Supporting Document 1
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Restriction Endonucleases
Molecular Biology Problem Solver: A Laboratory Guide. Edited by Alan S. Gerstein Copyright © 2001 by Wiley-Liss, Inc. ISBNs: 0-471-37972-7 (Paper); 0-471-22390-5 (Electronic) 9 Restriction Endonucleases Derek Robinson, Paul R. Walsh, and Joseph A. Bonventre Background Information . 226 Which Restriction Enzymes Are Commercially Available? . 226 Why Are Some Enzymes More Expensive Than Others? . 227 What Can You Do to Reduce the Cost of Working with Restriction Enzymes? . 228 If You Could Select among Several Restriction Enzymes for Your Application, What Criteria Should You Consider to Make the Most Appropriate Choice? . 229 What Are the General Properties of Restriction Endonucleases? . 232 What Insight Is Provided by a Restriction Enzyme’s Quality Control Data? . 233 How Stable Are Restriction Enzymes? . 236 How Stable Are Diluted Restriction Enzymes? . 236 Simple Digests . 236 How Should You Set up a Simple Restriction Digest? . 236 Is It Wise to Modify the Suggested Reaction Conditions? . 237 Complex Restriction Digestions . 239 How Can a Substrate Affect the Restriction Digest? . 239 Should You Alter the Reaction Volume and DNA Concentration? . 241 Double Digests: Simultaneous or Sequential? . 242 225 Genomic Digests . 244 When Preparing Genomic DNA for Southern Blotting, How Can You Determine If Complete Digestion Has Been Obtained? . 244 What Are Your Options If You Must Create Additional Rare or Unique Restriction Sites? . 247 Troubleshooting . 255 What Can Cause a Simple Restriction Digest to Fail? . 255 The Volume of Enzyme in the Vial Appears Very Low. Did Leakage Occur during Shipment? . 259 The Enzyme Shipment Sat on the Shipping Dock for Two Days. -

SUPPLEMENTARY INFORMATION in Silico Signature Prediction
SUPPLEMENTARY INFORMATION In Silico Signature Prediction Modeling in Cytolethal Distending Toxin-Producing Escherichia coli Strains Maryam Javadi, Mana Oloomi*, Saeid Bouzari Department of Molecular Biology, Pasteur Institute of Iran, Tehran 13164, Iran http://www.genominfo.org/src/sm/gni-15-69-s001.pdf Supplementary Table 6. Aalphabetic abbreviation and description of putative conserved domains Alphabetic Abbreviation Description 17 Large terminase protein 2_A_01_02 Multidrug resistance protein 2A0115 Benzoate transport; [Transport and binding proteins, Carbohydrates, organic alcohols] 52 DNA topisomerase II medium subunit; Provisional AAA_13 AAA domain; This family of domains contain a P-loop motif AAA_15 AAA ATPase domain; This family of domains contain a P-loop motif AAA_21 AAA domain AAA_23 AAA domain ABC_RecF ATP-binding cassette domain of RecF; RecF is a recombinational DNA repair ATPase ABC_SMC_barmotin ATP-binding cassette domain of barmotin, a member of the SMC protein family AcCoA-C-Actrans Acetyl-CoA acetyltransferases AHBA_syn 3-Amino-5-hydroxybenzoic acid synthase family (AHBA_syn) AidA Type V secretory pathway, adhesin AidA [Cell envelope biogenesis] Ail_Lom Enterobacterial Ail/Lom protein; This family consists of several bacterial and phage Ail_Lom proteins AIP3 Actin interacting protein 3; Aip3p/Bud6p is a regulator of cell and cytoskeletal polarity Aldose_epim_Ec_YphB Aldose 1-epimerase, similar to Escherichia coli YphB AlpA Predicted transcriptional regulator [Transcription] AntA AntA/AntB antirepressor AraC AraC-type -

Open Matthew R Moreau Ph.D. Dissertation Finalfinal.Pdf
The Pennsylvania State University The Graduate School Department of Veterinary and Biomedical Sciences Pathobiology Program PATHOGENOMICS AND SOURCE DYNAMICS OF SALMONELLA ENTERICA SEROVAR ENTERITIDIS A Dissertation in Pathobiology by Matthew Raymond Moreau 2015 Matthew R. Moreau Submitted in Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy May 2015 The Dissertation of Matthew R. Moreau was reviewed and approved* by the following: Subhashinie Kariyawasam Associate Professor, Veterinary and Biomedical Sciences Dissertation Adviser Co-Chair of Committee Bhushan M. Jayarao Professor, Veterinary and Biomedical Sciences Dissertation Adviser Co-Chair of Committee Mary J. Kennett Professor, Veterinary and Biomedical Sciences Vijay Kumar Assistant Professor, Department of Nutritional Sciences Anthony Schmitt Associate Professor, Veterinary and Biomedical Sciences Head of the Pathobiology Graduate Program *Signatures are on file in the Graduate School iii ABSTRACT Salmonella enterica serovar Enteritidis (SE) is one of the most frequent common causes of morbidity and mortality in humans due to consumption of contaminated eggs and egg products. The association between egg contamination and foodborne outbreaks of SE suggests egg derived SE might be more adept to cause human illness than SE from other sources. Therefore, there is a need to understand the molecular mechanisms underlying the ability of egg- derived SE to colonize the chicken intestinal and reproductive tracts and cause disease in the human host. To this end, the present study was carried out in three objectives. The first objective was to sequence two egg-derived SE isolates belonging to the PFGE type JEGX01.0004 to identify the genes that might be involved in SE colonization and/or pathogenesis. -

Octopine Synthase Mrna Isolated from Sunflower Crown Gall Callus Is
Proc. Nati Acad. Sci. USA Vol. 79, pp. 86-90, January 1982 Biochemistry Octopine synthase mRNA isolated from sunflower crown gall callus is homologous to the Ti plasmid of Agrobacterium tumefaciens (recombinant plasmid/hybridization/mRNA size/in vitro translation/immunoprecipitation) NORIMOTO MURAI AND JOHN D. KEMP Department of Plant Pathology, University ofWisconsin, and Plant Disease Research Unit, ARS, USDA, Madison, Wisconsin 53706 Communicated by Folke Skoog, September 14, 1981 ABSTRACT We have shown that the structural gene for oc- families have been identified. The enzymes responsible for the topine synthase (a crown gall-specific enzyme) is located in a cen- synthesis of the first two have been purified and characterized tral portion ofthe T-DNA that came from the Ti plasmid ofAgro- (17, 18). bacterium tumefaciens and is expressed after it has been trans- The particular opine family present in a crown gall cell cor- ferred to the plant cells. Polyadenylylated RNA was prepared relates with a particular Ti plasmid rather than with the host from polysomes isolated from an octopine-producing crown gall plant (19, 20). Analysis ofdeletion mutants of an octopine-type callus and purified by selective hybridization to one offive recom- Ti plasmid has shown that a gene controlling octopine produc- binant plasmids. Each such plasmid contained a different frag- tion is located on a 14.6-kbp fragment ofthe T-DNA generated ment ofT-DNA ofpTi-15955 (octopine-type Ti plasmid). Purified by Sin 1 (21) (Fig. 1). Detailed examination ofthe physical or- mRNA was translated in vitro in rabbit reticulocyte lysates, and ganization of T-DNA in octopine-type tumor lines has shown the translation products were immunoprecipitated with antibody that octopine production is correlated with the presence ofthe against octopine synthase. -

Roles of Methylation and Sequestration in the Mechanisms of DNA Replication in Some Members of the Enterobacteriaceae Family
Chapter 12 Roles of Methylation and Sequestration in the Mechanisms of DNA Replication in some Members of the Enterobacteriaceae Family Amine Aloui, Alya El May, Saloua Kouass Sahbani and Ahmed Landoulsi Additional information is available at the end of the chapter http://dx.doi.org/10.5772/51724 1. Introduction When growing cells divide, they need to copy their genetic material and distribute it to en‐ sure that each daughter cell receives one copy. This is a challenging task especially when the enormous length of the DNA compared to the cell size is considered. During DNA replica‐ tion, organization of the chromosomes is even more demanding, since replication forks con‐ tinuously produce new DNA. This DNA contains all the information required to build the cells and tissues of a prokaryotic or an eukaryotic organism. The exact replication of this in‐ formation in any species assures its genetic continuity from generation to generation and is critical to the normal development of an individual. The information stored in DNA is ar‐ ranged in hereditary units known as genes that control the identifiable traits of an organism. Discovery of the structure of DNA and subsequent elucidation of how DNA directs synthe‐ sis of RNA, which then directs assembly of proteins -the so-called central dogma - were monumental achievements that marked the early days of molecular biology. However, the simplified representation of the central dogma as DNA → RNA → protein does not reflect the role of proteins in the synthesis of nucleic acids. Moreover, proteins are largely responsi‐ ble for regulating DNA replication and gene expression, the entire process whereby the in‐ formation encoded in DNA is decoded into the proteins that characterize various cell types. -

Deciphering Bacterial Epigenomes Using Modern Sequencing Technologies
REVIEWS Deciphering bacterial epigenomes using modern sequencing technologies John Beaulaurier, Eric E. Schadt and Gang Fang * Abstract | Prokaryotic DNA contains three types of methylation: N6-methyladenine, N4-methylcytosine and 5-methylcytosine. The lack of tools to analyse the frequency and distribution of methylated residues in bacterial genomes has prevented a full understanding of their functions. Now , advances in DNA sequencing technology , including single- molecule, real- time sequencing and nanopore- based sequencing, have provided new opportunities for systematic detection of all three forms of methylated DNA at a genome- wide scale and offer unprecedented opportunities for achieving a more complete understanding of bacterial epigenomes. Indeed, as the number of mapped bacterial methylomes approaches 2,000, increasing evidence supports roles for methylation in regulation of gene expression, virulence and pathogen–host interactions. Phase variation DNA methylation was discovered in bacteria more than MTases has been found to play important regulatory 1 2–12 A means by which reversible a half century ago . It is now known that modification roles in bacteria . RM systems protect cells from variation of protein expression of the four canonical DNA bases by methylation can act invading DNA by methylating endogenous DNA and is achieved in bacteria, often in as an epigenetic regulator — that is, it can impart dis- cleaving non- methylated foreign DNA2,4. RM sys- an ON/OFF manner. The process creates phenotypic tinct and reversible regulatory states to identical genetic tems are divided into three main categories based diversity in a clonally expanded sequences. In eukaryotes, epigenetic regulation can on the subunits involved and the precise site of DNA population and allows the occur at multiple levels: DNA methylation, nucleosome restriction13–16 (Fig. -

The Microbiota-Produced N-Formyl Peptide Fmlf Promotes Obesity-Induced Glucose
Page 1 of 230 Diabetes Title: The microbiota-produced N-formyl peptide fMLF promotes obesity-induced glucose intolerance Joshua Wollam1, Matthew Riopel1, Yong-Jiang Xu1,2, Andrew M. F. Johnson1, Jachelle M. Ofrecio1, Wei Ying1, Dalila El Ouarrat1, Luisa S. Chan3, Andrew W. Han3, Nadir A. Mahmood3, Caitlin N. Ryan3, Yun Sok Lee1, Jeramie D. Watrous1,2, Mahendra D. Chordia4, Dongfeng Pan4, Mohit Jain1,2, Jerrold M. Olefsky1 * Affiliations: 1 Division of Endocrinology & Metabolism, Department of Medicine, University of California, San Diego, La Jolla, California, USA. 2 Department of Pharmacology, University of California, San Diego, La Jolla, California, USA. 3 Second Genome, Inc., South San Francisco, California, USA. 4 Department of Radiology and Medical Imaging, University of Virginia, Charlottesville, VA, USA. * Correspondence to: 858-534-2230, [email protected] Word Count: 4749 Figures: 6 Supplemental Figures: 11 Supplemental Tables: 5 1 Diabetes Publish Ahead of Print, published online April 22, 2019 Diabetes Page 2 of 230 ABSTRACT The composition of the gastrointestinal (GI) microbiota and associated metabolites changes dramatically with diet and the development of obesity. Although many correlations have been described, specific mechanistic links between these changes and glucose homeostasis remain to be defined. Here we show that blood and intestinal levels of the microbiota-produced N-formyl peptide, formyl-methionyl-leucyl-phenylalanine (fMLF), are elevated in high fat diet (HFD)- induced obese mice. Genetic or pharmacological inhibition of the N-formyl peptide receptor Fpr1 leads to increased insulin levels and improved glucose tolerance, dependent upon glucagon- like peptide-1 (GLP-1). Obese Fpr1-knockout (Fpr1-KO) mice also display an altered microbiome, exemplifying the dynamic relationship between host metabolism and microbiota. -

(12) United States Patent (10) Patent No.: US 7,112,717 B2 Valentin Et Al
US007 112717B2 (12) United States Patent (10) Patent No.: US 7,112,717 B2 Valentin et al. (45) Date of Patent: Sep. 26, 2006 (54) HOMOGENTISATE PRENYLTRANSFERASE 2004/0045051 A1 3/2004 Norris et al. GENE (HPT2). FROM ARABIDOPSIS AND USES THEREOF FOREIGN PATENT DOCUMENTS (75) Inventors: Henry E. Valentin, Chesterfield, MO CA 2339519 2, 2000 (US); Tyamagondlu V. Venkatesh, St. CA 2343919 3, 2000 Louis, MO (US); Karunanandaa CA 237,2332 11 2000 Balasulojini, Creve Coeur, MO (US) DE 19835 219 A1 8, 1998 (73) Assignee: Monsanto Technology LLC, St. Louis, EP O 531 639 A2 3, 1993 MO (US) E. 8. G. O. (*) Notice: Subject to any disclaimer, the term of this EP O 723 O17 A2 T 1996 patent is extended or adjusted under 35 EP O 763 542 A2 3, 1997 U.S.C. 154(b) by 404 days. EP 1 033. 405 A2 9, 2000 EP 1 063. 297 A1 12/2000 (21) Appl. No.: 10/391,363 FR 2 778 527 11, 1999 GB 560.529 4f1944 (22) Filed: Mar. 18, 2003 WO WO 91/02059 2, 1991 WO WO 91,09128 6, 1991 (65) Prior Publication Data WO WO 91/13078 9, 1991 WO WO 93/18158 9, 1993 US 2003/0213017 A1 Nov. 13, 2003 WO WO 94,11516 5, 1994 O O WO WO 94/12014 6, 1994 Related U.S. Application Data WO WO94, 18337 8, 1994 (60) Provisional application No. 60/365,202, filed on Mar. WO WO95/08914 4f1995 19, 2002. WO WO95/18220 7, 1995 WO WO95/23863 9, 1995 (51) Int. -
Q 297 Suppl USE
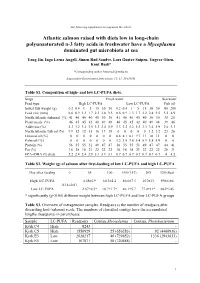
The following supplement accompanies the article Atlantic salmon raised with diets low in long-chain polyunsaturated n-3 fatty acids in freshwater have a Mycoplasma dominated gut microbiota at sea Yang Jin, Inga Leena Angell, Simen Rød Sandve, Lars Gustav Snipen, Yngvar Olsen, Knut Rudi* *Corresponding author: [email protected] Aquaculture Environment Interactions 11: 31–39 (2019) Table S1. Composition of high- and low LC-PUFA diets. Stage Fresh water Sea water Feed type High LC-PUFA Low LC-PUFA Fish oil Initial fish weight (g) 0.2 0.4 1 5 15 30 50 0.2 0.4 1 5 15 30 50 80 200 Feed size (mm) 0.6 0.9 1.3 1.7 2.2 2.8 3.5 0.6 0.9 1.3 1.7 2.2 2.8 3.5 3.5 4.9 North Atlantic fishmeal (%) 41 40 40 40 40 30 30 41 40 40 40 40 30 30 35 25 Plant meals (%) 46 45 45 42 40 49 48 46 45 45 42 40 49 48 39 46 Additives (%) 3.3 3.2 3.2 3.5 3.3 3.4 3.9 3.3 3.2 3.2 3.5 3.3 3.4 3.9 2.6 3.3 North Atlantic fish oil (%) 9.9 12 12 15 16 17 18 0 0 0 0 0 1.2 1.2 23 26 Linseed oil (%) 0 0 0 0 0 0 0 6.8 8.1 8.1 9.7 11 10 11 0 0 Palm oil (%) 0 0 0 0 0 0 0 3.2 3.8 3.8 5.4 5.9 5.8 5.9 0 0 Protein (%) 56 55 55 51 49 47 47 56 55 55 51 49 47 47 44 41 Fat (%) 16 18 18 21 22 22 22 16 18 18 21 22 22 22 28 31 EPA+DHA (% diet) 2.2 2.4 2.4 2.9 3.1 3.1 3.1 0.7 0.7 0.7 0.7 0.7 0.7 0.7 4 4.2 Table S2. -

By USDA APHIS BRS Document Control Officer at 2:18 Pm, Dec 11
CBI Deleted EXECUTIVE SUMMARY The Animal and Plant Health Inspection Service (APHIS) of the United States (U.S.) Department of Agriculture (USDA) has responsibility under the Plant Protection Act (Title IV Pub. L. 106-224, 114 Stat. 438, 7 U.S.C. § 7701-7772) to prevent the introduction and dissemination of plant pests into the U.S. APHIS regulation 7 CFR § 340.6 provides that an applicant may petition APHIS to evaluate submitted data to determine that a particular regulated article does not present a plant pest risk and no longer should be regulated. If APHIS determines that the regulated article does not present a plant pest risk, the petition is granted, thereby allowing unrestricted introduction of the article. Monsanto Company is submitting this request to APHIS for a determination of nonregulated status for the new biotechnology-derived maize product, MON 87429, any progeny derived from crosses between MON 87429 and conventional maize, and any progeny derived from crosses of MON 87429 with biotechnology-derived maize that have previously been granted nonregulated status under 7 CFR Part 340. Product Description Monsanto Company has developed herbicide tolerant MON 87429 maize, which is tolerant to the herbicides dicamba, glufosinate, aryloxyphenoxypropionate (AOPP) acetyl coenzyme A carboxylase (ACCase) inhibitors (so called “FOPs” herbicides such as quizalofop) and 2,4-dichlorophenoxyacetic acid (2,4-D). In addition, it provides tissue-specific glyphosate tolerance to facilitate the production of hybrid maize seeds. MON 87429 contains -

FULLTEXT01.Pdf
ARTICLE https://doi.org/10.1038/s41467-019-11179-9 OPEN Genome-wide systematic identification of methyltransferase recognition and modification patterns Torbjørn Ølshøj Jensen1, Christian Tellgren-Roth2, Stephanie Redl1,3, Jérôme Maury1, Simo Abdessamad Baallal Jacobsen1, Lasse Ebdrup Pedersen 1 & Alex Toftgaard Nielsen 1 1234567890():,; Genome-wide analysis of DNA methylation patterns using single molecule real-time DNA sequencing has boosted the number of publicly available methylomes. However, there is a lack of tools coupling methylation patterns and the corresponding methyltransferase genes. Here we demonstrate a high-throughput method for coupling methyltransferases with their respective motifs, using automated cloning and analysing the methyltransferases in vectors carrying a strain-specific cassette containing all potential target sites. To validate the method, we analyse the genomes of the thermophile Moorella thermoacetica and the mesophile Acetobacterium woodii, two acetogenic bacteria having substantially modified genomes with 12 methylation motifs and a total of 23 methyltransferase genes. Using our method, we characterize the 23 methyltransferases, assign motifs to the respective enzymes and verify activity for 11 of the 12 motifs. 1 The Novo Nordisk Foundation Center for Biosustainability (CfB), Technical University of Denmark (DTU), DK-2800 Lyngby, Denmark. 2 Uppsala Genome Center, National Genomics Infrastructure, SciLifeLab, SE-751 08 Uppsala, Sweden. 3 Massachusetts Institute of Technology, Cambridge, MA 02139, USA. Correspondence -

Methylation of Adenine in the Nuclear DNA of "Tetrahymena" Is Internucleosomal and Independent of Histone H1 Kathleen M
Marquette University e-Publications@Marquette Biological Sciences Faculty Research and Biological Sciences, Department of Publications 1-1-2002 Methylation of Adenine in the Nuclear DNA of "Tetrahymena" is Internucleosomal and Independent of Histone H1 Kathleen M. Karrer Marquette University, [email protected] Teresa Ann Van Nuland Marquette University Published version. Nucleic Acids Research, Vol. 30, No. 6 (2002): 1354-1370. DOI. © 2002 Oxford University Press. Used with permission. 1364-1370 Nucleic Acids Research, 2002, Vol. 30, No. 6 © 2002 Oxford University Press Methylation of adenine in the nuclear DNA of Tetrahymena is internucleosomal and independent of histone H1 Kathleen M. Karrer* and Teresa A. VanNuland Department of Biology, Wehr Life Sciences, Marquette University, PO Box 1881, Milwaukee, WI 53201·1881, USA Received November 19. 2001; Revised and Accepted January 28. 2002 DDBJ/EMBUGenBank accession nos+ ABSTRACT In the somatic macronucleus of Tetrahymena -0.8% of the There are about 50 copies of each chromosome in the adenine residues are methylated. Because the genome is -75% A-T, this amounts to approximately one methylated adenine somatic macronucleus of the ciliated protozoan per 165 bp of DNA. In vivo, methylation occurs at the Tetrahymena. Approximately 0.8% of the adenine resi sequence 5'-NAT-3' (9). A small fraction of the methylated dues in the macronuclear DNA of Tetrahymena are adenines occur in the sequence GA TC, and can be assayed by methylated to N~methyladenine. The degree of methy digestion with restriction enzyme isoschizomers that are differ lation varies between sites from a very low percentage entially sensitive to adenine methylation.