Anti-SIX1 Picoband Antibody Catalog # ABO12575
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

GRIM19 Impedes Obesity by Regulating Inflammatory White Fat
cells Article GRIM19 Impedes Obesity by Regulating Inflammatory White Fat Browning and Promoting Th17/Treg Balance JooYeon Jhun 1,†, Jin Seok Woo 1,† , Seung Hoon Lee 2, Jeong-Hee Jeong 1, KyungAh Jung 3, Wonhee Hur 4, Seon-Yeong Lee 1, Jae Yoon Ryu 1, Young-Mee Moon 1, Yoon Ju Jung 5, Kyo Young Song 5, Kiyuk Chang 6, Seung Kew Yoon 4,7 , Sung-Hwan Park 1,8 and Mi-La Cho 1,8,* 1 The Rheumatism Research Center, Catholic Research Institute of Medical Science, The Catholic University of Korea, Seoul 137-040, Korea; [email protected] (J.J.); [email protected] (J.S.W.); [email protected] (J.-H.J.); [email protected] (S.-Y.L.); [email protected] (J.Y.R.); [email protected] (Y.-M.M.); [email protected] (S.-H.P.) 2 Division of Immunology, Department of Microbiology and Immunobiology, Harvard Medical School, Boston, MA 02115, USA; [email protected] 3 Research Center, Impact Biotech, Seoul 137-040, Korea; [email protected] 4 The Catholic University Liver Research Center & WHO Collaborating Center of Viral Hepatitis, College of Medicine, The Catholic University of Korea, Seoul 137-040, Korea; [email protected] (W.H.); [email protected] (S.K.Y.) 5 Division of Gastrointestinal Surgery, Department of General Surgery, Seoul St. Mary’s Hospital, The Catholic University of Korea, Seoul 137-040, Korea; [email protected] (Y.J.J.); [email protected] (K.Y.S.) 6 Cardiovascular Center and Cardiology Division, Seoul St. Mary’s Hospital, College of Medicine, The Catholic University of Korea, Seoul 137-040, Korea; [email protected] 7 Department of Internal Medicine, Seoul St. -

Characterization of the Human CIDEA Promoter in Fat Cells
International Journal of Obesity (2008) 32, 1380–1387 & 2008 Macmillan Publishers Limited All rights reserved 0307-0565/08 $32.00 www.nature.com/ijo ORIGINAL ARTICLE Characterization of the human CIDEA promoter in fat cells AT Pettersson1, J Laurencikiene1, EA Nordstro¨m1, BM Stenson1, V van Harmelen1, C Murphy2, I Dahlman1 and M Ryde´n1 1Department of Medicine, Huddinge, Lipid Laboratory, Novum, Karolinska Institutet, Stockholm, Sweden and 2Department of Laboratory Medicine, Karolinska Institutet, Stockholm, Sweden. Background: Cell death-inducing DFFA (DNA fragmentation factor-a)-like effector A (CIDEA) is a protein that regulates lipolysis in human adipocytes through cross-talk involving tumor necrosis factor-a (TNF-a). TNF-a downregulates CIDEA mRNA although it is unclear whether this is mediated through transcriptional or post-transcriptional mechanisms. CIDEA has important metabolic effects in human fat cells and genetic variations in the human CIDEA gene have been correlated to the development of obesity. However, little is known about the factors regulating CIDEA expression in human adipocytes. We set out to describe the transcriptional control of human CIDEA. Methods: A 1.1-kb genomic fragment upstream of the transcriptional start site (TSS) of human CIDEA was cloned and deletion fragments were generated. Transcriptional activity of the promoter was analyzed by luciferase reporter assays in in vitro- differentiated human adipocytes. The effect of TNF-a was assessed in human adipocytes and murine 3T3-L1 cells transfected with deletion fragments of the CIDEA promoter. Protein–DNA interactions were analyzed by electrophoretic mobility shift assays (EMSA). Results: Basal transcriptional activity was found in a 97-bp region upstream of the TSS. -
![Anti-CIDE a Antibody [V62P1E3*B10] (ARG10830)](https://docslib.b-cdn.net/cover/1546/anti-cide-a-antibody-v62p1e3-b10-arg10830-181546.webp)
Anti-CIDE a Antibody [V62P1E3*B10] (ARG10830)
Product datasheet [email protected] ARG10830 Package: 100 μg anti-CIDE A antibody [V62P1E3*B10] Store at: -20°C Summary Product Description Mouse Monoclonal antibody recognizes CIDE A Tested Reactivity Hu Tested Application IHC-P, WB Host Mouse Clonality Monoclonal Clone V62P1E3*B10 Isotype IgG1, kappa Target Name CIDE A Antigen Species Human Immunogen Ovalbumin-conjugated synthetic peptide. (QAKGRFTCG) Conjugation Un-conjugated Alternate Names CIDE-A; Cell death-inducing DFFA-like effector A; Cell death activator CIDE-A Application Instructions Application table Application Dilution IHC-P Assay-dependent WB Assay-dependent Application Note Antigen Retrieval: Boil tissue section for 10 - 20 min at 800 - 950W microwave with 10 mM Citrate buffer or Sodium citrate buffer (pH 6.0). * The dilutions indicate recommended starting dilutions and the optimal dilutions or concentrations should be determined by the scientist. Calculated Mw 25 kDa Properties Form Liquid Purification Affinity purification with immunogen. Storage instruction For continuous use, store undiluted antibody at 2-8°C for up to a week. For long-term storage, aliquot and store at -20°C or below. Storage in frost free freezers is not recommended. Avoid repeated freeze/thaw cycles. Suggest spin the vial prior to opening. The antibody solution should be gently mixed before use. Note For laboratory research only, not for drug, diagnostic or other use. Bioinformation www.arigobio.com 1/3 Gene Symbol CIDEA Gene Full Name cell death-inducing DFFA-like effector a Background This gene encodes the homolog of the mouse protein Cidea that has been shown to activate apoptosis. This activation of apoptosis is inhibited by the DNA fragmentation factor DFF45 but not by caspase inhibitors. -

Análise Integrativa De Perfis Transcricionais De Pacientes Com
UNIVERSIDADE DE SÃO PAULO FACULDADE DE MEDICINA DE RIBEIRÃO PRETO PROGRAMA DE PÓS-GRADUAÇÃO EM GENÉTICA ADRIANE FEIJÓ EVANGELISTA Análise integrativa de perfis transcricionais de pacientes com diabetes mellitus tipo 1, tipo 2 e gestacional, comparando-os com manifestações demográficas, clínicas, laboratoriais, fisiopatológicas e terapêuticas Ribeirão Preto – 2012 ADRIANE FEIJÓ EVANGELISTA Análise integrativa de perfis transcricionais de pacientes com diabetes mellitus tipo 1, tipo 2 e gestacional, comparando-os com manifestações demográficas, clínicas, laboratoriais, fisiopatológicas e terapêuticas Tese apresentada à Faculdade de Medicina de Ribeirão Preto da Universidade de São Paulo para obtenção do título de Doutor em Ciências. Área de Concentração: Genética Orientador: Prof. Dr. Eduardo Antonio Donadi Co-orientador: Prof. Dr. Geraldo A. S. Passos Ribeirão Preto – 2012 AUTORIZO A REPRODUÇÃO E DIVULGAÇÃO TOTAL OU PARCIAL DESTE TRABALHO, POR QUALQUER MEIO CONVENCIONAL OU ELETRÔNICO, PARA FINS DE ESTUDO E PESQUISA, DESDE QUE CITADA A FONTE. FICHA CATALOGRÁFICA Evangelista, Adriane Feijó Análise integrativa de perfis transcricionais de pacientes com diabetes mellitus tipo 1, tipo 2 e gestacional, comparando-os com manifestações demográficas, clínicas, laboratoriais, fisiopatológicas e terapêuticas. Ribeirão Preto, 2012 192p. Tese de Doutorado apresentada à Faculdade de Medicina de Ribeirão Preto da Universidade de São Paulo. Área de Concentração: Genética. Orientador: Donadi, Eduardo Antonio Co-orientador: Passos, Geraldo A. 1. Expressão gênica – microarrays 2. Análise bioinformática por module maps 3. Diabetes mellitus tipo 1 4. Diabetes mellitus tipo 2 5. Diabetes mellitus gestacional FOLHA DE APROVAÇÃO ADRIANE FEIJÓ EVANGELISTA Análise integrativa de perfis transcricionais de pacientes com diabetes mellitus tipo 1, tipo 2 e gestacional, comparando-os com manifestações demográficas, clínicas, laboratoriais, fisiopatológicas e terapêuticas. -

The Genetics of Bipolar Disorder
Molecular Psychiatry (2008) 13, 742–771 & 2008 Nature Publishing Group All rights reserved 1359-4184/08 $30.00 www.nature.com/mp FEATURE REVIEW The genetics of bipolar disorder: genome ‘hot regions,’ genes, new potential candidates and future directions A Serretti and L Mandelli Institute of Psychiatry, University of Bologna, Bologna, Italy Bipolar disorder (BP) is a complex disorder caused by a number of liability genes interacting with the environment. In recent years, a large number of linkage and association studies have been conducted producing an extremely large number of findings often not replicated or partially replicated. Further, results from linkage and association studies are not always easily comparable. Unfortunately, at present a comprehensive coverage of available evidence is still lacking. In the present paper, we summarized results obtained from both linkage and association studies in BP. Further, we indicated new potential interesting genes, located in genome ‘hot regions’ for BP and being expressed in the brain. We reviewed published studies on the subject till December 2007. We precisely localized regions where positive linkage has been found, by the NCBI Map viewer (http://www.ncbi.nlm.nih.gov/mapview/); further, we identified genes located in interesting areas and expressed in the brain, by the Entrez gene, Unigene databases (http://www.ncbi.nlm.nih.gov/entrez/) and Human Protein Reference Database (http://www.hprd.org); these genes could be of interest in future investigations. The review of association studies gave interesting results, as a number of genes seem to be definitively involved in BP, such as SLC6A4, TPH2, DRD4, SLC6A3, DAOA, DTNBP1, NRG1, DISC1 and BDNF. -

Comparative Transcriptomics Reveals Similarities and Differences
Seifert et al. BMC Cancer (2015) 15:952 DOI 10.1186/s12885-015-1939-9 RESEARCH ARTICLE Open Access Comparative transcriptomics reveals similarities and differences between astrocytoma grades Michael Seifert1,2,5*, Martin Garbe1, Betty Friedrich1,3, Michel Mittelbronn4 and Barbara Klink5,6,7 Abstract Background: Astrocytomas are the most common primary brain tumors distinguished into four histological grades. Molecular analyses of individual astrocytoma grades have revealed detailed insights into genetic, transcriptomic and epigenetic alterations. This provides an excellent basis to identify similarities and differences between astrocytoma grades. Methods: We utilized public omics data of all four astrocytoma grades focusing on pilocytic astrocytomas (PA I), diffuse astrocytomas (AS II), anaplastic astrocytomas (AS III) and glioblastomas (GBM IV) to identify similarities and differences using well-established bioinformatics and systems biology approaches. We further validated the expression and localization of Ang2 involved in angiogenesis using immunohistochemistry. Results: Our analyses show similarities and differences between astrocytoma grades at the level of individual genes, signaling pathways and regulatory networks. We identified many differentially expressed genes that were either exclusively observed in a specific astrocytoma grade or commonly affected in specific subsets of astrocytoma grades in comparison to normal brain. Further, the number of differentially expressed genes generally increased with the astrocytoma grade with one major exception. The cytokine receptor pathway showed nearly the same number of differentially expressed genes in PA I and GBM IV and was further characterized by a significant overlap of commonly altered genes and an exclusive enrichment of overexpressed cancer genes in GBM IV. Additional analyses revealed a strong exclusive overexpression of CX3CL1 (fractalkine) and its receptor CX3CR1 in PA I possibly contributing to the absence of invasive growth. -

The Brown Adipocyte Protein CIDEA Promotes Lipid Droplet Fusion
1 The brown adipocyte protein CIDEA promotes lipid droplet 2 fusion via a phosphatidic acid-binding amphipathic helix 3 David Barneda1, Joan Planas-Iglesias2, Maria L. Gaspar3, Dariush Mohammadyani4, 4 Sunil Prasannan2, Dirk Dormann5, Gil-Soo Han5, Stephen A. Jesch3, George M. 5 Carman6, Valerian Kagan4, Malcolm G. Parker1, Nicholas T. Ktistakis7, Judith Klein- 6 Seetharaman2, 4, Ann M. Dixon8, Susan A. Henry3, Mark Christian1,2*. 7 1 Institute of Reproductive and Developmental Biology, Imperial College London, London W12 ONN, 8 UK 9 2 Warwick Medical School, University of Warwick, Coventry, CV4 7AL, UK. 10 3 Department of Molecular Biology and Genetics, Cornell University, Ithaca, New York 14853, USA. 11 4 Department of Bioengineering, University of Pittsburgh, Pittsburgh, Pennsylvania 15219, USA. 12 5 Microscopy Facility, MRC Clinical Sciences Centre, Imperial College London, London W12 0NN, UK 13 6 Department of Food Science, Rutgers Center for Lipid Research, Rutgers University, New Brunswick, 14 New Jersey 08901, USA. 15 7 Signalling Programme, Babraham Institute, Cambridge CB22 3AT, UK. 16 8 Department of Chemistry, University of Warwick, Coventry, CV4 7AL, UK. 17 18 19 *Corresponding author. 20 E-mail: [email protected] 21 Phone number: 44 2476 96 8585 1 22 23 Summary 24 Maintenance of energy homeostasis depends on the highly regulated storage and 25 release of triacylglycerol primarily in adipose tissue and excessive storage is a feature of 26 common metabolic disorders. CIDEA is a lipid droplet (LD)-protein enriched in brown 27 adipocytes promoting the enlargement of LDs which are dynamic, ubiquitous organelles 28 specialized for storing neutral lipids. -
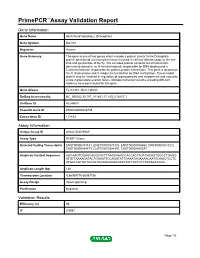
Download Validation Data
PrimePCR™Assay Validation Report Gene Information Gene Name dachshund homolog 2 (Drosophila) Gene Symbol DACH2 Organism Human Gene Summary This gene is one of two genes which encode a protein similar to the Drosophila protein dachshund a transcription factor involved in cell fate determination in the eye limb and genital disc of the fly. The encoded protein contains two characteristic dachshund domains: an N-terminal domain responsible for DNA binding and a C-terminal domain responsible for protein-protein interactions. This gene is located on the X chromosome and is subject to inactivation by DNA methylation. The encoded protein may be involved in regulation of organogenesis and myogenesis and may play a role in premature ovarian failure. Multiple transcript variants encoding different isoforms have been found for this gene. Gene Aliases FLJ31391, MGC138545 RefSeq Accession No. NC_000023.10, NT_011651.17, NG_012817.1 UniGene ID Hs.86603 Ensembl Gene ID ENSG00000126733 Entrez Gene ID 117154 Assay Information Unique Assay ID qHsaCID0009489 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000373131, ENST00000373125, ENST00000508860, ENST00000510272, ENST00000484479, ENST00000344497, ENST00000400297 Amplicon Context Sequence AGCAAGTGGAGCAGGCACTTAAGCAAGCCACCACTAGTGACAGTGGCCTGAGG ATGTTAAAAGATACTGGAATTCCAGATATTGAAATAGAAAACAATGGGACTCCTC ATGATAGTGCTGCTATGCAAGGAGGTAACTATTACTGTTTAGAAATGGC Amplicon Length (bp) 126 Chromosome Location X:86069775-86087136 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) -

Nº Ref Uniprot Proteína Péptidos Identificados Por MS/MS 1 P01024
Document downloaded from http://www.elsevier.es, day 26/09/2021. This copy is for personal use. Any transmission of this document by any media or format is strictly prohibited. Nº Ref Uniprot Proteína Péptidos identificados 1 P01024 CO3_HUMAN Complement C3 OS=Homo sapiens GN=C3 PE=1 SV=2 por 162MS/MS 2 P02751 FINC_HUMAN Fibronectin OS=Homo sapiens GN=FN1 PE=1 SV=4 131 3 P01023 A2MG_HUMAN Alpha-2-macroglobulin OS=Homo sapiens GN=A2M PE=1 SV=3 128 4 P0C0L4 CO4A_HUMAN Complement C4-A OS=Homo sapiens GN=C4A PE=1 SV=1 95 5 P04275 VWF_HUMAN von Willebrand factor OS=Homo sapiens GN=VWF PE=1 SV=4 81 6 P02675 FIBB_HUMAN Fibrinogen beta chain OS=Homo sapiens GN=FGB PE=1 SV=2 78 7 P01031 CO5_HUMAN Complement C5 OS=Homo sapiens GN=C5 PE=1 SV=4 66 8 P02768 ALBU_HUMAN Serum albumin OS=Homo sapiens GN=ALB PE=1 SV=2 66 9 P00450 CERU_HUMAN Ceruloplasmin OS=Homo sapiens GN=CP PE=1 SV=1 64 10 P02671 FIBA_HUMAN Fibrinogen alpha chain OS=Homo sapiens GN=FGA PE=1 SV=2 58 11 P08603 CFAH_HUMAN Complement factor H OS=Homo sapiens GN=CFH PE=1 SV=4 56 12 P02787 TRFE_HUMAN Serotransferrin OS=Homo sapiens GN=TF PE=1 SV=3 54 13 P00747 PLMN_HUMAN Plasminogen OS=Homo sapiens GN=PLG PE=1 SV=2 48 14 P02679 FIBG_HUMAN Fibrinogen gamma chain OS=Homo sapiens GN=FGG PE=1 SV=3 47 15 P01871 IGHM_HUMAN Ig mu chain C region OS=Homo sapiens GN=IGHM PE=1 SV=3 41 16 P04003 C4BPA_HUMAN C4b-binding protein alpha chain OS=Homo sapiens GN=C4BPA PE=1 SV=2 37 17 Q9Y6R7 FCGBP_HUMAN IgGFc-binding protein OS=Homo sapiens GN=FCGBP PE=1 SV=3 30 18 O43866 CD5L_HUMAN CD5 antigen-like OS=Homo -

Proteomic Analysis of Monolayer-Integrated Proteins on Lipid Droplets Identifies Amphipathic Interfacial Α-Helical Membrane Anchors
Proteomic analysis of monolayer-integrated proteins on lipid droplets identifies amphipathic interfacial α-helical membrane anchors Camille I. Patakia, João Rodriguesb, Lichao Zhangc, Junyang Qiand, Bradley Efrond, Trevor Hastied, Joshua E. Eliasc, Michael Levittb, and Ron R. Kopitoe,1 aDepartment of Biochemistry, Stanford University, Stanford, CA 94305; bDepartment of Structural Biology, Stanford University, Stanford, CA 94305; cDepartment of Chemical and Systems Biology, Stanford University, Stanford, CA 94305; dDepartment of Statistics, Stanford University, Stanford, CA 94305; and eDepartment of Biology, Stanford University, Stanford, CA 94305 Edited by Jennifer Lippincott-Schwartz, Howard Hughes Medical Institute, Ashburn, VA, and approved July 23, 2018 (received for review May 9, 2018) Despite not spanning phospholipid bilayers, monotopic integral The first step in understanding the structural diversity of proteins (MIPs) play critical roles in organizing biochemical reac- monotopic HMDs is to identify a sufficiently large set of proteins tions on membrane surfaces. Defining the structural basis by with unequivocal monotopic topology. In this study, we exploited which these proteins are anchored to membranes has been the fact that lipid droplets (LDs) are enveloped by phospholipid hampered by the paucity of unambiguously identified MIPs and a monolayers that separate a highly hydrophobic lipid core from lack of computational tools that accurately distinguish monolayer- the aqueous cytoplasm (8). Because TMDs are flanked on both integrating motifs from bilayer-spanning transmembrane domains ends by soluble, hydrophilic domains, bilayer-spanning proteins (TMDs). We used quantitative proteomics and statistical modeling are strongly disfavored from integrating into the monolayer to identify 87 high-confidence candidate MIPs in lipid droplets, membranes of LDs. -

Loss of DACH2 Is a Candidate Early Event in Breast and Ovarian Carcinogenesis
Loss of DACH2 is a candidate early event in breast and ovarian carcinogenesis By Rania Chehade A thesis submitted in conformity with the requirements for Degree of Master of Science Medical Biophysics Department University of Toronto Copyright by Rania Chehade 2013 Loss of DACH2 is a candidate early event in breast and ovarian carcinogenesis Rania Chehade Master of Science Medical Biophysics Department University of Toronto 2013 Abstract Mechanistic insights into how enduring menstrual cycle hormonal signaling promotes tumorigenesis are emerging. We performed a genome-wide screen in primary epithelial cells to identify hormonally-regulated candidates that initiate pro-tumorigenic phenotypes in normal cells. One candidate, DACH2, has been described as part of a network that regulates organogenesis during development. In vitro, we find that DACH2 expression is regulated by estrogen and progesterone in a dosage dependent manner. Lentiviral-mediated shRNA silencing of DACH2 in hormonally responsive tissues promotes expansion of cells with progenitor characteristics. Within gene expression profiles of fallopian tubes, DACH2 is a member of a minimal gene classifier that distinguishes follicular versus luteal phases. Decreased DACH2 is characteristic of luteal-phase tubal cells and the majority of ovarian serous carcinomas. These studies suggest that DACH2 may orchestrate a physiological program that, when deregulated, locks cells in a progenitor-state, induces uncontrolled proliferation, and predisposes cells for breast and ovarian carcinogenesis. ii Acknowledgements Graduate school has been a bountiful journey of learning opportunities in academic, work and life relationships. I have developed various skills and I have matured both as a researcher and a human being. First and foremost, I would like to acknowledge my supervisor Dr. -

PRDM16 Binds MED1 and Controls Chromatin Architecture to Determine a Brown Fat Transcriptional Program
Downloaded from genesdev.cshlp.org on September 24, 2021 - Published by Cold Spring Harbor Laboratory Press PRDM16 binds MED1 and controls chromatin architecture to determine a brown fat transcriptional program Matthew J. Harms,1,2,4 Hee-Woong Lim,1,3,4 Yugong Ho,3 Suzanne N. Shapira,1,2 Jeff Ishibashi,1,2 Sona Rajakumari,1,2 David J. Steger,1 Mitchell A. Lazar,1 Kyoung-Jae Won,1,3 and Patrick Seale1,2 1Institute for Diabetes, Obesity, and Metabolism, 2Department of Cell and Developmental Biology, 3Genetics Department, Smilow Center for Translational Research, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania 19104, USA PR (PRD1–BF1–RIZ1 homologous) domain-containing 16 (PRDM16) drives a brown fat differentiation program, but the mechanisms by which PRDM16 activates brown fat-selective genes have been unclear. Through chromatin immunoprecipitation (ChIP) followed by deep sequencing (ChIP-seq) analyses in brown adipose tissue (BAT), we reveal that PRDM16 binding is highly enriched at a broad set of brown fat-selective genes. Importantly, we found that PRDM16 physically binds to MED1, a component of the Mediator complex, and recruits it to superenhancers at brown fat-selective genes. PRDM16 deficiency in BAT reduces MED1 binding at PRDM16 target sites and causes a fundamental change in chromatin architecture at key brown fat-selective genes. Together, these data indicate that PRDM16 controls chromatin architecture and superenhancer activity in BAT. [Keywords: PRDM16; PRDM3; MED1; Mediator; brown adipose tissue; UCP1] Supplemental material is available for this article. Received September 17, 2014; revised version accepted December 10, 2014. Obesity is a leading cause of preventable death in the United required for beige fat differentiation and the maintenance of States due to its link to many diseases, including type 2 brown adipose tissue (BAT) fate in adult mice (Cohen diabetes, cardiovascular disease, stroke, and certain cancers et al.