular extensions from chloroplasts called stromules have recently been equally to this work shown to connect with nuclei and function during innate immunity. We demonstrate that stromules Present address: ‡Department extend along microtubules (MTs) and MT organization directly affects stromule dynamics since of Plant Sciences, College of stabilization of MTs chemically or genetically -

CRISPR-Enhanced Human Adipocyte “Browning” As Cell Therapy for Metabolic Disease
bioRxiv preprint doi: https://doi.org/10.1101/2020.10.13.337923; this version posted October 13, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. CRISPR-enhanced human adipocyte “browning” as cell therapy for metabolic disease Emmanouela Tsagkaraki1,6*, Sarah Nicoloro1*, Tiffany De Souza1, Javier Solivan-Rivera1, Anand Desai1, Yuefei Shen1, Mark Kelly1, Adilson Guilherme1, Felipe Henriques1, Raed Ibraheim2, Nadia Amrani2, Kevin Luk4, Stacy Maitland4, Randall H. Friedline1, Lauren Tauer1, Xiaodi Hu1, Jason K. Kim1,3, Scot A. Wolfe4,5, Erik J. Sontheimer2,5, Silvia Corvera1**, Michael P. Czech1**. 1Program in Molecular Medicine, University of Massachusetts Medical School, Worcester, MA 01605 2RNA Therapeutics Institute, University of Massachusetts Medical School, Worcester MA 01605 3Division of Endocrinology, Metabolism and Diabetes, Department of Medicine, University of Massachusetts Medical School Worcester, MA 01605 4Department of Molecular,Cell and Cancer Biology, University of Massachusetts Medical School, Worcester MA 01605 5Li Weibo Institute for Rare Diseases Research, University of Massachusetts Medical School, Worcester, MA 01605 6University of Crete School of Medicine, Crete 71003 *These authors contributed equally **Co-corresponding authors Michael P. Czech 373 Plantation Street Program in Molecular Medicine University of Massachusetts Medical School Worcester, MA 01605 [email protected] Tel: 1 508 856 2254 Fax: 1 508 856 1617 Silvia Corvera 373 Plantation Street Program in Molecular Medicine University of Massachusetts Medical School Worcester, MA 01605 [email protected] Tel: 1 508 856 6869 Fax: 1 508 856 1617 bioRxiv preprint doi: https://doi.org/10.1101/2020.10.13.337923; this version posted October 13, 2020. -

Supp Table 6.Pdf
Supplementary Table 6. Processes associated to the 2037 SCL candidate target genes ID Symbol Entrez Gene Name Process NM_178114 AMIGO2 adhesion molecule with Ig-like domain 2 adhesion NM_033474 ARVCF armadillo repeat gene deletes in velocardiofacial syndrome adhesion NM_027060 BTBD9 BTB (POZ) domain containing 9 adhesion NM_001039149 CD226 CD226 molecule adhesion NM_010581 CD47 CD47 molecule adhesion NM_023370 CDH23 cadherin-like 23 adhesion NM_207298 CERCAM cerebral endothelial cell adhesion molecule adhesion NM_021719 CLDN15 claudin 15 adhesion NM_009902 CLDN3 claudin 3 adhesion NM_008779 CNTN3 contactin 3 (plasmacytoma associated) adhesion NM_015734 COL5A1 collagen, type V, alpha 1 adhesion NM_007803 CTTN cortactin adhesion NM_009142 CX3CL1 chemokine (C-X3-C motif) ligand 1 adhesion NM_031174 DSCAM Down syndrome cell adhesion molecule adhesion NM_145158 EMILIN2 elastin microfibril interfacer 2 adhesion NM_001081286 FAT1 FAT tumor suppressor homolog 1 (Drosophila) adhesion NM_001080814 FAT3 FAT tumor suppressor homolog 3 (Drosophila) adhesion NM_153795 FERMT3 fermitin family homolog 3 (Drosophila) adhesion NM_010494 ICAM2 intercellular adhesion molecule 2 adhesion NM_023892 ICAM4 (includes EG:3386) intercellular adhesion molecule 4 (Landsteiner-Wiener blood group)adhesion NM_001001979 MEGF10 multiple EGF-like-domains 10 adhesion NM_172522 MEGF11 multiple EGF-like-domains 11 adhesion NM_010739 MUC13 mucin 13, cell surface associated adhesion NM_013610 NINJ1 ninjurin 1 adhesion NM_016718 NINJ2 ninjurin 2 adhesion NM_172932 NLGN3 neuroligin