Laboratory Investigation of Platelet Function in Patients with Mild Bleeding Disorders
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Analysis of Trans Esnps Infers Regulatory Network Architecture
Analysis of trans eSNPs infers regulatory network architecture Anat Kreimer Submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Graduate School of Arts and Sciences COLUMBIA UNIVERSITY 2014 © 2014 Anat Kreimer All rights reserved ABSTRACT Analysis of trans eSNPs infers regulatory network architecture Anat Kreimer eSNPs are genetic variants associated with transcript expression levels. The characteristics of such variants highlight their importance and present a unique opportunity for studying gene regulation. eSNPs affect most genes and their cell type specificity can shed light on different processes that are activated in each cell. They can identify functional variants by connecting SNPs that are implicated in disease to a molecular mechanism. Examining eSNPs that are associated with distal genes can provide insights regarding the inference of regulatory networks but also presents challenges due to the high statistical burden of multiple testing. Such association studies allow: simultaneous investigation of many gene expression phenotypes without assuming any prior knowledge and identification of unknown regulators of gene expression while uncovering directionality. This thesis will focus on such distal eSNPs to map regulatory interactions between different loci and expose the architecture of the regulatory network defined by such interactions. We develop novel computational approaches and apply them to genetics-genomics data in human. We go beyond pairwise interactions to define network motifs, including regulatory modules and bi-fan structures, showing them to be prevalent in real data and exposing distinct attributes of such arrangements. We project eSNP associations onto a protein-protein interaction network to expose topological properties of eSNPs and their targets and highlight different modes of distal regulation. -

Defining Functional Interactions During Biogenesis of Epithelial Junctions
ARTICLE Received 11 Dec 2015 | Accepted 13 Oct 2016 | Published 6 Dec 2016 | Updated 5 Jan 2017 DOI: 10.1038/ncomms13542 OPEN Defining functional interactions during biogenesis of epithelial junctions J.C. Erasmus1,*, S. Bruche1,*,w, L. Pizarro1,2,*, N. Maimari1,3,*, T. Poggioli1,w, C. Tomlinson4,J.Lees5, I. Zalivina1,w, A. Wheeler1,w, A. Alberts6, A. Russo2 & V.M.M. Braga1 In spite of extensive recent progress, a comprehensive understanding of how actin cytoskeleton remodelling supports stable junctions remains to be established. Here we design a platform that integrates actin functions with optimized phenotypic clustering and identify new cytoskeletal proteins, their functional hierarchy and pathways that modulate E-cadherin adhesion. Depletion of EEF1A, an actin bundling protein, increases E-cadherin levels at junctions without a corresponding reinforcement of cell–cell contacts. This unexpected result reflects a more dynamic and mobile junctional actin in EEF1A-depleted cells. A partner for EEF1A in cadherin contact maintenance is the formin DIAPH2, which interacts with EEF1A. In contrast, depletion of either the endocytic regulator TRIP10 or the Rho GTPase activator VAV2 reduces E-cadherin levels at junctions. TRIP10 binds to and requires VAV2 function for its junctional localization. Overall, we present new conceptual insights on junction stabilization, which integrate known and novel pathways with impact for epithelial morphogenesis, homeostasis and diseases. 1 National Heart and Lung Institute, Faculty of Medicine, Imperial College London, London SW7 2AZ, UK. 2 Computing Department, Imperial College London, London SW7 2AZ, UK. 3 Bioengineering Department, Faculty of Engineering, Imperial College London, London SW7 2AZ, UK. 4 Department of Surgery & Cancer, Faculty of Medicine, Imperial College London, London SW7 2AZ, UK. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Communication Pathways in Human Nonmuscle Myosin-2C 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 Authors: 25 Krishna Chinthalapudia,B,C,1, Sarah M
1 Mechanistic Insights into the Active Site and Allosteric 2 Communication Pathways in Human Nonmuscle Myosin-2C 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 Authors: 25 Krishna Chinthalapudia,b,c,1, Sarah M. Heisslera,d,1, Matthias Prellera,e, James R. Sellersd,2, and 26 Dietmar J. Mansteina,b,2 27 28 Author Affiliations 29 aInstitute for Biophysical Chemistry, OE4350 Hannover Medical School, 30625 Hannover, 30 Germany. 31 bDivision for Structural Biochemistry, OE8830, Hannover Medical School, 30625 Hannover, 32 Germany. 33 cCell Adhesion Laboratory, Department of Integrative Structural and Computational Biology, The 34 Scripps Research Institute, Jupiter, Florida 33458, USA. 35 dLaboratory of Molecular Physiology, NHLBI, National Institutes of Health, Bethesda, Maryland 36 20892, USA. 37 eCentre for Structural Systems Biology (CSSB), German Electron Synchrotron (DESY), 22607 38 Hamburg, Germany. 39 1K.C. and S.M.H. contributed equally to this work 40 2To whom correspondence may be addressed: E-mail: [email protected] or 41 [email protected] 42 43 1 44 Abstract 45 Despite a generic, highly conserved motor domain, ATP turnover kinetics and their activation by 46 F-actin vary greatly between myosin-2 isoforms. Here, we present a 2.25 Å crystal pre- 47 powerstroke state (ADPVO4) structure of the human nonmuscle myosin-2C motor domain, one 48 of the slowest myosins characterized. In combination with integrated mutagenesis, ensemble- 49 solution kinetics, and molecular dynamics simulation approaches, the structure reveals an 50 allosteric communication pathway that connects the distal end of the motor domain with the 51 active site. -

Insights from Conventional and Unconventional Myosins A
Structure-Function Analysis of Motor Proteins: Insights from Conventional and Unconventional Myosins A Thesis SUBMITTED TO THE FACULTY OF UNIVERSITY OF MINNESOTA BY Karl J. Petersen IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE OF DOCTOR OF PHILOSOPHY Margaret A. Titus, Advisor David D. Thomas, Co-Advisor December 2016 © Karl J. Petersen 2016 Acknowledgements This thesis would not exist without the patient support of my advisors, Meg Titus and Dave Thomas. Any shortcomings are my own. Collaborators Holly Goodson, Anne Houdusse, and Gant Luxton also provided invaluable training and support. I am also grateful for essential services provided by departmental staff: Sarah Blakely Anderson, Octavian Cornea, Sarah Dittrich, Karen Hawkinson, Michelle Lewis, Mary Muwahid, Laurie O’Neill, Darlene Toedter, with apologies to others not listed. Thanks to friends and colleagues at the University of Minnesota: Ashley Arthur, Kelly Bower, Brett Colson, Sinziana Cornea, Chi Meng Fong, Greg Gillispie, Piyali Guhathakurta, Tejas Gupte, Tom Hays, Norma Jiménez Ramírez, Dawn Lowe, Allison MacLean, Santiago Martínez Cifuentes, Jared Matzke, Megan McCarthy, Joachim Mueller, Joe Muretta, Kurt Peterson, Mary Porter, Ewa Prochniewicz, Mike Ritt, Cosmo Saunders, Shiv Sivaramakrishnan, Ruth Sommese, Doug Tritschler, Brian Woolums. i Abstract Myosin motor proteins play fundamental roles in a multitude of cellular processes. Myosin generates force on cytoskeletal actin filaments to control cell shape, most dramatically during cytokinesis, and has a conserved role in defining cell polarity. Myosin contracts the actin cytoskeleton, ensuring prompt turnover of cellular adhesion sites, retracting the cell body during migration and development, and contracting muscle among diverse other functions. How myosins work, and why force generation is essential for their function, is in many cases an open question. -

A New Efficient Method to Detect Genetic Interactions for Lung Cancer
Luyapan et al. BMC Med Genomics (2020) 13:162 https://doi.org/10.1186/s12920-020-00807-9 TECHNICAL ADVANCE Open Access A new efcient method to detect genetic interactions for lung cancer GWAS Jennifer Luyapan1,2, Xuemei Ji2, Siting Li1,2, Xiangjun Xiao3, Dakai Zhu2,3, Eric J. Duell4, David C. Christiani5,6, Matthew B. Schabath7, Susanne M. Arnold8, Shanbeh Zienolddiny9, Hans Brunnström10, Olle Melander11, Mark D. Thornquist12, Todd A. MacKenzie1,2, Christopher I. Amos1,2,3* and Jiang Gui1,2* Abstract Background: Genome-wide association studies (GWAS) have proven successful in predicting genetic risk of disease using single-locus models; however, identifying single nucleotide polymorphism (SNP) interactions at the genome- wide scale is limited due to computational and statistical challenges. We addressed the computational burden encountered when detecting SNP interactions for survival analysis, such as age of disease-onset. To confront this problem, we developed a novel algorithm, called the Efcient Survival Multifactor Dimensionality Reduction (ES-MDR) method, which used Martingale Residuals as the outcome parameter to estimate survival outcomes, and imple- mented the Quantitative Multifactor Dimensionality Reduction method to identify signifcant interactions associated with age of disease-onset. Methods: To demonstrate efcacy, we evaluated this method on two simulation data sets to estimate the type I error rate and power. Simulations showed that ES-MDR identifed interactions using less computational workload and allowed for adjustment of covariates. We applied ES-MDR on the OncoArray-TRICL Consortium data with 14,935 cases and 12,787 controls for lung cancer (SNPs 108,254) to search over all two-way interactions to identify genetic interactions associated with lung cancer age-of-onset.= We tested the best model in an independent data set from the OncoArray-TRICL data. -

The Genetics of Bipolar Disorder
Molecular Psychiatry (2008) 13, 742–771 & 2008 Nature Publishing Group All rights reserved 1359-4184/08 $30.00 www.nature.com/mp FEATURE REVIEW The genetics of bipolar disorder: genome ‘hot regions,’ genes, new potential candidates and future directions A Serretti and L Mandelli Institute of Psychiatry, University of Bologna, Bologna, Italy Bipolar disorder (BP) is a complex disorder caused by a number of liability genes interacting with the environment. In recent years, a large number of linkage and association studies have been conducted producing an extremely large number of findings often not replicated or partially replicated. Further, results from linkage and association studies are not always easily comparable. Unfortunately, at present a comprehensive coverage of available evidence is still lacking. In the present paper, we summarized results obtained from both linkage and association studies in BP. Further, we indicated new potential interesting genes, located in genome ‘hot regions’ for BP and being expressed in the brain. We reviewed published studies on the subject till December 2007. We precisely localized regions where positive linkage has been found, by the NCBI Map viewer (http://www.ncbi.nlm.nih.gov/mapview/); further, we identified genes located in interesting areas and expressed in the brain, by the Entrez gene, Unigene databases (http://www.ncbi.nlm.nih.gov/entrez/) and Human Protein Reference Database (http://www.hprd.org); these genes could be of interest in future investigations. The review of association studies gave interesting results, as a number of genes seem to be definitively involved in BP, such as SLC6A4, TPH2, DRD4, SLC6A3, DAOA, DTNBP1, NRG1, DISC1 and BDNF. -

In This Table Protein Name, Uniprot Code, Gene Name P-Value
Supplementary Table S1: In this table protein name, uniprot code, gene name p-value and Fold change (FC) for each comparison are shown, for 299 of the 301 significantly regulated proteins found in both comparisons (p-value<0.01, fold change (FC) >+/-0.37) ALS versus control and FTLD-U versus control. Two uncharacterized proteins have been excluded from this list Protein name Uniprot Gene name p value FC FTLD-U p value FC ALS FTLD-U ALS Cytochrome b-c1 complex P14927 UQCRB 1.534E-03 -1.591E+00 6.005E-04 -1.639E+00 subunit 7 NADH dehydrogenase O95182 NDUFA7 4.127E-04 -9.471E-01 3.467E-05 -1.643E+00 [ubiquinone] 1 alpha subcomplex subunit 7 NADH dehydrogenase O43678 NDUFA2 3.230E-04 -9.145E-01 2.113E-04 -1.450E+00 [ubiquinone] 1 alpha subcomplex subunit 2 NADH dehydrogenase O43920 NDUFS5 1.769E-04 -8.829E-01 3.235E-05 -1.007E+00 [ubiquinone] iron-sulfur protein 5 ARF GTPase-activating A0A0C4DGN6 GIT1 1.306E-03 -8.810E-01 1.115E-03 -7.228E-01 protein GIT1 Methylglutaconyl-CoA Q13825 AUH 6.097E-04 -7.666E-01 5.619E-06 -1.178E+00 hydratase, mitochondrial ADP/ATP translocase 1 P12235 SLC25A4 6.068E-03 -6.095E-01 3.595E-04 -1.011E+00 MIC J3QTA6 CHCHD6 1.090E-04 -5.913E-01 2.124E-03 -5.948E-01 MIC J3QTA6 CHCHD6 1.090E-04 -5.913E-01 2.124E-03 -5.948E-01 Protein kinase C and casein Q9BY11 PACSIN1 3.837E-03 -5.863E-01 3.680E-06 -1.824E+00 kinase substrate in neurons protein 1 Tubulin polymerization- O94811 TPPP 6.466E-03 -5.755E-01 6.943E-06 -1.169E+00 promoting protein MIC C9JRZ6 CHCHD3 2.912E-02 -6.187E-01 2.195E-03 -9.781E-01 Mitochondrial 2- -

Human Induced Pluripotent Stem Cell–Derived Podocytes Mature Into Vascularized Glomeruli Upon Experimental Transplantation
BASIC RESEARCH www.jasn.org Human Induced Pluripotent Stem Cell–Derived Podocytes Mature into Vascularized Glomeruli upon Experimental Transplantation † Sazia Sharmin,* Atsuhiro Taguchi,* Yusuke Kaku,* Yasuhiro Yoshimura,* Tomoko Ohmori,* ‡ † ‡ Tetsushi Sakuma, Masashi Mukoyama, Takashi Yamamoto, Hidetake Kurihara,§ and | Ryuichi Nishinakamura* *Department of Kidney Development, Institute of Molecular Embryology and Genetics, and †Department of Nephrology, Faculty of Life Sciences, Kumamoto University, Kumamoto, Japan; ‡Department of Mathematical and Life Sciences, Graduate School of Science, Hiroshima University, Hiroshima, Japan; §Division of Anatomy, Juntendo University School of Medicine, Tokyo, Japan; and |Japan Science and Technology Agency, CREST, Kumamoto, Japan ABSTRACT Glomerular podocytes express proteins, such as nephrin, that constitute the slit diaphragm, thereby contributing to the filtration process in the kidney. Glomerular development has been analyzed mainly in mice, whereas analysis of human kidney development has been minimal because of limited access to embryonic kidneys. We previously reported the induction of three-dimensional primordial glomeruli from human induced pluripotent stem (iPS) cells. Here, using transcription activator–like effector nuclease-mediated homologous recombination, we generated human iPS cell lines that express green fluorescent protein (GFP) in the NPHS1 locus, which encodes nephrin, and we show that GFP expression facilitated accurate visualization of nephrin-positive podocyte formation in -
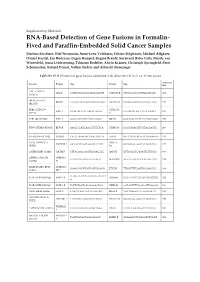
RNA-Based Detection of Gene Fusions in Formalin- Fixed And
Supplementary Materials RNA-Based Detection of Gene Fusions in Formalin- Fixed and Paraffin-Embedded Solid Cancer Samples Martina Kirchner, Olaf Neumann, Anna-Lena Volckmar, Fabian Stögbauer, Michael Allgäuer, Daniel Kazdal, Jan Budczies, Eugen Rempel, Regine Brandt, Suranand Babu Talla, Moritz von Winterfeld, Jonas Leichsenring, Tilmann Bochtler, Alwin Krämer, Christoph Springfeld, Peter Schirmacher, Roland Penzel, Volker Endris and Albrecht Stenzinger Table S1. PCR Primers for gene fusions identified with either the OCAv3- or Archer-panel. Amplicon Fusion Primer Seq Primer Seq [bp] AXL::CAPN15 AXL F CATGGATGAGGGTGGAGGTT CAPN15 R CTGGGCACACGTGAATCAC 178 (A19C2) BRD3::NUTM1 BRD3 F AAGAAACAGGCAGCCAAGTC NUTM1 R CTGGTGGGTCAGAAGTTGGT 217 (B11N2) ESR1-CCDC170 CCDC170 ESR1 F GGAGACTCGCTACTGTGCA CCCAGACTCCTTTCCCAACT 167 (E2C7) R ESR1-QKI (E2Q5) ESR1 F GGAGACTCGCTACTGTGCA QKI R GGCTGGTGATTTAATGTTGGC 197 ETV6::NTRK3 (E5N15) ETV6 F AAGCCCATCAACCTCTCTCA NTRK3 R GGGCTGAGGTTGTAGCACTC 206 FGFR2::INA (F17I2) FGFR2 F CTCCCAGAGACCAACGTTCA INA R GTCCTGGTATTCCCGAAAGGT 148 FNDC3B-PIK3CA PIK3CA FNDC3B F GCAGCTCAGCAGGTTATTCT GTCGTGGAGGCATTGTTCTG 177 (F3P2) R1 GATM::RAF1 (G2R8) GATM F CTTACAACGAATGGGACCCC RAF1 R GTTGGGCTCAGATTGTTGGG 160 GPBP1L1::MAST2 GPBP1L1 CGTAGTGGAGGTGGCACA MAST2 R1 AGGTGATGTGCTAGAGGTCA 178 (G6M4) F1 HNRNPA2B1::ETV1 HNRNPA GGAGGATATGGTGGTGGAGG ETV1 R TTGATTTTCAGTGGCAGGCC 164 (H9E6) 2B1 F TGATGAATCTGGAATTGTTGCT MYB::NFIB (M12N9) MYB12 F NFIB9 R CGTAATTTTGGACATTGGCCG 150 G MYB::NFIB (M13N9) MYB13 F TCTTCTGCTCACACCACTGG NFIB9 R CGTAATTTTGGACATTGGCCG 160 SND1::BRAF (S9B9) SND1 F CGATTCACCTGTCCAGCATC BRAF R CGCTGAGGTCCTGGAGATTT 184 TBL1XR1::PIK3CA TBL1XR1 F TTTCCTTGTGCCTCCATTCC PIK3CA R GTCGTGGAGGCATTGTTCTG 195 (T1P2) TMPRSS2 TMPRSS2::ERG (T2E4) CGCGGCAGGTCATATTGAA ERG R CCTTCCCATCGATGTTCTGG 190 F WHSC1L1::FGFR1 WHSC1L1 TGATCGCACTGACACGGC FGFR1 R ACAAGGCTCCACATCTCCAT 108 (W1F2) F Table S2. Clinical and diagnostic implications of the detected gene fusions. Reclassification Entities Where Entity Where Fusion Is Based on Molecular Fusion Is Fusion Drug Ref. -

SARS-Cov-2 Infection of Human Ipsc-Derived Cardiac Cells Predicts Novel Cytopathic Features in Hearts of COVID-19 Patients
bioRxiv preprint doi: https://doi.org/10.1101/2020.08.25.265561; this version posted September 12, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-ND 4.0 International license. SARS-CoV-2 infection of human iPSC-derived cardiac cells predicts novel cytopathic features in hearts of COVID-19 patients Juan A. Pérez-Bermejo*1, Serah Kang*1, Sarah J. Rockwood*1, Camille R. Simoneau*1,2, David A. Joy1,3, Gokul N. Ramadoss1,2, Ana C. Silva1, Will R. Flanigan1,3, Huihui Li1, Ken Nakamura1,4, Jeffrey D. Whitman5, Melanie Ott†,1, Bruce R. Conklin†,1,6,7,8, Todd C. McDevitt†1,9 * These authors contributed equally to this work. † Co-corresponding authors. 1 Gladstone Institutes, San Francisco, CA 2 Biomedical Sciences PhD Program, University of California, San Francisco, CA 3 UC Berkeley UCSF Joint Program in Bioengineering, Berkeley, CA 4 UCSF Department of Neurology, San Francisco, CA 5 UCSF Department of Laboratory Medicine, San Francisco, CA 6 Innovative Genomics Institute, Berkeley, CA 7 UCSF Department of Ophthalmology, San Francisco, CA 8 UCSF Department of Medicine, San Francisco, CA 9 UCSF Department of Bioengineering and Therapeutic Sciences, San Francisco, CA ABSTRACT Although COVID-19 causes cardiac dysfunction in up to 25% of patients, its pathogenesis remains unclear. Exposure of human iPSC-derived heart cells to SARS-CoV-2 revealed productive infection and robust transcriptomic and morphological signatures of damage, particularly in cardiomyocytes. -

Cytogenetic and Molecular Characterization of the Macro- And
University of Ulm Department of Human Genetics Prof. Dr. med. Walther Vogel Cytogenetic and Molecular Characterization of the Macro- and Micro-inversions, which Distinguish the Human and the Chimpanzee Karyotypes - from Speciation to Polymorphism Thesis Applying for the Degree of Doctor of Human Biology (Dr. hum. biol.) Faculty of Medicine University of Ulm Presented by Justyna Monika Szamalek from Wrze śnia in Poland 2006 Amtierender Dekan: Prof. Dr. Klaus-Michael Debatin 1. Berichterstatter: Prof. Dr. med. Horst Hameister 2. Berichterstatter: Prof. Dr. med. Konstanze Döhner Tag der Promotion: 28.07.2006 Content Content 1. Introduction ...................................................................................................................7 1.1. Primate phylogeny........................................................................................................7 1.2. Africa as the place of human origin and the living area of the present-day chimpanzee populations .................................................................9 1.3. Cytogenetic and molecular differences between human and chimpanzee genomes.............................................................................................10 1.4. Cytogenetic and molecular differences between common chimpanzee and bonobo genomes................................................................................17 1.5. Theory of speciation .....................................................................................................18 1.6. Theory of selection