Proteomic Profiling of Mitochondrial-Derived Vesicles In
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Analysis of Trans Esnps Infers Regulatory Network Architecture
Analysis of trans eSNPs infers regulatory network architecture Anat Kreimer Submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Graduate School of Arts and Sciences COLUMBIA UNIVERSITY 2014 © 2014 Anat Kreimer All rights reserved ABSTRACT Analysis of trans eSNPs infers regulatory network architecture Anat Kreimer eSNPs are genetic variants associated with transcript expression levels. The characteristics of such variants highlight their importance and present a unique opportunity for studying gene regulation. eSNPs affect most genes and their cell type specificity can shed light on different processes that are activated in each cell. They can identify functional variants by connecting SNPs that are implicated in disease to a molecular mechanism. Examining eSNPs that are associated with distal genes can provide insights regarding the inference of regulatory networks but also presents challenges due to the high statistical burden of multiple testing. Such association studies allow: simultaneous investigation of many gene expression phenotypes without assuming any prior knowledge and identification of unknown regulators of gene expression while uncovering directionality. This thesis will focus on such distal eSNPs to map regulatory interactions between different loci and expose the architecture of the regulatory network defined by such interactions. We develop novel computational approaches and apply them to genetics-genomics data in human. We go beyond pairwise interactions to define network motifs, including regulatory modules and bi-fan structures, showing them to be prevalent in real data and exposing distinct attributes of such arrangements. We project eSNP associations onto a protein-protein interaction network to expose topological properties of eSNPs and their targets and highlight different modes of distal regulation. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

A New Efficient Method to Detect Genetic Interactions for Lung Cancer
Luyapan et al. BMC Med Genomics (2020) 13:162 https://doi.org/10.1186/s12920-020-00807-9 TECHNICAL ADVANCE Open Access A new efcient method to detect genetic interactions for lung cancer GWAS Jennifer Luyapan1,2, Xuemei Ji2, Siting Li1,2, Xiangjun Xiao3, Dakai Zhu2,3, Eric J. Duell4, David C. Christiani5,6, Matthew B. Schabath7, Susanne M. Arnold8, Shanbeh Zienolddiny9, Hans Brunnström10, Olle Melander11, Mark D. Thornquist12, Todd A. MacKenzie1,2, Christopher I. Amos1,2,3* and Jiang Gui1,2* Abstract Background: Genome-wide association studies (GWAS) have proven successful in predicting genetic risk of disease using single-locus models; however, identifying single nucleotide polymorphism (SNP) interactions at the genome- wide scale is limited due to computational and statistical challenges. We addressed the computational burden encountered when detecting SNP interactions for survival analysis, such as age of disease-onset. To confront this problem, we developed a novel algorithm, called the Efcient Survival Multifactor Dimensionality Reduction (ES-MDR) method, which used Martingale Residuals as the outcome parameter to estimate survival outcomes, and imple- mented the Quantitative Multifactor Dimensionality Reduction method to identify signifcant interactions associated with age of disease-onset. Methods: To demonstrate efcacy, we evaluated this method on two simulation data sets to estimate the type I error rate and power. Simulations showed that ES-MDR identifed interactions using less computational workload and allowed for adjustment of covariates. We applied ES-MDR on the OncoArray-TRICL Consortium data with 14,935 cases and 12,787 controls for lung cancer (SNPs 108,254) to search over all two-way interactions to identify genetic interactions associated with lung cancer age-of-onset.= We tested the best model in an independent data set from the OncoArray-TRICL data. -

The Genetics of Bipolar Disorder
Molecular Psychiatry (2008) 13, 742–771 & 2008 Nature Publishing Group All rights reserved 1359-4184/08 $30.00 www.nature.com/mp FEATURE REVIEW The genetics of bipolar disorder: genome ‘hot regions,’ genes, new potential candidates and future directions A Serretti and L Mandelli Institute of Psychiatry, University of Bologna, Bologna, Italy Bipolar disorder (BP) is a complex disorder caused by a number of liability genes interacting with the environment. In recent years, a large number of linkage and association studies have been conducted producing an extremely large number of findings often not replicated or partially replicated. Further, results from linkage and association studies are not always easily comparable. Unfortunately, at present a comprehensive coverage of available evidence is still lacking. In the present paper, we summarized results obtained from both linkage and association studies in BP. Further, we indicated new potential interesting genes, located in genome ‘hot regions’ for BP and being expressed in the brain. We reviewed published studies on the subject till December 2007. We precisely localized regions where positive linkage has been found, by the NCBI Map viewer (http://www.ncbi.nlm.nih.gov/mapview/); further, we identified genes located in interesting areas and expressed in the brain, by the Entrez gene, Unigene databases (http://www.ncbi.nlm.nih.gov/entrez/) and Human Protein Reference Database (http://www.hprd.org); these genes could be of interest in future investigations. The review of association studies gave interesting results, as a number of genes seem to be definitively involved in BP, such as SLC6A4, TPH2, DRD4, SLC6A3, DAOA, DTNBP1, NRG1, DISC1 and BDNF. -

In This Table Protein Name, Uniprot Code, Gene Name P-Value
Supplementary Table S1: In this table protein name, uniprot code, gene name p-value and Fold change (FC) for each comparison are shown, for 299 of the 301 significantly regulated proteins found in both comparisons (p-value<0.01, fold change (FC) >+/-0.37) ALS versus control and FTLD-U versus control. Two uncharacterized proteins have been excluded from this list Protein name Uniprot Gene name p value FC FTLD-U p value FC ALS FTLD-U ALS Cytochrome b-c1 complex P14927 UQCRB 1.534E-03 -1.591E+00 6.005E-04 -1.639E+00 subunit 7 NADH dehydrogenase O95182 NDUFA7 4.127E-04 -9.471E-01 3.467E-05 -1.643E+00 [ubiquinone] 1 alpha subcomplex subunit 7 NADH dehydrogenase O43678 NDUFA2 3.230E-04 -9.145E-01 2.113E-04 -1.450E+00 [ubiquinone] 1 alpha subcomplex subunit 2 NADH dehydrogenase O43920 NDUFS5 1.769E-04 -8.829E-01 3.235E-05 -1.007E+00 [ubiquinone] iron-sulfur protein 5 ARF GTPase-activating A0A0C4DGN6 GIT1 1.306E-03 -8.810E-01 1.115E-03 -7.228E-01 protein GIT1 Methylglutaconyl-CoA Q13825 AUH 6.097E-04 -7.666E-01 5.619E-06 -1.178E+00 hydratase, mitochondrial ADP/ATP translocase 1 P12235 SLC25A4 6.068E-03 -6.095E-01 3.595E-04 -1.011E+00 MIC J3QTA6 CHCHD6 1.090E-04 -5.913E-01 2.124E-03 -5.948E-01 MIC J3QTA6 CHCHD6 1.090E-04 -5.913E-01 2.124E-03 -5.948E-01 Protein kinase C and casein Q9BY11 PACSIN1 3.837E-03 -5.863E-01 3.680E-06 -1.824E+00 kinase substrate in neurons protein 1 Tubulin polymerization- O94811 TPPP 6.466E-03 -5.755E-01 6.943E-06 -1.169E+00 promoting protein MIC C9JRZ6 CHCHD3 2.912E-02 -6.187E-01 2.195E-03 -9.781E-01 Mitochondrial 2- -

Cytogenetic and Molecular Characterization of the Macro- And
University of Ulm Department of Human Genetics Prof. Dr. med. Walther Vogel Cytogenetic and Molecular Characterization of the Macro- and Micro-inversions, which Distinguish the Human and the Chimpanzee Karyotypes - from Speciation to Polymorphism Thesis Applying for the Degree of Doctor of Human Biology (Dr. hum. biol.) Faculty of Medicine University of Ulm Presented by Justyna Monika Szamalek from Wrze śnia in Poland 2006 Amtierender Dekan: Prof. Dr. Klaus-Michael Debatin 1. Berichterstatter: Prof. Dr. med. Horst Hameister 2. Berichterstatter: Prof. Dr. med. Konstanze Döhner Tag der Promotion: 28.07.2006 Content Content 1. Introduction ...................................................................................................................7 1.1. Primate phylogeny........................................................................................................7 1.2. Africa as the place of human origin and the living area of the present-day chimpanzee populations .................................................................9 1.3. Cytogenetic and molecular differences between human and chimpanzee genomes.............................................................................................10 1.4. Cytogenetic and molecular differences between common chimpanzee and bonobo genomes................................................................................17 1.5. Theory of speciation .....................................................................................................18 1.6. Theory of selection -

Genomic Sequence Analysis of a Potential QTL Region for Fat Trait on Pig Chromosome 6I
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector Genomics 87 (2006) 218 – 224 www.elsevier.com/locate/ygeno Genomic sequence analysis of a potential QTL region for fat trait on pig chromosome 6i Kyung-Tai Lee a,b,1, Eung-Woo Park a,1, Sunjin Moon c,1, Hye-Sook Park a, Hyung-Yong Kim a, Gil-Won Jang a, Bong-Hwan Choi a, H.Y. Chung a, Ji-Woong Lee a, Il-Cheong Cheong a, Sung-Jong Oh a, Heebal Kim c, Dong-Sang Suh b, Tae-Hun Kim a,* a Division of Animal Genomics and Bioinformatics, National Livestock Research Institute, Rural Development Administration, Omokchun-dong 564, Kwonsun-gu, Suwon, Korea b Department of Genetic Engineering, Sungkyunkwan University, Chunchun-dong 300, Jangan-gu, Suwon, Korea c School of Agricultural Biotechnology, Seoul National University San 56-1, Sillim-dong, Gwanak-gu, Seoul 151-742, Korea Received 1 June 2005; accepted 3 September 2005 Available online 2 December 2005 Abstract On pig chromosome 6, the SW71 microsatellite is located in the region corresponding to several quantitative trait loci (QTL), such as those for intramuscular fat content and for body weight at 4 weeks of age. The genomic sequence of approximately 909 kb was obtained from seven BAC clones encompassing the SW71 region corresponding to human 18q11.21–q11.22. By searching the NCBI GenBank using BLASTX and BLASTN, this 909-kb segment was found to contain eight genes, RAB31, TXNDC2, VAPA, APCDD1, NAPG, FAM38B, C18orf30, and C18orf58, and one putative gene (DN119777). -
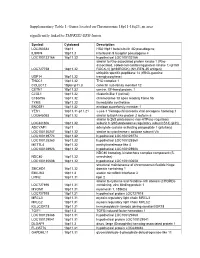
Supplementary Table 1: Genes Located on Chromosome 18P11-18Q23, an Area Significantly Linked to TMPRSS2-ERG Fusion
Supplementary Table 1: Genes located on Chromosome 18p11-18q23, an area significantly linked to TMPRSS2-ERG fusion Symbol Cytoband Description LOC260334 18p11 HSA18p11 beta-tubulin 4Q pseudogene IL9RP4 18p11.3 interleukin 9 receptor pseudogene 4 LOC100132166 18p11.32 hypothetical LOC100132166 similar to Rho-associated protein kinase 1 (Rho- associated, coiled-coil-containing protein kinase 1) (p160 LOC727758 18p11.32 ROCK-1) (p160ROCK) (NY-REN-35 antigen) ubiquitin specific peptidase 14 (tRNA-guanine USP14 18p11.32 transglycosylase) THOC1 18p11.32 THO complex 1 COLEC12 18pter-p11.3 collectin sub-family member 12 CETN1 18p11.32 centrin, EF-hand protein, 1 CLUL1 18p11.32 clusterin-like 1 (retinal) C18orf56 18p11.32 chromosome 18 open reading frame 56 TYMS 18p11.32 thymidylate synthetase ENOSF1 18p11.32 enolase superfamily member 1 YES1 18p11.31-p11.21 v-yes-1 Yamaguchi sarcoma viral oncogene homolog 1 LOC645053 18p11.32 similar to BolA-like protein 2 isoform a similar to 26S proteasome non-ATPase regulatory LOC441806 18p11.32 subunit 8 (26S proteasome regulatory subunit S14) (p31) ADCYAP1 18p11 adenylate cyclase activating polypeptide 1 (pituitary) LOC100130247 18p11.32 similar to cytochrome c oxidase subunit VIc LOC100129774 18p11.32 hypothetical LOC100129774 LOC100128360 18p11.32 hypothetical LOC100128360 METTL4 18p11.32 methyltransferase like 4 LOC100128926 18p11.32 hypothetical LOC100128926 NDC80 homolog, kinetochore complex component (S. NDC80 18p11.32 cerevisiae) LOC100130608 18p11.32 hypothetical LOC100130608 structural maintenance -

NAPG Mutation in Family Members with Hereditary Hemorrhagic Telangiectasia in China
Xu et al. BMC Pulm Med (2021) 21:197 https://doi.org/10.1186/s12890-021-01524-4 RESEARCH ARTICLE Open Access NAPG mutation in family members with hereditary hemorrhagic telangiectasia in China Yu Xu1†, Yong‑Biao Zhang2†, Li‑Jun Liang1, Jia‑Li Tian1, Jin‑Ming Lin3, Pan‑Pan Wang3, Rong‑Hui Li3, Ming‑Liang Gu3,4* and Zhan‑Cheng Gao1* Abstract Background: Hereditary hemorrhagic telangiectasia (HHT) is a disease characterized by arteriovenous malforma‑ tions in the skin and mucous membranes. We enrolled a large pedigree comprising 32 living members, and screened for mutations responsible for HHT. Methods: We performed whole‑exome sequencing to identify novel mutations in the pedigree after excluding three previously reported HHT‑related genes using Sanger sequencing. We then performed in silico functional analysis of candidate mutations that were obtained using a variant fltering strategy to identify mutations responsible for HHT. Results: After screening the HHT‑related genes, activin A receptor‑like type 1 (ACVRL1), endoglin (ENG), and SMAD family member 4 (SMAD4), we did not detect any co‑segregated mutations in this pedigree. Whole‑exome sequencing analysis of 7 members and Sanger sequencing analysis of 16 additional members identifed a mutation (c.784A > G) in the NSF attachment protein gamma (NAPG) gene that co‑segregated with the disease. Functional prediction showed that the mutation was deleterious and might change the conformational stability of the NAPG protein. Conclusions: NAPG c.784A > G may potentially lead to HHT. These results expand the current understanding of the genetic contributions to HHT pathogenesis. Keywords: Hereditary hemorrhagic telangiectasia, Whole‑exome sequencing, NAPG Background membranes. -

Laboratory Investigation of Platelet Function in Patients with Mild Bleeding Disorders
LABORATORY INVESTIGATION OF PLATELET FUNCTION IN PATIENTS WITH MILD BLEEDING DISORDERS by Rashid Hafidh Rashid Al Ghaithi A thesis submitted to the University of Birmingham for the degree of DOCTOR OF PHILOSOPHY Institute of Inflammation and Ageing College of Medical and Dental Sciences University of Birmingham January 2018 University of Birmingham Research Archive e-theses repository This unpublished thesis/dissertation is copyright of the author and/or third parties. The intellectual property rights of the author or third parties in respect of this work are as defined by The Copyright Designs and Patents Act 1988 or as modified by any successor legislation. Any use made of information contained in this thesis/dissertation must be in accordance with that legislation and must be properly acknowledged. Further distribution or reproduction in any format is prohibited without the permission of the copyright holder. Abstract ABSTRACT Platelets play a crucial role in haemostasis by preventing bleeding at sites of vascular injury. Inherited or acquired platelet defects can impair haemostasis resulting in bleeding symptoms of varying severity ranging from mild to excessive which can be life threatening. Diagnosis of mild platelet-based bleeding disorders is challenging due to the absence of a gold standard technique and their variable bleeding symptoms and bleeding phenotypes observed in healthy individual as well as other haemostatic disorders. Furthermore their bleeding symptoms often only manifest after haemostatic challenge. The work in this thesis built on the previous studies in the genotyping and platelet phenotyping project allowing further characterization of inherited platelet function defects in individuals with mild bleeding disorders. The bleeding symptoms of patients were evaluated using the bleeding assessment tool, and its likelihood in diagnosing platelet function defects was assessed and recorded. -

Contextual Analysis of Large-Scale Biomedical Associations for the Elucidation and Prioritization of Genes and Their Roles in Complex Disease Jeremy J
The University of Maine DigitalCommons@UMaine Electronic Theses and Dissertations Fogler Library 12-2013 Contextual Analysis of Large-Scale Biomedical Associations for the Elucidation and Prioritization of Genes and their Roles in Complex Disease Jeremy J. Jay Follow this and additional works at: http://digitalcommons.library.umaine.edu/etd Part of the Computer Sciences Commons Recommended Citation Jay, Jeremy J., "Contextual Analysis of Large-Scale Biomedical Associations for the Elucidation and Prioritization of Genes and their Roles in Complex Disease" (2013). Electronic Theses and Dissertations. 2140. http://digitalcommons.library.umaine.edu/etd/2140 This Open-Access Dissertation is brought to you for free and open access by DigitalCommons@UMaine. It has been accepted for inclusion in Electronic Theses and Dissertations by an authorized administrator of DigitalCommons@UMaine. CONTEXTUAL ANALYSIS OF LARGE-SCALE BIOMEDICAL ASSOCIATIONS FOR THE ELUCIDATION AND PRIORITIZATION OF GENES AND THEIR ROLES IN COMPLEX DISEASE By Jeremy J. Jay B.S.I. Baylor University, 2006 M.S. University of Tennessee, 2009 A DISSERTATION Submitted in Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy (in Computer Science) The Graduate School The University of Maine December 2013 Advisory Committee: George Markowsky, Professor, Advisor Elissa J Chesler, Associate Professor, The Jackson Laboratory Erich J Baker, Associate Professor, Baylor University Judith Blake, Associate Professor, The Jackson Laboratory James Fastook, Professor DISSERTATION ACCEPTANCE STATEMENT On behalf of the Graduate Committee for Jeremy J. Jay, I affirm that this manuscript is the final and accepted dissertation. Signatures of all committee members are on file with the Graduate School at the University of Maine, 42 Stodder Hall, Orono, Maine. -

Patterns of Sequence Conservation in Presynaptic Neural Genes
University of Pennsylvania ScholarlyCommons Departmental Papers (CIS) Department of Computer & Information Science November 2006 Patterns of Sequence Conservation in Presynaptic Neural Genes Dexter Hadley University of Pennsylvania Tara Murphy University of Pennsylvania Otto Valladares University of Pennsylvania Sridhar Hannenhalli University of Pennsylvania Lyle H. Ungar University of Pennsylvania, [email protected] See next page for additional authors Follow this and additional works at: https://repository.upenn.edu/cis_papers Recommended Citation Dexter Hadley, Tara Murphy, Otto Valladares, Sridhar Hannenhalli, Lyle H. Ungar, Junhyong Kim, and Maja Bucan, "Patterns of Sequence Conservation in Presynaptic Neural Genes", . November 2006. Reprinted from Genome Biology, Volume 7, Issue 11, November 2006, pages R105.1-R105.19. Publisher URL: http://genomebiology.com/2006/7/11/R105 This paper is posted at ScholarlyCommons. https://repository.upenn.edu/cis_papers/282 For more information, please contact [email protected]. Patterns of Sequence Conservation in Presynaptic Neural Genes Abstract Background: The neuronal synapse is a fundamental functional unit in the central nervous system of animals. Because synaptic function is evolutionarily conserved, we reasoned that functional sequences of genes and related genomic elements known to play important roles in neurotransmitter release would also be conserved. Results: Evolutionary rate analysis revealed that presynaptic proteins evolve slowly, although some members of large gene families exhibit accelerated evolutionary rates relative to other family members. Comparative sequence analysis of 46 megabases spanning 150 presynaptic genes identified more than 26,000 elements that are highly conserved in eight vertebrate species, as well as a small subset of sequences (6%) that are shared among unrelated presynaptic genes.