Contextual Analysis of Large-Scale Biomedical Associations for the Elucidation and Prioritization of Genes and Their Roles in Complex Disease Jeremy J
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Deletion of the SNARE Vti1b in Mice Results in the Loss of a Single
MOLECULAR AND CELLULAR BIOLOGY, Aug. 2003, p. 5198–5207 Vol. 23, No. 15 0270-7306/03/$08.00ϩ0 DOI: 10.1128/MCB.23.15.5198–5207.2003 Copyright © 2003, American Society for Microbiology. All Rights Reserved. Deletion of the SNARE vti1b in Mice Results in the Loss of Downloaded from a Single SNARE Partner, Syntaxin 8 Vadim Atlashkin,1 Vera Kreykenbohm,1 Eeva-Liisa Eskelinen,2 Dirk Wenzel,3 Afshin Fayyazi,4 and Gabriele Fischer von Mollard1* Zentrum Biochemie und Molekulare Zellbiologie, Abteilung Biochemie II,1 and Abteilung Pathologie,4 Universita¨t Go¨ttingen, and Abteilung Neurobiologie, Max-Planck Institut fu¨r Biophysikalische Chemie,3 Go¨ttingen, and http://mcb.asm.org/ Biochemisches Institut, Universita¨t Kiel, Kiel,2 Germany Received 13 February 2003/Accepted 26 April 2003 SNARE proteins participate in recognition and fusion of membranes. A SNARE complex consisting of vti1b, syntaxin 8, syntaxin 7, and endobrevin/VAMP-8 which is required for fusion of late endosomes in vitro has been identified recently. Here, we generated mice deficient in vti1b to study the function of this protein in vivo. vti1b-deficient mice had reduced amounts of syntaxin 8 due to degradation of the syntaxin 8 protein, while the amounts of syntaxin 7 and endobrevin did not change. These data indicate that vti1b is specifically required for the stability of a single SNARE partner. vti1b-deficient mice were viable and fertile. Most vti1b-deficient on February 22, 2016 by MAX PLANCK INSTITUT F BIOPHYSIKALISCHE CHEMIE mice were indistinguishable from wild-type mice and did not display defects in transport to the lysosome. -

Analysis of Trans Esnps Infers Regulatory Network Architecture
Analysis of trans eSNPs infers regulatory network architecture Anat Kreimer Submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Graduate School of Arts and Sciences COLUMBIA UNIVERSITY 2014 © 2014 Anat Kreimer All rights reserved ABSTRACT Analysis of trans eSNPs infers regulatory network architecture Anat Kreimer eSNPs are genetic variants associated with transcript expression levels. The characteristics of such variants highlight their importance and present a unique opportunity for studying gene regulation. eSNPs affect most genes and their cell type specificity can shed light on different processes that are activated in each cell. They can identify functional variants by connecting SNPs that are implicated in disease to a molecular mechanism. Examining eSNPs that are associated with distal genes can provide insights regarding the inference of regulatory networks but also presents challenges due to the high statistical burden of multiple testing. Such association studies allow: simultaneous investigation of many gene expression phenotypes without assuming any prior knowledge and identification of unknown regulators of gene expression while uncovering directionality. This thesis will focus on such distal eSNPs to map regulatory interactions between different loci and expose the architecture of the regulatory network defined by such interactions. We develop novel computational approaches and apply them to genetics-genomics data in human. We go beyond pairwise interactions to define network motifs, including regulatory modules and bi-fan structures, showing them to be prevalent in real data and exposing distinct attributes of such arrangements. We project eSNP associations onto a protein-protein interaction network to expose topological properties of eSNPs and their targets and highlight different modes of distal regulation. -

CHRISTINA “TINA” WARINNER (Last Updated October 18, 2018)
CHRISTINA “TINA” WARINNER (last updated October 18, 2018) Max Planck Institute University of Oklahoma for the Science of Human History (MPI-SHH) Department of Anthropology Department of Archaeogenetics Laboratories of Molecular Anthropology Kahlaische Strasse 10, 07743 Jena, Germany And Microbiome Research (LMAMR) +49 3641686620 101 David L. Boren Blvd, [email protected] Norman, OK 73019 USA www.christinawarinner.com [email protected] http://www.shh.mpg.de/employees/50506/25522 www.lmamr.org APPOINTMENTS W2 Group Leader, Max Planck Institute for the Science of Human History, Germany 2016-present University Professor, Friedrich Schiller University, Jena, Germany 2018-present Presidential Research Professor, Univ. of Oklahoma, USA 2014-present Assistant Professor, Dept. of Anthropology, Univ. of Oklahoma, USA 2014-present Adjunct Professor, Dept. of Periodontics, College of Dentistry, Univ. of Oklahoma, USA 2014-present Visiting Associate Professor, Dept. of Systems Biology, Technical University of Denmark 2015 Research Associate, Dept. of Anthropology, Univ. of Oklahoma, USA 2012-2014 Acting Head of Group, Centre for Evolutionary Medicine, Univ. of Zürich, Switzerland 2011-2012 Research Assistant, Centre for Evolutionary Medicine, Univ. of Zürich, Switzerland 2010-2011 EDUCATION Ph.D., Anthropology, Dept. of Anthropology, Harvard University 2010 Thesis Title: “Life and Death at Teposcolula Yucundaa: Mortuary, Archaeogenetic, and Isotopic Investigations of the Early Colonial Period in Mexico” A.M., Anthropology, Dept. of Anthropology, Harvard University 2008 B.A., with Honors, Anthropology, University of Kansas 2003 B.A., Germanic Literatures and Languages, University of Kansas 2003 SELECTED HONORS, AWARDS, AND FELLOWSHIPS Invited speaker, British Academy, Albert Reckitt Archaeological Lecture (forthcoming) 2019 Invited speaker, EMBL Science and Society (forthcoming, Nov. -

Ayshwarya Subramanian Updated June 10, 2021
Ayshwarya Subramanian Updated June 10, 2021 Contact Kuchroo Lab and Klarman Cell Observatory Information Broad Institute twitter: @ayshwaryas 5407G, 415 Main Street website: ayshwaryas.github.io Cambridge, MA 02412 USA email:[email protected] Areas of Computational Biology (Kidney disease, Cancer, Inflammatory Bowel Disease, RNA Biology), Ge- expertise nomic Data Analysis (Single-cell and Bulk-RNAseq, Metagenomics, Exome and single-nucleus DNA sequencing), Machine Learning, Probabilistic modeling, Phylogenetics, Applied Statistics Education 2013 Ph.D., Biological Sciences Carnegie Mellon University, Pittsburgh, PA USA Doctoral Advisor: Russell Schwartz, Ph.D. Dissertation: Inferring tumor evolution using computational phylogenetics 2007 M.Sc. (Hons), Biological Sciences (Undergraduate degree) Birla Institute of Technology and Science (BITS{Pilani), Rajasthan, India CGPA 9.21/10, Major GPA 10/10, with Distinction Undergraduate Honors Thesis: A mathematical model for phototactic responses in Halobac- terium salinarium, Max Planck Institute for Complex Technical Systems, Germany. Current Computational Scientist, Cambridge MA 2017{Present Appointment Mentors: Aviv Regev, Ph.D. & Vijay Kuchroo, Ph.D., D.V.M Research Summary: Single-cell portraits of disease and normal states using human data, mouse and organoid models Klarman Cell Observatory Broad Institute, Cambridge, MA 02142 Publications Pre-prints/Under review [1] Subramanian A†, Vernon KA†, Zhou Y† et al. Obesity-instructed TREM2high macrophages identified by comparative analysis of diabetic mouse and human kidney at single cell resolution. bioRxiv 2021.05.30.446342; doi: https://doi.org/10.1101/2021.05.30.446342. [2] Subramanian A†,Vernon KA†, Slyper M, Waldman J, et al. RAAS blockade, kidney dis- ease, and expression of ACE2, the entry receptor for SARS-CoV-2, in kidney epithelial and endothelial cells. -

Case Studies in Bayesian Statistics Workshop 9 Poster Session
Case Studies in Bayesian Statistics Workshop 9 Poster Session Following is a tentative list of posters being presented during the workshop: 1. Edoardo Airoldi, Curtis Huttenhower, Olga Troyanskaya and David Botstein 2. Dipankar Bandyopadhyay, Elizabeth Slate, Debajyoti Sinha, Dikpak Dey and Jyotika Fernandes 3. Brenda Betancourt and Maria-Eglee Perez 4. Sham Bhat, Murali Haran, Julio Molineros and Erick Dewolf 5. Jen-hwa Chu, Merlise A. Clyde and Feng Liang 6. Jason Connor, Scott Berry and Don Berry 7. J. Mark Donovan, Michael R. Elliott and Daniel F. Heitjan 8. Elena A. Erosheva, Donatello Telesca, Ross L. Matsueda and Derek Kreager 9. Xiaodan Fan and Jun S. Liu 10. Jairo A. Fuquene and Luis Raul Pericchi 11. Marti Font, Josep Ginebra, Xavier Puig 12. Isobel Claire Gormley and Thomas Brendan Murphy 13. Cari G. Kaufman and Stephan R. Sain 14. Alex Lenkoski 15. Herbie Lee 16. Fei Liu 17. Jingchen Liu, Xiao-Li Meng, Chih-nan Chen, Margarita Alegria 18. Christian Macaro 19. Il-Chul Moon, Eunice J. Kim and Kathleen M. Carley 20. Christopher Paciorek 21. Susan M. Paddock and Patricia Ebener 22. Nicholas M. Pajewski, L. Thomas Johnson, Thomas Radmer, and Purushottam W. Laud 23. Mark W. Perlin, Joseph B. Kadane, Robin W. Cotton and Alexander Sinelnikov 24. Alicia Quiros, Raquel Montes Diez and Dani Gamerman 25. Eiki Satake and Philip Amato 26. James Scott 27. Russell Steele, Robert Platt and Michelle Ross 28. Alejandro Villagran, Gabriel Huerta, Charles S. Jackson and Mrinal K. Sen 29. Dawn Woodard 30. David C. Wheeler, Lance A. Waller and John O. -

Protein Interaction Network of Alternatively Spliced Isoforms from Brain Links Genetic Risk Factors for Autism
ARTICLE Received 24 Aug 2013 | Accepted 14 Mar 2014 | Published 11 Apr 2014 DOI: 10.1038/ncomms4650 OPEN Protein interaction network of alternatively spliced isoforms from brain links genetic risk factors for autism Roser Corominas1,*, Xinping Yang2,3,*, Guan Ning Lin1,*, Shuli Kang1,*, Yun Shen2,3, Lila Ghamsari2,3,w, Martin Broly2,3, Maria Rodriguez2,3, Stanley Tam2,3, Shelly A. Trigg2,3,w, Changyu Fan2,3, Song Yi2,3, Murat Tasan4, Irma Lemmens5, Xingyan Kuang6, Nan Zhao6, Dheeraj Malhotra7, Jacob J. Michaelson7,w, Vladimir Vacic8, Michael A. Calderwood2,3, Frederick P. Roth2,3,4, Jan Tavernier5, Steve Horvath9, Kourosh Salehi-Ashtiani2,3,w, Dmitry Korkin6, Jonathan Sebat7, David E. Hill2,3, Tong Hao2,3, Marc Vidal2,3 & Lilia M. Iakoucheva1 Increased risk for autism spectrum disorders (ASD) is attributed to hundreds of genetic loci. The convergence of ASD variants have been investigated using various approaches, including protein interactions extracted from the published literature. However, these datasets are frequently incomplete, carry biases and are limited to interactions of a single splicing isoform, which may not be expressed in the disease-relevant tissue. Here we introduce a new interactome mapping approach by experimentally identifying interactions between brain-expressed alternatively spliced variants of ASD risk factors. The Autism Spliceform Interaction Network reveals that almost half of the detected interactions and about 30% of the newly identified interacting partners represent contribution from splicing variants, emphasizing the importance of isoform networks. Isoform interactions greatly contribute to establishing direct physical connections between proteins from the de novo autism CNVs. Our findings demonstrate the critical role of spliceform networks for translating genetic knowledge into a better understanding of human diseases. -

Genome-Wide Approach to Identify Risk Factors for Therapy-Related Myeloid Leukemia
Leukemia (2006) 20, 239–246 & 2006 Nature Publishing Group All rights reserved 0887-6924/06 $30.00 www.nature.com/leu ORIGINAL ARTICLE Genome-wide approach to identify risk factors for therapy-related myeloid leukemia A Bogni1, C Cheng2, W Liu2, W Yang1, J Pfeffer1, S Mukatira3, D French1, JR Downing4, C-H Pui4,5,6 and MV Relling1,6 1Department of Pharmaceutical Sciences, The University of Tennessee, Memphis, TN, USA; 2Department of Biostatistics, The University of Tennessee, Memphis, TN, USA; 3Hartwell Center, The University of Tennessee, Memphis, TN, USA; 4Department of Pathology, The University of Tennessee, Memphis, TN, USA; 5Department of Hematology/Oncology St Jude Children’s Research Hospital, The University of Tennessee, Memphis, TN, USA; and 6Colleges of Medicine and Pharmacy, The University of Tennessee, Memphis, TN, USA Using a target gene approach, only a few host genetic risk therapy increases, the importance of identifying host factors for factors for treatment-related myeloid leukemia (t-ML) have been secondary neoplasms increases. defined. Gene expression microarrays allow for a more 4 genome-wide approach to assess possible genetic risk factors Because DNA microarrays interrogate multiple ( 10 000) for t-ML. We assessed gene expression profiles (n ¼ 12 625 genes in one experiment, they allow for a ‘genome-wide’ probe sets) in diagnostic acute lymphoblastic leukemic cells assessment of genes that may predispose to leukemogenesis. from 228 children treated on protocols that included leukemo- DNA microarray analysis of gene expression has been used to genic agents such as etoposide, 13 of whom developed t-ML. identify distinct expression profiles that are characteristic of Expression of 68 probes, corresponding to 63 genes, was different leukemia subtypes.13,14 Studies using this method have significantly related to risk of t-ML. -

WO 2Ull/13162O Al
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date Χ 1 / A 1 27 October 2011 (27.10.2011) WO 2Ull/13162o Al (51) International Patent Classification: AO, AT, AU, AZ, BA, BB, BG, BH, BR, BW, BY, BZ, C12N 9/02 (2006.01) A61K 38/44 (2006.01) CA, CH, CL, CN, CO, CR, CU, CZ, DE, DK, DM, DO, A61K 38/17 (2006.01) DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, HN, HR, HU, ID, IL, IN, IS, JP, KE, KG, KM, KN, KP, (21) International Application Number: KR, KZ, LA, LC, LK, LR, LS, LT, LU, LY, MA, MD, PCT/EP20 11/056142 ME, MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, (22) International Filing Date: NO, NZ, OM, PE, PG, PH, PL, PT, RO, RS, RU, SC, SD, 18 April 201 1 (18.04.201 1) SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. (25) Filing Language: English (84) Designated States (unless otherwise indicated, for every (26) Publication Langi English kind of regional protection available): ARIPO (BW, GH, (30) Priority Data: GM, KE, LR, LS, MW, MZ, NA, SD, SL, SZ, TZ, UG, 10160368.6 19 April 2010 (19.04.2010) EP ZM, ZW), Eurasian (AM, AZ, BY, KG, KZ, MD, RU, TJ, TM), European (AL, AT, BE, BG, CH, CY, CZ, DE, DK, (71) Applicants (for all designated States except US): MEDI- EE, ES, FI, FR, GB, GR, HR, HU, IE, IS, FT, LT, LU, ZINISCHE UNIVERSITAT INNSBRUCK [AT/AT]; LV, MC, MK, MT, NL, NO, PL, PT, RO, RS, SE, SI, SK, Christoph-Probst-Platz, Innrain 52, A-6020 Innsbruck SM, TR), OAPI (BF, BJ, CF, CG, CI, CM, GA, GN, GQ, (AT). -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Genome Informatics 4–8 September 2002, Wellcome Trust Genome Campus, Hinxton, Cambridge, UK
Comparative and Functional Genomics Comp Funct Genom 2003; 4: 509–514. Published online in Wiley InterScience (www.interscience.wiley.com). DOI: 10.1002/cfg.300 Feature Meeting Highlights: Genome Informatics 4–8 September 2002, Wellcome Trust Genome Campus, Hinxton, Cambridge, UK Jo Wixon1* and Jennifer Ashurst2 1MRC UK HGMP-RC, Hinxton, Cambridge CB10 1SB, UK 2The Wellcome Trust Sanger Institute, Wellcome Trust Genome Campus, Hinxton, Cambridge CB10 1SA, UK *Correspondence to: Abstract Jo Wixon, MRC UK HGMP-RC, Hinxton, Cambridge CB10 We bring you the highlights of the second Joint Cold Spring Harbor Laboratory 1SB, UK. and Wellcome Trust ‘Genome Informatics’ Conference, organized by Ewan Birney, E-mail: [email protected] Suzanna Lewis and Lincoln Stein. There were sessions on in silico data discovery, comparative genomics, annotation pipelines, functional genomics and integrative biology. The conference included a keynote address by Sydney Brenner, who was awarded the 2002 Nobel Prize in Physiology or Medicine (jointly with John Sulston and H. Robert Horvitz) a month later. Copyright 2003 John Wiley & Sons, Ltd. In silico data discovery background set was genes which had a log ratio of ∼0 in liver. Their approach found 17 of 17 known In the first of two sessions on this topic, Naoya promoters with a specificity of 17/28. None of the Hata (Cold Spring Harbor Laboratory, USA) sites they identified was located downstream of a spoke about motif searching for tissue specific TSS and all showed an excess in the foreground promoters. The first step in the process is to sample compared to the background sample. -

Towards a Knowledge Graph for Science
Towards a Knowledge Graph for Science Invited Article∗ Sören Auer Viktor Kovtun Manuel Prinz TIB Leibniz Information Centre for L3S Research Centre, Leibniz TIB Leibniz Information Centre for Science and Technology and L3S University of Hannover Science and Technology Research Centre at University of Hannover, Germany Hannover, Germany Hannover [email protected] [email protected] Hannover, Germany [email protected] Anna Kasprzik Markus Stocker TIB Leibniz Information Centre for TIB Leibniz Information Centre for Science and Technology Science and Technology Hannover, Germany Hannover, Germany [email protected] [email protected] ABSTRACT KEYWORDS The document-centric workflows in science have reached (or al- Knowledge Graph, Science and Technology, Research Infrastructure, ready exceeded) the limits of adequacy. This is emphasized by Libraries, Information Science recent discussions on the increasing proliferation of scientific lit- ACM Reference Format: erature and the reproducibility crisis. This presents an opportu- Sören Auer, Viktor Kovtun, Manuel Prinz, Anna Kasprzik, and Markus nity to rethink the dominant paradigm of document-centric schol- Stocker. 2018. Towards a Knowledge Graph for Science: Invited Article. arly information communication and transform it into knowledge- In WIMS ’18: 8th International Conference on Web Intelligence, Mining and based information flows by representing and expressing informa- Semantics, June 25–27, 2018, Novi Sad, Serbia. ACM, New York, NY, USA, tion through semantically rich, interlinked knowledge graphs. At 6 pages. https://doi.org/10.1145/3227609.3227689 the core of knowledge-based information flows is the creation and evolution of information models that establish a common under- 1 INTRODUCTION standing of information communicated between stakeholders as The communication of scholarly information is document-centric. -
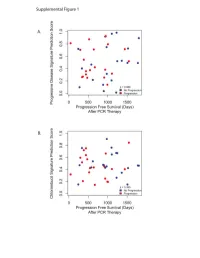
Supplementary Data
Progressive Disease Signature Upregulated probes with progressive disease U133Plus2 ID Gene Symbol Gene Name 239673_at NR3C2 nuclear receptor subfamily 3, group C, member 2 228994_at CCDC24 coiled-coil domain containing 24 1562245_a_at ZNF578 zinc finger protein 578 234224_at PTPRG protein tyrosine phosphatase, receptor type, G 219173_at NA NA 218613_at PSD3 pleckstrin and Sec7 domain containing 3 236167_at TNS3 tensin 3 1562244_at ZNF578 zinc finger protein 578 221909_at RNFT2 ring finger protein, transmembrane 2 1552732_at ABRA actin-binding Rho activating protein 59375_at MYO15B myosin XVB pseudogene 203633_at CPT1A carnitine palmitoyltransferase 1A (liver) 1563120_at NA NA 1560098_at AKR1C2 aldo-keto reductase family 1, member C2 (dihydrodiol dehydrogenase 2; bile acid binding pro 238576_at NA NA 202283_at SERPINF1 serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), m 214248_s_at TRIM2 tripartite motif-containing 2 204766_s_at NUDT1 nudix (nucleoside diphosphate linked moiety X)-type motif 1 242308_at MCOLN3 mucolipin 3 1569154_a_at NA NA 228171_s_at PLEKHG4 pleckstrin homology domain containing, family G (with RhoGef domain) member 4 1552587_at CNBD1 cyclic nucleotide binding domain containing 1 220705_s_at ADAMTS7 ADAM metallopeptidase with thrombospondin type 1 motif, 7 232332_at RP13-347D8.3 KIAA1210 protein 1553618_at TRIM43 tripartite motif-containing 43 209369_at ANXA3 annexin A3 243143_at FAM24A family with sequence similarity 24, member A 234742_at SIRPG signal-regulatory protein gamma