Structure-Based Subfamily Classification of Homeodomains
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Detecting Remote, Functional Conserved Domains in Entire Genomes by Combining Relaxed Sequence-Database Searches with Fold Recognition
HMMerThread: Detecting Remote, Functional Conserved Domains in Entire Genomes by Combining Relaxed Sequence-Database Searches with Fold Recognition Charles Richard Bradshaw1¤a, Vineeth Surendranath1, Robert Henschel2,3, Matthias Stefan Mueller2, Bianca Hermine Habermann1,4*¤b 1 Bioinformatics Laboratory, Max Planck Institute of Molecular Cell Biology and Genetics, Dresden, Saxony, Germany, 2 Center for Information Services and High Performance Computing (ZIH), Technical University, Dresden, Saxony, Germany, 3 High Performance Applications, Pervasive Technology Institute, Indiana University, Bloomington, Indiana, United States of America, 4 Bioinformatics Laboratory, Scionics c/o Max Planck Institute of Molecular Cell Biology and Genetics, Dresden, Saxony, Germany Abstract Conserved domains in proteins are one of the major sources of functional information for experimental design and genome-level annotation. Though search tools for conserved domain databases such as Hidden Markov Models (HMMs) are sensitive in detecting conserved domains in proteins when they share sufficient sequence similarity, they tend to miss more divergent family members, as they lack a reliable statistical framework for the detection of low sequence similarity. We have developed a greatly improved HMMerThread algorithm that can detect remotely conserved domains in highly divergent sequences. HMMerThread combines relaxed conserved domain searches with fold recognition to eliminate false positive, sequence-based identifications. With an accuracy of 90%, our software is able to automatically predict highly divergent members of conserved domain families with an associated 3-dimensional structure. We give additional confidence to our predictions by validation across species. We have run HMMerThread searches on eight proteomes including human and present a rich resource of remotely conserved domains, which adds significantly to the functional annotation of entire proteomes. -
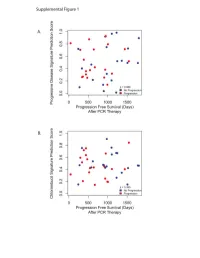
Supplementary Data
Progressive Disease Signature Upregulated probes with progressive disease U133Plus2 ID Gene Symbol Gene Name 239673_at NR3C2 nuclear receptor subfamily 3, group C, member 2 228994_at CCDC24 coiled-coil domain containing 24 1562245_a_at ZNF578 zinc finger protein 578 234224_at PTPRG protein tyrosine phosphatase, receptor type, G 219173_at NA NA 218613_at PSD3 pleckstrin and Sec7 domain containing 3 236167_at TNS3 tensin 3 1562244_at ZNF578 zinc finger protein 578 221909_at RNFT2 ring finger protein, transmembrane 2 1552732_at ABRA actin-binding Rho activating protein 59375_at MYO15B myosin XVB pseudogene 203633_at CPT1A carnitine palmitoyltransferase 1A (liver) 1563120_at NA NA 1560098_at AKR1C2 aldo-keto reductase family 1, member C2 (dihydrodiol dehydrogenase 2; bile acid binding pro 238576_at NA NA 202283_at SERPINF1 serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), m 214248_s_at TRIM2 tripartite motif-containing 2 204766_s_at NUDT1 nudix (nucleoside diphosphate linked moiety X)-type motif 1 242308_at MCOLN3 mucolipin 3 1569154_a_at NA NA 228171_s_at PLEKHG4 pleckstrin homology domain containing, family G (with RhoGef domain) member 4 1552587_at CNBD1 cyclic nucleotide binding domain containing 1 220705_s_at ADAMTS7 ADAM metallopeptidase with thrombospondin type 1 motif, 7 232332_at RP13-347D8.3 KIAA1210 protein 1553618_at TRIM43 tripartite motif-containing 43 209369_at ANXA3 annexin A3 243143_at FAM24A family with sequence similarity 24, member A 234742_at SIRPG signal-regulatory protein gamma -

Supplementary Table 2
Supplementary Table 2. Differentially Expressed Genes following Sham treatment relative to Untreated Controls Fold Change Accession Name Symbol 3 h 12 h NM_013121 CD28 antigen Cd28 12.82 BG665360 FMS-like tyrosine kinase 1 Flt1 9.63 NM_012701 Adrenergic receptor, beta 1 Adrb1 8.24 0.46 U20796 Nuclear receptor subfamily 1, group D, member 2 Nr1d2 7.22 NM_017116 Calpain 2 Capn2 6.41 BE097282 Guanine nucleotide binding protein, alpha 12 Gna12 6.21 NM_053328 Basic helix-loop-helix domain containing, class B2 Bhlhb2 5.79 NM_053831 Guanylate cyclase 2f Gucy2f 5.71 AW251703 Tumor necrosis factor receptor superfamily, member 12a Tnfrsf12a 5.57 NM_021691 Twist homolog 2 (Drosophila) Twist2 5.42 NM_133550 Fc receptor, IgE, low affinity II, alpha polypeptide Fcer2a 4.93 NM_031120 Signal sequence receptor, gamma Ssr3 4.84 NM_053544 Secreted frizzled-related protein 4 Sfrp4 4.73 NM_053910 Pleckstrin homology, Sec7 and coiled/coil domains 1 Pscd1 4.69 BE113233 Suppressor of cytokine signaling 2 Socs2 4.68 NM_053949 Potassium voltage-gated channel, subfamily H (eag- Kcnh2 4.60 related), member 2 NM_017305 Glutamate cysteine ligase, modifier subunit Gclm 4.59 NM_017309 Protein phospatase 3, regulatory subunit B, alpha Ppp3r1 4.54 isoform,type 1 NM_012765 5-hydroxytryptamine (serotonin) receptor 2C Htr2c 4.46 NM_017218 V-erb-b2 erythroblastic leukemia viral oncogene homolog Erbb3 4.42 3 (avian) AW918369 Zinc finger protein 191 Zfp191 4.38 NM_031034 Guanine nucleotide binding protein, alpha 12 Gna12 4.38 NM_017020 Interleukin 6 receptor Il6r 4.37 AJ002942 -

LN-EPC Vs CEPC List
Supplementary Information Table 5. List of genes upregulated on LN-EPC (LCB represents the variation of gene expression comparing LN-EPC with CEPC) Gene dystrophin (muscular dystrophy, Duchenne and Becker types) regulator of G-protein signalling 13 chemokine (C-C motif) ligand 8 vascular cell adhesion molecule 1 matrix metalloproteinase 9 (gelatinase B, 92kDa gelatinase, 92kDa type IV collagenase) chemokine (C-C motif) ligand 2 solute carrier family 2 (facilitated glucose/fructose transporter), member 5 eukaryotic translation initiation factor 1A, Y-linked regulator of G-protein signalling 1 ubiquitin D chemokine (C-X-C motif) ligand 3 transcription factor 4 chemokine (C-X-C motif) ligand 13 (B-cell chemoattractant) solute carrier family 7, (cationic amino acid transporter, y+ system) member 11 transcription factor 4 apolipoprotein D RAS guanyl releasing protein 3 (calcium and DAG-regulated) matrix metalloproteinase 1 (interstitial collagenase) DEAD (Asp-Glu-Ala-Asp) box polypeptide 3, Y-linked /// DEAD (Asp-Glu-Ala-Asp) box polypeptide 3, Y-linked transcription factor 4 regulator of G-protein signalling 1 B-cell linker interleukin 8 POU domain, class 2, associating factor 1 CD24 antigen (small cell lung carcinoma cluster 4 antigen) Consensus includes gb:AK000168.1 /DEF=Homo sapiens cDNA FLJ20161 fis, clone COL09252, highly similar to L33930 Homo sapiens CD24 signal transducer mRNA. /FEA=mRNA /DB_XREF=gi:7020079 /UG=Hs.332045 Homo sapiens cDNA FLJ20161 fis, clone COL09252, highly similar to L33930 Homo sapiens CD24 signal transducer mRNA -

Supplementary Figures
Supplementary Figures Supplementary Figure 1 | Sampling locality, genome size estimation, and GC content. (a) Sampling locality in Amami Island (i.e., Amami Oshima, Japan) and its relative location to Okinawa are shown with coordinates (adapted from Google Maps). (b) Sperm cells collected from gravid male gonads were stained with DAPI and subjected to fluorescence-activated cell sorting (FACS) flow cytometry analysis. Sperm with known genome size from zebrafish (Danio rerio) were used as an internal standard to estimate the Lingula genome size. (c) The analysis of stepwise assembly shows that the saturation point is achieved when input sequences reach 10 Gbp from 454 and Illumina reads. (d) K-mer analysis (17-mer) using Illumina reads shows two peaks, in which the homozygous peak coverage is twice the heterozygous peak. The estimated heterozygosity rate calculating the ratio of the peaks, is 1.6%. (e) Distribution of GC content calculated from 3,830 scaffolds. (f) Comparison of GC content in selected lophotrochozoans. Error bars, standard deviation. Supplementary Figure 2 | Schematic flow of sequencing and assembly of the Lingula genome. (a) Genomic DNA from a male gonad was extracted for genome sequencing using Roche 454, Illumina, and PacBio platforms. A total of 96-Gb of data was obtained with approximately 226- fold coverage of the 425-Mb Lingula genome. (b) Ten embryonic stages from egg to larva and seven adult tissues were collected for RNA-seq and reads were assembled de novo using Trinity. (c) Transcript information from RNA-seq was used to generate hints by spliced alignment with PASA and BLAT. Gene models were predicted with trained AUGUSTUS. -

Protein Classification Based on Text Document Classification Techniques
PROTEINS: Structure, Function, and Bioinformatics 58:955–970 (2005) Protein Classification Based on Text Document Classification Techniques Betty Yee Man Cheng,1 Jaime G. Carbonell,1 and Judith Klein-Seetharaman1,2* 1Language Technologies Institute, Carnegie Mellon University, 5000 Forbes Avenue, Pittsburgh, Pennsylvania 2Department of Pharmacology, University of Pittsburgh School of Medicine, Biomedical Science Tower E1058, Pittsburgh, Pennsylvania ABSTRACT The need for accurate, automated The first category of methods (Table I-A) searches a protein classification methods continues to increase database of known sequences for the one most similar to as advances in biotechnology uncover new proteins. the query sequence and assigns its classification to the G-protein coupled receptors (GPCRs) are a particu- query sequence. The similarity search is accomplished by larly difficult superfamily of proteins to classify due performing a pairwise sequence alignment between the to extreme diversity among its members. Previous query sequence and every sequence in the database using comparisons of BLAST, k-nearest neighbor (k-NN), an amino acid similarity matrix. Smith–Waterman1 and hidden markov model (HMM) and support vector Needleman–Wunsch2 are dynamic programming algo- machine (SVM) using alignment-based features have rithms guaranteed to find the optimal local and global suggested that classifiers at the complexity of SVM alignment respectively, but they are extremely slow and are needed to attain high accuracy. Here, analogous thus impossible to use in a database-wide search. A to document classification, we applied Decision Tree number of heuristic algorithms have been developed, of and Naı¨ve Bayes classifiers with chi-square feature which BLAST3 is the most prevalent. -

Supplemental Material 1
Supplemental gure 1 Lin-Sca-1- Marker Thymus Bone Skin 41% ±1 64% ±8 92% ±4 CD29 % of max % of max % of max 50% ±10 58% ±9 87% ±4 CD51 % of max % of max % of max 3% ±1 47% ±3 72% ±7 CD140a % of max % of max % of max 39% ±6 51% ±3 71% ±3 CD140b % of max % of max % of max 4% ±2 1% ±1 7% ±1 CD34 % of max % of max % of max 21% ±16 30% ±10 42% ±10 gp38 % of max % of max % of max 50% ±4 43% ±7 55% ±5 Ly51 % of max % of max % of max 49% ±3 8% ±1 73% ±6 CD90.2 % of max % of max % of max 36% ±10 35% ±10 77% ±5 CD105 % of max % of max % of max 2% ±2 47% ±8 78% ±6 CD73 % of max % of max % of max 9% ±8 8% ±3 38% ±2 CD44 % of max % of max % of max 48% ±3 27% ±1 52% ±4 CD146 % of max % of max % of max 17% ±13 11% ±8 28% ±7 Nestin % of max % of max % of max 6% ±5 23% ±29 37% ±25 CXCL12 % of max % of max % of max SUPPLEMENTAL FIGURE 1. Flow cytometry analysis of Lin- Sca-1- cells from the thymus, bone and skin. Overlay histograms illustrate staining with the relevant antibody (in blue) and an isotype control (in red). Each overlay histogram is representative of three independent experiments (3-5 mice per biological replicate). Numbers represent the mean percentage of positive cells (+/- SD). Supplemental gure 2 A B 1000 250k M 100 tMC 98.2% K P bMC 200k R 10 sMC 1 A - 150k 1 9 1 a 4 8 1 0 5 A 2 5 0 3 3 5 9 0 C S CD CD CD gp Ly CD CD1 FSC 100k CD14 CD140b 1000 tMC 50k bMC M 100 sMC K 0 P Thymocytes R 10 -103 0 103 104 105 cTEC 1 mTEC SCA-1 t 5 e 1 m 1 i 4 3 3 n -k ca x c CD CD CD p o E F C tMC vs bMC tMC vs sMC bMC vs sMC 7 7 ) ) ) 6 695 6 6 365 5 426 sMC sMC bMC 5 5 4 4 4 3 3 3 2 2 2 (ReadCount (ReadCount 362 10 10 1 849 1 1 573 log log 0 log10(ReadCount 0 r = 0.476 0 r = 0.629 r = 0.607 0 1 2 3 4 5 6 0 1 2 3 4 5 6 0 1 2 3 4 5 6 log (ReadCount tMC) log10(ReadCount bMC) log10(ReadCount tMC) 10 SUPPLEMENTAL FIGURE 2. -

SUPPLEMENTARY INFORMATION Gotree/Goalign
SUPPLEMENTARY INFORMATION Gotree/Goalign : Toolkit and Go API to facilitate the development of phylogenetic workflows Frédéric Lemoine1,2∗ and Olivier Gascuel1,3 1 Unité de Bioinformatique Évolutive, Département de Biologie Computationnelle, Institut Pasteur, Paris, FRANCE, 2 Hub de Bioinformatique et Biostatistique, Département de Biologie Computationnelle, Institut Pasteur, Paris, FRANCE, 3 Current address: Institut de Systématique, Evolution, Biodiversité (ISYEB - UMR 7205), CNRS & Muséum National d’Histoire Naturelle, Paris, FRANCE *To whom correspondence should be addressed: [email protected] Supp. Text 1: Examples of Gotree/Goalign commands pp. 2-4 Supp Figure 1: Representation of the use case workflow and command templates pp. 5-6 Supp. Data 1: Nextflow implementation of the use case pp. 7-8 Supp. Data 2: List of analyzed primate species pp. 9 Supp. Data 3: List of 1,315 orthologous groups from OrthoDB pp. 10-15 1 Supplementary Text 1: Examples of Gotree/Goalign commands The comprehensive list of Gotree/Goalign commands is given on their respective GitHub repositories: https://github.com/evolbioinfo/gotree/blob/master/docs/index.md https://github.com/evolbioinfo/goalign/blob/master/docs/index.md 1) Reformatting a tree from newick to nexus1 gotree reformat nexus -i itol://129215302173073111930481660 The input tree is directly downloaded from iTOL, using its identifier and reformatted in Newick locally. 2) Reformatting an alignment from Fasta to Phylip1 goalign reformat phylip -i https://github.com/evolbioinfo/goalign/raw/master/tests/data/test_xz.xz -

Targeted Disruption of Lynx2 Reveals Distinct Functions for Lynx Homologues in Learning and Behavior Ayse Begum Tekinay
Rockefeller University Digital Commons @ RU Student Theses and Dissertations 2007 Targeted Disruption of Lynx2 Reveals Distinct Functions for Lynx Homologues in Learning and Behavior Ayse Begum Tekinay Follow this and additional works at: http://digitalcommons.rockefeller.edu/ student_theses_and_dissertations Part of the Life Sciences Commons Recommended Citation Tekinay, Ayse Begum, "Targeted Disruption of Lynx2 Reveals Distinct Functions for Lynx Homologues in Learning and Behavior" (2007). Student Theses and Dissertations. Paper 28. This Thesis is brought to you for free and open access by Digital Commons @ RU. It has been accepted for inclusion in Student Theses and Dissertations by an authorized administrator of Digital Commons @ RU. For more information, please contact [email protected]. TARGETED DISRUPTION OF LYNX2 REVEALS DISTINCT FUNCTIONS FOR LYNX HOMOLOGUES IN LEARNING AND BEHAVIOR A Thesis Presented to the Faculty of The Rockefeller University In Partial Fulfillment of the Requirements for the degree of Doctor of Philosophy by Ayse Begum Tekinay June 2007 © Copyright by Ayse Begum Tekinay 2007 TARGETED DISRUPTION OF LYNX2 REVEALS DISTINCT FUNCTIONS FOR LYNX HOMOLOGUES IN LEARNING AND BEHAVIOR Ayse Begum Tekinay, Ph.D. The Rockefeller University 2007 Endogenous short peptide modulators of ion channels provide a new level of regulation of nervous function. Lynx1 was identified as an endogenous mammalian homologue of snake venom peptide neurotoxins capable of binding to and functionally modulating nicotinic acetylcholine receptors (nAChR). Lynx1 is a member of the Ly6- α−neurotoxin superfamily (Ly6SF) of genes. Through extensive database searches, I identified 85 members of this superfamily including previously unidentified vertebrate and invertebrate family members. I show that these proteins are very divergent in their sequences, and identify two conserved subfamilies, snake toxins and immune system expressed Ly6 genes through phylogenetic inference. -
Evolutionary Analysis of the CAP Superfamily of Proteins Using Amino Acid Sequences
Evolutionary Analysis of the CAP Superfamily of Proteins using Amino Acid Sequences and Splice Sites by Anup Abraham A Thesis Presented in Partial Fulfillment of the Requirements for the Degree Master of Science Approved April 2016 by the Graduate Supervisory Committee: Douglas E. Chandler, Chair Kenneth H. Buetow Robert W. Roberson ARIZONA STATE UNIVERSITY May 2016 ABSTRACT Here I document the breadth of the CAP (Cysteine-RIch Secretory Proteins (CRISP), Antigen 5 (Ag5), and the Pathogenesis-Related 1 (PR)) protein superfamily and trace some of the major events in the evolution of this family with particular focus on vertebrate CRISP proteins. Specifically, I sought to study the origin of these CAP subfamilies using both amino acid sequence data and gene structure data, more precisely the positions of exon/intron borders within their genes. Counter to current scientific understanding, I find that the wide variety of CAP subfamilies present in mammals, where they were originally discovered and characterized, have distinct homologues in the invertebrate phyla contrary to the common assumption that these are vertebrate protein subfamilies. In addition, I document the fact that primitive eukaryotic CAP genes contained only one exon, likely inherited from prokaryotic SCP-domain containing genes which were, by nature, free of introns. As evolution progressed, an increasing number of introns were inserted into CAP genes, reaching 2 to 5 in the invertebrate world, and 5 to 15 in the vertebrate world. Lastly, phylogenetic relationships between these proteins appear to be traceable not only by amino acid sequence homology but also by preservation of exon number and exon borders within their genes. -

4 353 Skin Oral 1 B
A B Supplementary Figure S1: Differentially expressed piRNAs during skin and oral mucosal wound healing. (A) piRNA 0hr-1 0hr-3 0hr-2 24hr-2 24hr-1 24hr-3 5day-3 5day-1 5day-2 0hr-1 0hr-2 0hr-3 24hr-2 24hr-1 24hr-3 5day-1 5day-2 5day-3 profiles were obtained on mouse skin and oral mucosal (palate) wound healing time course (0hr, 24 hr, and 5 day). A total of 357 differentially expressed piRNA were identified during skin wound healing (Bonferroni adjusted P value <0.05). See Supplementary Table 1A for the full list. C (B) Five differentially expressed piRNA skin were identified during oral mucosal wound healing (P value <0.01, list presented in Supplementary Table 1B). Note: more 353 stringent statistical cut-off ((Bonferroni adjusted P value) yield 0 differentially expressed piRNA gene. (C) Venn diagram illustrates overlaps between differentially 4 expressedpiRNAsinskinandoral oral 1 mucosal wound healing. min max Supplementary Table S1a: Differentially expressed piRNAs in skin wound healing Mean StDev piRNA 0 hr 24 hr 5 day 0 hr 24 hr 5 day pVal adj P piR‐mmu‐15927330 5.418351 11.39746 10.799 0.34576 0.253492 0.154802 6.32E‐14 6.97E‐11 piR‐mmu‐49559417 5.301647 10.10777 9.816878 0.441719 0.222335 0.032479 3.95E‐12 4.36E‐09 piR‐mmu‐30053093 6.32531 11.26384 4.020902 0.280841 1.057847 0.178798 5.21E‐12 5.76E‐09 piR‐mmu‐29303577 5.15005 10.47662 9.52175 0.554877 0.26622 0.163283 1.53E‐11 1.69E‐08 piR‐mmu‐49254706 5.187673 10.19644 9.622374 0.520671 0.330378 0.191116 1.96E‐11 2.16E‐08 piR‐mmu‐49005170 5.415133 9.639725 9.565967 0.411507 0.281143 -

GPRASP/ARMCX Protein Family: Potential Involvement in Health And
GPRASP/ARMCX protein family: potential involvement in health and diseases revealed by their novel interacting partners Juliette Kaeffer, Gabrielle Zeder-Lutz, Frederic Simonin, Sandra Lecat To cite this version: Juliette Kaeffer, Gabrielle Zeder-Lutz, Frederic Simonin, Sandra Lecat. GPRASP/ARMCX pro- tein family: potential involvement in health and diseases revealed by their novel interacting part- ners. Current Topics in Medicinal Chemistry, Bentham Science Publishers, 2021, 21 (3), pp.227-254. 10.2174/1568026620666201202102448. hal-03095663 HAL Id: hal-03095663 https://hal.archives-ouvertes.fr/hal-03095663 Submitted on 4 Jan 2021 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. GPRASP/ARMCX protein family: potential involvement in health and diseases revealed by their novel interacting partners Juliette Kaeffer, Gabrielle Zeder-Lutz, Frédéric Simonin* and Sandra Lecat* Affiliation : Biotechnologie et Signalisation Cellulaire, UMR7242 CNRS, Université de Strasbourg, Illkirch-Graffenstaden, France. * Corresponding authors: Sandra Lecat ([email protected]) and Frédéric Simonin ([email protected]) : Biotechnologie et Signalisation Cellulaire, UMR7242 CNRS / Université de Strasbourg, 300 boulevard Sébastien Brant, CS 10413, 67412 Illkirch-Graffenstaden, Cedex, France. 1 Abstract GPRASP (GPCR-associated sorting protein)/ARMCX (ARMadillo repeat-Containing proteins on the X chromosome) family is composed of 10 proteins, which genes are located on a small locus of the X chromosome except one.