Genetic Analysis of Chromosome Replication Timing
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Alterations of Genetic Variants and Transcriptomic Features of Response to Tamoxifen in the Breast Cancer Cell Line
Alterations of Genetic Variants and Transcriptomic Features of Response to Tamoxifen in the Breast Cancer Cell Line Mahnaz Nezamivand-Chegini Shiraz University Hamed Kharrati-Koopaee Shiraz University https://orcid.org/0000-0003-2345-6919 seyed taghi Heydari ( [email protected] ) Shiraz University of Medical Sciences https://orcid.org/0000-0001-7711-1137 Hasan Giahi Shiraz University Ali Dehshahri Shiraz University of Medical Sciences Mehdi Dianatpour Shiraz University of Medical Sciences Kamran Bagheri Lankarani Shiraz University of Medical Sciences Research Keywords: Tamoxifen, breast cancer, genetic variants, RNA-seq. Posted Date: August 17th, 2021 DOI: https://doi.org/10.21203/rs.3.rs-783422/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/33 Abstract Background Breast cancer is one of the most important causes of mortality in the world, and Tamoxifen therapy is known as a medication strategy for estrogen receptor-positive breast cancer. In current study, two hypotheses of Tamoxifen consumption in breast cancer cell line (MCF7) were investigated. First, the effect of Tamoxifen on genes expression prole at transcriptome level was evaluated between the control and treated samples. Second, due to the fact that Tamoxifen is known as a mutagenic factor, there may be an association between the alterations of genetic variants and Tamoxifen treatment, which can impact on the drug response. Methods In current study, the whole-transcriptome (RNA-seq) dataset of four investigations (19 samples) were derived from European Bioinformatics Institute (EBI). At transcriptome level, the effect of Tamoxifen was investigated on gene expression prole between control and treatment samples. -

Aneuploidy: Using Genetic Instability to Preserve a Haploid Genome?
Health Science Campus FINAL APPROVAL OF DISSERTATION Doctor of Philosophy in Biomedical Science (Cancer Biology) Aneuploidy: Using genetic instability to preserve a haploid genome? Submitted by: Ramona Ramdath In partial fulfillment of the requirements for the degree of Doctor of Philosophy in Biomedical Science Examination Committee Signature/Date Major Advisor: David Allison, M.D., Ph.D. Academic James Trempe, Ph.D. Advisory Committee: David Giovanucci, Ph.D. Randall Ruch, Ph.D. Ronald Mellgren, Ph.D. Senior Associate Dean College of Graduate Studies Michael S. Bisesi, Ph.D. Date of Defense: April 10, 2009 Aneuploidy: Using genetic instability to preserve a haploid genome? Ramona Ramdath University of Toledo, Health Science Campus 2009 Dedication I dedicate this dissertation to my grandfather who died of lung cancer two years ago, but who always instilled in us the value and importance of education. And to my mom and sister, both of whom have been pillars of support and stimulating conversations. To my sister, Rehanna, especially- I hope this inspires you to achieve all that you want to in life, academically and otherwise. ii Acknowledgements As we go through these academic journeys, there are so many along the way that make an impact not only on our work, but on our lives as well, and I would like to say a heartfelt thank you to all of those people: My Committee members- Dr. James Trempe, Dr. David Giovanucchi, Dr. Ronald Mellgren and Dr. Randall Ruch for their guidance, suggestions, support and confidence in me. My major advisor- Dr. David Allison, for his constructive criticism and positive reinforcement. -

Download Thesis
This electronic thesis or dissertation has been downloaded from the King’s Research Portal at https://kclpure.kcl.ac.uk/portal/ The Genetics and Spread of Amyotrophic Lateral Sclerosis Jones, Ashley Richard Awarding institution: King's College London The copyright of this thesis rests with the author and no quotation from it or information derived from it may be published without proper acknowledgement. END USER LICENCE AGREEMENT Unless another licence is stated on the immediately following page this work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International licence. https://creativecommons.org/licenses/by-nc-nd/4.0/ You are free to copy, distribute and transmit the work Under the following conditions: Attribution: You must attribute the work in the manner specified by the author (but not in any way that suggests that they endorse you or your use of the work). Non Commercial: You may not use this work for commercial purposes. No Derivative Works - You may not alter, transform, or build upon this work. Any of these conditions can be waived if you receive permission from the author. Your fair dealings and other rights are in no way affected by the above. Take down policy If you believe that this document breaches copyright please contact [email protected] providing details, and we will remove access to the work immediately and investigate your claim. Download date: 07. Oct. 2021 THE GENETICS AND SPREAD OF AMYOTROPHIC LATERAL SCLEROSIS Ashley Richard Jones PhD in Clinical Neuroscience - 1 - Abstract Our knowledge of the genetic contribution to Amyotrophic Lateral Sclerosis (ALS) is rapidly growing, and there is increasing research into how ALS spreads through the motor system and beyond. -

Regular Article
From www.bloodjournal.org by guest on April 6, 2015. For personal use only. Regular Article IMMUNOBIOLOGY Hemophagocytic lymphohistiocytosis caused by dominant-negative mutations in STXBP2 that inhibit SNARE-mediated membrane fusion Waldo A. Spessott,1 Maria L. Sanmillan,1 Margaret E. McCormick,1 Nishant Patel,2 Joyce Villanueva,3 Kejian Zhang,4 Kim E. Nichols,5 and Claudio G. Giraudo1 1Department of Pathology and Laboratory Medicine, and 2Division of Oncology, Department of Pediatrics, The Children’s Hospital of Philadelphia, University of Pennsylvania, Philadelphia, PA; 3Division of Bone Marrow Transplant and Immune Deficiency, and 4Division of Human Genetics, Cincinnati Children’s Hospital Medical Center, Department of Pediatrics, University of Cincinnati College of Medicine, Cincinnati, OH; and 5Division of Cancer Predisposition, Department of Oncology, St. Jude Children’s Research Hospital, Memphis, TN Key Points Familial hemophagocytic lymphohistiocytosis (F-HLH) and Griscelli syndrome type 2 (GS) are life-threatening immunodeficiencies characterized by impaired cytotoxic T lymphocyte • Monoallelic STXBP2 mutations (CTL) and natural killer (NK) cell lytic activity. In the majority of cases, these disorders are affecting codon 65 impair caused by biallelic inactivating germline mutations in genes such as RAB27A (GS) and PRF1, lymphocyte cytotoxicity and UNC13D, STX11,andSTXBP2 (F-HLH). Although monoallelic (ie, heterozygous) mutations contribute to hemophagocytic have been identified in certain patients, the clinical significance and molecular mechanisms lymphohistiocytosis. by which these mutations influence CTL and NK cell function remain poorly understood. • Munc18-2R65Q/W mutant Here, we characterize 2 novel monoallelic hemophagocytic lymphohistiocytosis (HLH)- associated mutations affecting codon 65 of STXPB2, the gene encoding Munc18-2, a member proteins function in a dominant- of the SEC/MUNC18 family. -

University of Florida Thesis Or Dissertation Formatting
RNA-SEQ REVEALS NOVEL GENES AND PATHWAYS INVOLVED IN BOVINE MAMMARY INVOLUTION DURING THE DRY PERIOD AND UNDER ENVIRONMENTAL HEAT STRESS By BETHANY M. DADO SENN A THESIS PRESENTED TO THE GRADUATE SCHOOL OF THE UNIVERSITY OF FLORIDA IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE OF MASTER OF SCIENCE UNIVERSITY OF FLORIDA 2018 © 2018 Bethany M. Dado Senn To my family, the true dairy enthusiasts ACKNOWLEDGMENTS To my advisor, mentor, and friend Dr. Jimena Laporta, I am humbled and grateful to have served as your graduate student as you provided invaluable advice and kindness throughout my projects. Your open-door policy has facilitated my growth both personally and professionally. Thank you for the opportunity to research lactation physiology, volunteer and teach, and pursue a degree at the University of Florida. I extend my appreciation to my committee members Dr. Geoffrey Dahl and Dr. Pete Hansen for utilizing their many years of experience to provide useful critiques and additional insight into my analysis and interpretations. Thank you to Dr. Hansen for the use of Ingenuity Pathway Analysis® and to Dr. Dahl for his heat stress expertise. I thank the faculty and staff in the Department of Animal Sciences at the University of Florida, especially Dr. Francisco Peñagaricano for his vital RNA- sequencing and statistical contribution to my thesis project. Further thanks to Dr. Corwin Nelson, Dr. Stephanie Wohlgemuth, and Dr. John Bromfield for use of lab space and research support. Special appreciation goes to Joyce Hayen, Pam Krueger, and Renee Parks-James and the UF Dairy Unit staff. I also express appreciation to the Animal Molecular and Cellular Biology program, the Brélan E. -

Syntaxin Binding Mechanism and Disease-Causing Mutations In
Syntaxin binding mechanism and disease-causing PNAS PLUS mutations in Munc18-2 Yvonne Hackmanna, Stephen C. Grahama,1,2, Stephan Ehlb, Stefan Höningc, Kai Lehmbergd, Maurizio Aricòe, David J. Owena, and Gillian M. Griffithsa,2 aCambridge Institute for Medical Research, University of Cambridge Biomedical Campus, University of Cambridge, Cambridge CB2 0XY, United Kingdom; bCentre of Chronic Immunodeficiency, 79106 Freiburg, Germany; cInstitute for Biochemistry I and Center for Molecular Medicine Cologne, University of Cologne, 50931 Cologne, Germany; dDepartment of Paediatric Haematology and Oncology, University Medical Center Hamburg Eppendorf, 20246 Hamburg, Germany; and ePediatric Hematology Oncology Network, Istituto Toscana Tumori, 50139 Florence, Italy Edited* by Peter Cresswell, Yale University School of Medicine, New Haven, CT, and approved October 11, 2013 (received for review July 18, 2013) Mutations in either syntaxin 11 (Stx11) or Munc18-2 abolish Munc18-2 belongs to the Sec1/Munc18-like (SM) protein cytotoxic T lymphocytes (CTL) and natural killer cell (NK) cytotox- family, whose members are all ∼600 residues long and are in- icity, and give rise to familial hemophagocytic lymphohistiocytosis volved in regulation of SNARE-mediated membrane fusion (FHL4 or FHL5, respectively). Although Munc18-2 is known to events (9, 10). The two closest homologs of Munc18-2 are interact with Stx11, little is known about the molecular mecha- Munc18-1, which is crucial for neurotransmitter secretion in nisms governing the specificity of this interaction or how in vitro neurons (11, 12), and Munc18-3, which is more widely expressed IL-2 activation leads to compensation of CTL and NK cytotoxicity. and is involved in Glut4 translocation (13). -

De Novo Transcriptome Sequencing and Gene Expression Profiling With/Without B-Chromosome Plants of Lilium Amabile Doori Park, Jong-Hwa Kim, Nam-Soo Kim
www.genominfo.org eISSN 2234-0742 Volume 17 number 3, September 30, 2019 Aims and scope Genomics & Informatics is the official journal of the Korea Genome Organization (http://kogo.or.kr). Its abbreviated title is Genomics Inform. It was launched in 2003 by the Korea Genome Organization. It aims at making a substantial contribution to the understanding of any areas of genomics or informatics. Its scope includes novel data on the topics of gene discovery, comparative genome analyses, molecular and human evolution, informatics, genome structure and function, technological innovations and applications, statistical and mathematical methods, cutting-edge genetic and physical mapping, next generation sequencing and de novo assembly, and other topics that present data where sequence information is used to address biological concerns. Especially, Clinical genomics section is for a short report of all kinds of genome analysis data from clinical field such as cancer, diverse complex diseases and genetic diseases. It encourages submission of the cancer panel analysis data for a single cancer patient or a group of patients. It also encourages deposition of the genome data into designated database. Genome archives section is for a short manuscript announcing the genetic information of recently sequenced prokaryotic and eukaryotic genomes. These genome archives data can make the rationale for sequencing a specific organism. It is published and distributed quarterly at the last dates of March, June, September, and December. All submitted manuscripts will be reviewed and selected for publication after single blind review process. All manuscripts must be submitted online through the e-submission system available from: http://submit.genominfo.org. -
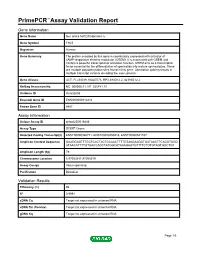
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name four and a half LIM domains 5 Gene Symbol FHL5 Organism Human Gene Summary The protein encoded by this gene is coordinately expressed with activator of cAMP-responsive element modulator (CREM). It is associated with CREM and confers a powerful transcriptional activation function. CREM acts as a transcription factor essential for the differentiation of spermatids into mature spermatozoa. There are multiple polyadenylation sites found in this gene. Alternative splicing results in multiple transcript variants encoding the same protein. Gene Aliases ACT, FLJ33049, KIAA0776, RP3-393D12.2, dJ393D12.2 RefSeq Accession No. NC_000006.11, NT_025741.15 UniGene ID Hs.632608 Ensembl Gene ID ENSG00000112214 Entrez Gene ID 9457 Assay Information Unique Assay ID qHsaCID0016446 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000326771, ENST00000450218, ENST00000541107 Amplicon Context Sequence GAAGGAGTTTGCTCACTACTGCAACTTTTGTAAGAAGGTGATAACTTCAGGTGGG ATAACATTTTGTGACCAGCTATGGCATAAAGAGTGTTTTCTGTGTAGTGGCTGT Amplicon Length (bp) 79 Chromosome Location 6:97053911-97058519 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 96 R2 0.9991 cDNA Cq Target not expressed in universal RNA cDNA Tm (Celsius) Target not expressed in universal RNA gDNA Cq Target not expressed in universal RNA Page 1/5 PrimePCR™Assay Validation Report Specificity (%) No cross reactivity detected Information to assist with data interpretation is provided at the end of this report. Page 2/5 -

Identification of a Deletion in Stxbp2
ndrom Sy es tic & e G n e e n G e f T o Coniglio et al., J Genet Syndr Gene Ther 2015, 6:3 Journal of Genetic Syndromes h l e a r n a DOI: 10.4172/2157-7412.1000276 r p u y o J & Gene Therapy ISSN: 2157-7412 Case Report Open Access “Identification of a Deletion in Stxbp2 Causative of Familial Hemophagocytic Lymphohistiocytosis Type 5” Coniglio ML1,3*, Cetica V1, Ciambotti B1,3, Grieve S2, Pantaleo M3, Rizzari C4, Sieni E1, Favre C1, Giglio S3, Griffiths GM2 and Aricò M5 1Department of Paediatric Oncohematology, Anna Meyer Children’s University Hospital, Florence, Italy 2Cambridge Institute for Medical Research, University of Cambridge Biomedical Campus, Addenbrooke’s Hospital, Cambridge, United Kingdom 3Genetics and Molecular Medicine Unit, Anna Meyer Children’s University Hospital, Florence, Italy 4Department of Pediatrics, Ospedale S. Gerardo, University of Milano-Bicocca, Fondazione MBBM, Monza, Italy 5Azienda Sanitaria Provinciale, Ragusa, Italy Abstract Hemophagocytic lymphohistiocytosis (HLH) is a rare, life-threatening immune deficiency, characterized by a hyper-inflammatory syndrome. The familial form of HLH (FHL) is caused by mutations in genes associated with lymphocyte granule-mediated cytotoxicity. Mutations in Stxbp2 (Sintaxin binding protein 2) gene result in defect of Munc18-2 protein, the causative defect of the subtype defined as FHL5. Functional tests as intracytoplasmic expression of perforin and surface expression of CD107a, help to direct genetic analysis. Different mutations have been described in the FHL-related genes known so far (PRF1, UNC13-D, STX11, Stxbp2): missense, nonsense, splicing, regulatory, small deletions/insertions. Recently a pathogenic inversion of 253 KB upstream of the 3’ UNC13D gene has been reported. -

Genome-Wide Characterization of Genetic Variants and Putative
Boschiero et al. BMC Genomics (2018) 19:83 DOI 10.1186/s12864-018-4444-0 RESEARCH ARTICLE Open Access Genome-wide characterization of genetic variants and putative regions under selection in meat and egg-type chicken lines Clarissa Boschiero1,4* , Gabriel Costa Monteiro Moreira1,AlmasAraGheyas2, Thaís Fernanda Godoy1, Gustavo Gasparin1, Pilar Drummond Sampaio Corrêa Mariani1,MarcelaPaduan1, Aline Silva Mello Cesar1, Mônica Corrêa Ledur3 and Luiz Lehmann Coutinho1 Abstract Background: Meat and egg-type chickens have been selected for several generations for different traits. Artificial and natural selection for different phenotypes can change frequency of genetic variants, leaving particular genomic footprints throghtout the genome. Thus, the aims of this study were to sequence 28 chickens from two Brazilian lines (meat and white egg-type) and use this information to characterize genome-wide genetic variations, identify putative regions under selection using Fst method, and find putative pathways under selection. Results: A total of 13.93 million SNPs and 1.36 million INDELs were identified, with more variants detected from the broiler (meat-type) line. Although most were located in non-coding regions, we identified 7255 intolerant non-synonymous SNPs, 512 stopgain/loss SNPs, 1381 frameshift and 1094 non-frameshift INDELs that may alter protein functions. Genes harboring intolerant non-synonymous SNPs affected metabolic pathways related mainly to reproduction and endocrine systems in the white-egg layer line, and lipid metabolism and metabolic diseases in the broiler line. Fst analysis in sliding windows, using SNPs and INDELs separately, identified over 300 putative regions of selection overlapping with more than 250 genes. For the first time in chicken, INDEL variants were considered for selection signature analysis, showing high level of correlation in results between SNP and INDEL data. -

Expression Profiling Associates Blood and Brain Glucocorticoid Receptor
Expression profiling associates blood and brain SEE COMMENTARY glucocorticoid receptor signaling with trauma-related individual differences in both sexes Nikolaos P. Daskalakisa,b,1, Hagit Cohenc, Guiqing Caia,d, Joseph D. Buxbauma,d,e, and Rachel Yehudaa,b,e Departments of aPsychiatry, dGenetics and Genomic Sciences, and eNeuroscience, Icahn School of Medicine at Mount Sinai, New York, NY 10029; bMental Health Patient Care Center, James J. Peters Veterans Affairs Medical Center, Bronx, NY 10468; and cAnxiety and Stress Research Unit, Ministry of Health Mental Health Center, Faculty of Health Sciences, Ben-Gurion University of the Negev, Beer Sheva 84170, Israel Edited by Bruce S. McEwen, The Rockefeller University, New York, NY, and approved July 14, 2014 (received for review February 7, 2014) Delineating the molecular basis of individual differences in the stress response to stress (4, 5). The emergence of system- and genome- response is critical to understanding the pathophysiology and treat- wide approaches permits the opportunity for unbiased identifi- ment of posttraumatic stress disorder (PTSD). In this study, 7 d after cation of novel pathways. Because PTSD is more prevalent in predator-scent-stress (PSS) exposure, male and female rats were women than men (1), and sex is a potential source of response classified into vulnerable (i.e., “PTSD-like”) and resilient (i.e., minimally variation to trauma in both animals (6) and humans (7), it is also affected) phenotypes on the basis of their performance on a variety of critical to include both sexes in such studies. behavioral measures. Genome-wide expression profiling in blood and In the present study, PSS-exposed male and female rats were two limbic brain regions (amygdala and hippocampus), followed by behaviorally tested in EPM and ASR tests a week after PSS and divided in EBR and MBR groups [at this point, the behavioral quantitative PCR validation, was performed in these two groups of response of the rats is stable in terms of prevalence of EBRs vs. -

Molecular Targeting and Enhancing Anticancer Efficacy of Oncolytic HSV-1 to Midkine Expressing Tumors
University of Cincinnati Date: 12/20/2010 I, Arturo R Maldonado , hereby submit this original work as part of the requirements for the degree of Doctor of Philosophy in Developmental Biology. It is entitled: Molecular Targeting and Enhancing Anticancer Efficacy of Oncolytic HSV-1 to Midkine Expressing Tumors Student's name: Arturo R Maldonado This work and its defense approved by: Committee chair: Jeffrey Whitsett Committee member: Timothy Crombleholme, MD Committee member: Dan Wiginton, PhD Committee member: Rhonda Cardin, PhD Committee member: Tim Cripe 1297 Last Printed:1/11/2011 Document Of Defense Form Molecular Targeting and Enhancing Anticancer Efficacy of Oncolytic HSV-1 to Midkine Expressing Tumors A dissertation submitted to the Graduate School of the University of Cincinnati College of Medicine in partial fulfillment of the requirements for the degree of DOCTORATE OF PHILOSOPHY (PH.D.) in the Division of Molecular & Developmental Biology 2010 By Arturo Rafael Maldonado B.A., University of Miami, Coral Gables, Florida June 1993 M.D., New Jersey Medical School, Newark, New Jersey June 1999 Committee Chair: Jeffrey A. Whitsett, M.D. Advisor: Timothy M. Crombleholme, M.D. Timothy P. Cripe, M.D. Ph.D. Dan Wiginton, Ph.D. Rhonda D. Cardin, Ph.D. ABSTRACT Since 1999, cancer has surpassed heart disease as the number one cause of death in the US for people under the age of 85. Malignant Peripheral Nerve Sheath Tumor (MPNST), a common malignancy in patients with Neurofibromatosis, and colorectal cancer are midkine- producing tumors with high mortality rates. In vitro and preclinical xenograft models of MPNST were utilized in this dissertation to study the role of midkine (MDK), a tumor-specific gene over- expressed in these tumors and to test the efficacy of a MDK-transcriptionally targeted oncolytic HSV-1 (oHSV).