NME Proteins Regulate Class Switch Recombination
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

Protein Identities in Evs Isolated from U87-MG GBM Cells As Determined by NG LC-MS/MS
Protein identities in EVs isolated from U87-MG GBM cells as determined by NG LC-MS/MS. No. Accession Description Σ Coverage Σ# Proteins Σ# Unique Peptides Σ# Peptides Σ# PSMs # AAs MW [kDa] calc. pI 1 A8MS94 Putative golgin subfamily A member 2-like protein 5 OS=Homo sapiens PE=5 SV=2 - [GG2L5_HUMAN] 100 1 1 7 88 110 12,03704523 5,681152344 2 P60660 Myosin light polypeptide 6 OS=Homo sapiens GN=MYL6 PE=1 SV=2 - [MYL6_HUMAN] 100 3 5 17 173 151 16,91913397 4,652832031 3 Q6ZYL4 General transcription factor IIH subunit 5 OS=Homo sapiens GN=GTF2H5 PE=1 SV=1 - [TF2H5_HUMAN] 98,59 1 1 4 13 71 8,048185945 4,652832031 4 P60709 Actin, cytoplasmic 1 OS=Homo sapiens GN=ACTB PE=1 SV=1 - [ACTB_HUMAN] 97,6 5 5 35 917 375 41,70973209 5,478027344 5 P13489 Ribonuclease inhibitor OS=Homo sapiens GN=RNH1 PE=1 SV=2 - [RINI_HUMAN] 96,75 1 12 37 173 461 49,94108966 4,817871094 6 P09382 Galectin-1 OS=Homo sapiens GN=LGALS1 PE=1 SV=2 - [LEG1_HUMAN] 96,3 1 7 14 283 135 14,70620005 5,503417969 7 P60174 Triosephosphate isomerase OS=Homo sapiens GN=TPI1 PE=1 SV=3 - [TPIS_HUMAN] 95,1 3 16 25 375 286 30,77169764 5,922363281 8 P04406 Glyceraldehyde-3-phosphate dehydrogenase OS=Homo sapiens GN=GAPDH PE=1 SV=3 - [G3P_HUMAN] 94,63 2 13 31 509 335 36,03039959 8,455566406 9 Q15185 Prostaglandin E synthase 3 OS=Homo sapiens GN=PTGES3 PE=1 SV=1 - [TEBP_HUMAN] 93,13 1 5 12 74 160 18,68541938 4,538574219 10 P09417 Dihydropteridine reductase OS=Homo sapiens GN=QDPR PE=1 SV=2 - [DHPR_HUMAN] 93,03 1 1 17 69 244 25,77302971 7,371582031 11 P01911 HLA class II histocompatibility antigen, -

The Function of NM23-H1/NME1 and Its Homologs in Major Processes Linked to Metastasis
University of Dundee The Function of NM23-H1/NME1 and Its Homologs in Major Processes Linked to Metastasis Mátyási, Barbara; Farkas, Zsolt; Kopper, László; Sebestyén, Anna; Boissan, Mathieu; Mehta, Anil Published in: Pathology and Oncology Research DOI: 10.1007/s12253-020-00797-0 Publication date: 2020 Licence: CC BY Document Version Publisher's PDF, also known as Version of record Link to publication in Discovery Research Portal Citation for published version (APA): Mátyási, B., Farkas, Z., Kopper, L., Sebestyén, A., Boissan, M., Mehta, A., & Takács-Vellai, K. (2020). The Function of NM23-H1/NME1 and Its Homologs in Major Processes Linked to Metastasis. Pathology and Oncology Research, 26(1), 49-61. https://doi.org/10.1007/s12253-020-00797-0 General rights Copyright and moral rights for the publications made accessible in Discovery Research Portal are retained by the authors and/or other copyright owners and it is a condition of accessing publications that users recognise and abide by the legal requirements associated with these rights. • Users may download and print one copy of any publication from Discovery Research Portal for the purpose of private study or research. • You may not further distribute the material or use it for any profit-making activity or commercial gain. • You may freely distribute the URL identifying the publication in the public portal. Take down policy If you believe that this document breaches copyright please contact us providing details, and we will remove access to the work immediately and investigate your -

Development and Validation of a Protein-Based Risk Score for Cardiovascular Outcomes Among Patients with Stable Coronary Heart Disease
Supplementary Online Content Ganz P, Heidecker B, Hveem K, et al. Development and validation of a protein-based risk score for cardiovascular outcomes among patients with stable coronary heart disease. JAMA. doi: 10.1001/jama.2016.5951 eTable 1. List of 1130 Proteins Measured by Somalogic’s Modified Aptamer-Based Proteomic Assay eTable 2. Coefficients for Weibull Recalibration Model Applied to 9-Protein Model eFigure 1. Median Protein Levels in Derivation and Validation Cohort eTable 3. Coefficients for the Recalibration Model Applied to Refit Framingham eFigure 2. Calibration Plots for the Refit Framingham Model eTable 4. List of 200 Proteins Associated With the Risk of MI, Stroke, Heart Failure, and Death eFigure 3. Hazard Ratios of Lasso Selected Proteins for Primary End Point of MI, Stroke, Heart Failure, and Death eFigure 4. 9-Protein Prognostic Model Hazard Ratios Adjusted for Framingham Variables eFigure 5. 9-Protein Risk Scores by Event Type This supplementary material has been provided by the authors to give readers additional information about their work. Downloaded From: https://jamanetwork.com/ on 10/02/2021 Supplemental Material Table of Contents 1 Study Design and Data Processing ......................................................................................................... 3 2 Table of 1130 Proteins Measured .......................................................................................................... 4 3 Variable Selection and Statistical Modeling ........................................................................................ -

Assessing the Human Canonical Protein Count[Version 1; Peer Review
F1000Research 2017, 6:448 Last updated: 15 JUL 2020 REVIEW Last rolls of the yoyo: Assessing the human canonical protein count [version 1; peer review: 1 approved, 2 approved with reservations] Christopher Southan IUPHAR/BPS Guide to Pharmacology, Centre for Integrative Physiology, University of Edinburgh, Edinburgh, EH8 9XD, UK First published: 07 Apr 2017, 6:448 Open Peer Review v1 https://doi.org/10.12688/f1000research.11119.1 Latest published: 07 Apr 2017, 6:448 https://doi.org/10.12688/f1000research.11119.1 Reviewer Status Abstract Invited Reviewers In 2004, when the protein estimate from the finished human genome was 1 2 3 only 24,000, the surprise was compounded as reviewed estimates fell to 19,000 by 2014. However, variability in the total canonical protein counts version 1 (i.e. excluding alternative splice forms) of open reading frames (ORFs) in 07 Apr 2017 report report report different annotation portals persists. This work assesses these differences and possible causes. A 16-year analysis of Ensembl and UniProtKB/Swiss-Prot shows convergence to a protein number of ~20,000. The former had shown some yo-yoing, but both have now plateaued. Nine 1 Michael Tress, Spanish National Cancer major annotation portals, reviewed at the beginning of 2017, gave a spread Research Centre (CNIO), Madrid, Spain of counts from 21,819 down to 18,891. The 4-way cross-reference concordance (within UniProt) between Ensembl, Swiss-Prot, Entrez Gene 2 Elspeth A. Bruford , European Molecular and the Human Gene Nomenclature Committee (HGNC) drops to 18,690, Biology Laboratory, Hinxton, UK indicating methodological differences in protein definitions and experimental existence support between sources. -
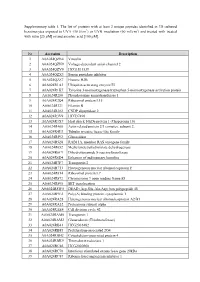
Supplementary Table 1. the List of Proteins with at Least 2 Unique
Supplementary table 1. The list of proteins with at least 2 unique peptides identified in 3D cultured keratinocytes exposed to UVA (30 J/cm2) or UVB irradiation (60 mJ/cm2) and treated with treated with rutin [25 µM] or/and ascorbic acid [100 µM]. Nr Accession Description 1 A0A024QZN4 Vinculin 2 A0A024QZN9 Voltage-dependent anion channel 2 3 A0A024QZV0 HCG1811539 4 A0A024QZX3 Serpin peptidase inhibitor 5 A0A024QZZ7 Histone H2B 6 A0A024R1A3 Ubiquitin-activating enzyme E1 7 A0A024R1K7 Tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein 8 A0A024R280 Phosphoserine aminotransferase 1 9 A0A024R2Q4 Ribosomal protein L15 10 A0A024R321 Filamin B 11 A0A024R382 CNDP dipeptidase 2 12 A0A024R3V9 HCG37498 13 A0A024R3X7 Heat shock 10kDa protein 1 (Chaperonin 10) 14 A0A024R408 Actin related protein 2/3 complex, subunit 2, 15 A0A024R4U3 Tubulin tyrosine ligase-like family 16 A0A024R592 Glucosidase 17 A0A024R5Z8 RAB11A, member RAS oncogene family 18 A0A024R652 Methylenetetrahydrofolate dehydrogenase 19 A0A024R6C9 Dihydrolipoamide S-succinyltransferase 20 A0A024R6D4 Enhancer of rudimentary homolog 21 A0A024R7F7 Transportin 2 22 A0A024R7T3 Heterogeneous nuclear ribonucleoprotein F 23 A0A024R814 Ribosomal protein L7 24 A0A024R872 Chromosome 9 open reading frame 88 25 A0A024R895 SET translocation 26 A0A024R8W0 DEAD (Asp-Glu-Ala-Asp) box polypeptide 48 27 A0A024R9E2 Poly(A) binding protein, cytoplasmic 1 28 A0A024RA28 Heterogeneous nuclear ribonucleoprotein A2/B1 29 A0A024RA52 Proteasome subunit alpha 30 A0A024RAE4 Cell division cycle 42 31 -

Adenovirus Strategies for Altering the Cellular Environment in Favor of Infection
University of Pennsylvania ScholarlyCommons Publicly Accessible Penn Dissertations 2019 Adenovirus Strategies For Altering The Cellular Environment In Favor Of Infection Christin Herrmann University of Pennsylvania Follow this and additional works at: https://repository.upenn.edu/edissertations Part of the Allergy and Immunology Commons, Immunology and Infectious Disease Commons, Medical Immunology Commons, and the Virology Commons Recommended Citation Herrmann, Christin, "Adenovirus Strategies For Altering The Cellular Environment In Favor Of Infection" (2019). Publicly Accessible Penn Dissertations. 3568. https://repository.upenn.edu/edissertations/3568 This paper is posted at ScholarlyCommons. https://repository.upenn.edu/edissertations/3568 For more information, please contact [email protected]. Adenovirus Strategies For Altering The Cellular Environment In Favor Of Infection Abstract Viruses, as obligate intracellular pathogens, rely on their host cell for successful replication. Viruses have evolved different strategies to hijack and redirect cellular processes to benefit infection and overcome host immune responses. Understanding the mechanisms by which viruses exploit their host cells will reveal new targets for antiviral therapies. In addition, these studies can provide insights into the regulation of fundamental cellular processes. While much progress has been made in this area, many unexpected nuances of virus-host interaction are still being discovered. Here, we employed several strategies to uncover new aspects of viral manipulation of the host environment by adenovirus, a nuclear-replicating DNA virus that commonly infects humans. The first project focused on how viral histone-like proteins impact cellular chromatin. Adenovirus encodes the small, basic protein VII that coats and condenses viral genomes. The effect of this viral DNA-binding protein on host chromatin structure and function had remained unexplored. -

S41467-020-17157-W.Pdf
ARTICLE https://doi.org/10.1038/s41467-020-17157-w OPEN Transcriptional activity and strain-specific history of mouse pseudogenes Cristina Sisu 1,2,3,15, Paul Muir4,5,15, Adam Frankish6, Ian Fiddes7, Mark Diekhans 7, David Thybert6,8, Duncan T. Odom 9,10, Paul Flicek 6,10, Thomas M. Keane 6, Tim Hubbard 11, Jennifer Harrow12 & ✉ Mark Gerstein 1,2,5,13,14 Pseudogenes are ideal markers of genome remodelling. In turn, the mouse is an ideal plat- 1234567890():,; form for studying them, particularly with the recent availability of strain-sequencing and transcriptional data. Here, combining both manual curation and automatic pipelines, we present a genome-wide annotation of the pseudogenes in the mouse reference genome and 18 inbred mouse strains (available via the mouse.pseudogene.org resource). We also annotate 165 unitary pseudogenes in mouse, and 303, in human. The overall pseudogene repertoire in mouse is similar to that in human in terms of size, biotype distribution, and family composition (e.g. with GAPDH and ribosomal proteins being the largest families). Notable differences arise in the pseudogene age distribution, with multiple retro- transpositional bursts in mouse evolutionary history and only one in human. Furthermore, in each strain about a fifth of all pseudogenes are unique, reflecting strain-specific evolution. Finally, we find that ~15% of the mouse pseudogenes are transcribed, and that highly tran- scribed parent genes tend to give rise to many processed pseudogenes. 1 Program in Computational Biology and Bioinformatics, Yale University, New Haven, CT 06520, USA. 2 Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, CT 06520, USA. -

Transcriptomic Analysis of Early Stages of Intestinal Regeneration in Holothuria Glaberrima David J
Transcriptomic Analysis of Early Stages of Intestinal Regeneration in Holothuria glaberrima David J. Quispe-Parra1, Joshua G. Medina-Feliciano1, Sebastián Cruz-González1, Humberto Ortiz-Zuazaga2, José E. García-Arrarás1* 1University of Puerto Rico, Biology Department, San Juan, 00925, Puerto Rico. 2University of Puerto Rico, Department of Computer Sciences, San Juan, 00925, Puerto Rico. Table S1. Results of transcriptome assessment with BUSCO Parameter BUSCO result Core genes queried 978 Complete core genes detected 99.1% Complete single copy core genes 27.2% Complete duplicated core genes 71.9% Fragmented core genes detected 0.4% Missing core genes 0.5% Table S2. Transcriptome length statistics and composition assessments with gVolante Parameter Result Number of sequences 491 436 Total length (nt) 408 930 895 Longest sequence (nt) 34 610 Shortest sequence (nt) 200 Mean sequence length (nt) 832 N50 sequence length (nt) 1 691 Table S3. RNA-seq data read statistic values Accession Quantity of reads Sample Mapped reads (SRA) Before Filtering After Filtering SRR12564573 NormalA 89 424 588.00 88 112 720.00 94.18% SRR12564572 NormalB 74 105 862.00 72 983 434.00 93.79% SRR12564570 NormalC 60 626 094.00 59 767 896.00 91.55% SRR12564564 Day1A 32 499 910.00 31 721 976.00 90.67% SRR12564563 Day1B 35 164 514.00 34 237 960.00 91.37% SRR12564571 Day1C 43 519 280.00 42 179 220.00 91.53% SRR12564567 Day3A 64 094 536.00 62 995 792.00 90.26% SRR12564566 Day3B 71 654 462.00 70 158 078.00 91.07% SRR12564565 Day3C 71 259 560.00 70 418 332.00 90.75% SRR12564569 Day3D* 107 688 184.00 106 083 898.00 - SRR12564568 Day3E* 108 372 334.00 106 767 522.00 - *Samples used for the assembly but not for differential expression analysis Table S4. -

A Chromosome-Centric Human Proteome Project (C-HPP) To
computational proteomics Laboratory for Computational Proteomics www.FenyoLab.org E-mail: [email protected] Facebook: NYUMC Computational Proteomics Laboratory Twitter: @CompProteomics Perspective pubs.acs.org/jpr A Chromosome-centric Human Proteome Project (C-HPP) to Characterize the Sets of Proteins Encoded in Chromosome 17 † ‡ § ∥ ‡ ⊥ Suli Liu, Hogune Im, Amos Bairoch, Massimo Cristofanilli, Rui Chen, Eric W. Deutsch, # ¶ △ ● § † Stephen Dalton, David Fenyo, Susan Fanayan,$ Chris Gates, , Pascale Gaudet, Marina Hincapie, ○ ■ △ ⬡ ‡ ⊥ ⬢ Samir Hanash, Hoguen Kim, Seul-Ki Jeong, Emma Lundberg, George Mias, Rajasree Menon, , ∥ □ △ # ⬡ ▲ † Zhaomei Mu, Edouard Nice, Young-Ki Paik, , Mathias Uhlen, Lance Wells, Shiaw-Lin Wu, † † † ‡ ⊥ ⬢ ⬡ Fangfei Yan, Fan Zhang, Yue Zhang, Michael Snyder, Gilbert S. Omenn, , Ronald C. Beavis, † # and William S. Hancock*, ,$, † Barnett Institute and Department of Chemistry and Chemical Biology, Northeastern University, Boston, Massachusetts 02115, United States ‡ Stanford University, Palo Alto, California, United States § Swiss Institute of Bioinformatics (SIB) and University of Geneva, Geneva, Switzerland ∥ Fox Chase Cancer Center, Philadelphia, Pennsylvania, United States ⊥ Institute for System Biology, Seattle, Washington, United States ¶ School of Medicine, New York University, New York, United States $Department of Chemistry and Biomolecular Sciences, Macquarie University, Sydney, NSW, Australia ○ MD Anderson Cancer Center, Houston, Texas, United States ■ Yonsei University College of Medicine, Yonsei University, -

Analysis of the Mouse Transcriptome Based on Functional Annotation of 60,770 Full-Length Cdnas
articles Analysis of the mouse transcriptome based on functional annotation of 60,770 full-length cDNAs The FANTOM Consortium and the RIKEN Genome Exploration Research Group Phase I & II Team* *A full list of authors appears at the end of this paper ........................................................................................................................................................................................................................... Only a small proportion of the mouse genome is transcribed into mature messenger RNA transcripts. There is an international collaborative effort to identify all full-length mRNA transcripts from the mouse, and to ensure that each is represented in a physical collection of clones. Here we report the manual annotation of 60,770 full-length mouse complementary DNA sequences. These are clustered into 33,409 ‘transcriptional units’, contributing 90.1% of a newly established mouse transcriptome database. Of these transcriptional units, 4,258 are new protein-coding and 11,665 are new non-coding messages, indicating that non-coding RNA is a major component of the transcriptome. 41% of all transcriptional units showed evidence of alternative splicing. In protein-coding transcripts, 79% of splice variations altered the protein product. Whole-transcriptome analyses resulted in the identification of 2,431 sense–antisense pairs. The present work, completely supported by physical clones, provides the most comprehensive survey of a mammalian transcriptome so far, and is a valuable resource for functional genomics. With the availability of draft sequences of the human genome1,2, 39,694 new cDNA clones (FANTOM2 new set) and the 21,076 increasing attention has focused on identifying the complete set of FANTOM1 clone set. We also present a global analysis of the mouse mammalian genes, both protein-coding and non-protein-coding. -

Automatic Functional Annotation of Predicted Active Sites: Combining PDB and Literature Mining
Automatic functional annotation of predicted active sites: combining PDB and literature mining Kevin Nagel Wolfson College A dissertation submitted to the University of Cambridge for the degree of Doctor of Philosophy European Molecular Biology Laboratory, European Bioinformatics Institute, Wellcome Trust Genome Campus, Hinxton, Cambridge CB10 1SD, United Kingdom. Email: [email protected] January 2009 Declaration This dissertation is the result of my own work, and includes nothing which is the outcome of work done in collaboration, except where specifically indicated in the text. The disser- tation does not exceed the specified length limit of 300 pages as defined by the Biology Degree Committee. This thesis has been typeset in 12pt font using LATEX 2"according to the specifications defined by the Board of Graduate Studies and the Biology Degree Committee. 1 Summary Kevin Nagel European Bioinformatics Institute University of Cambridge Dissertation title: Automatic functional annotation of predicted active sites: combining PDB and literature mining. Proteins are essential to cell functions, which is mainly identified in biological experiments. The structural models for proteins help to explain their function, but are not direct evidence for their function. Nonetheless, we can mine structural databases, such as Protein Data Bank (PDB), to filter out shared structural components that are meaningful with regards to the protein function. This thesis applied mining techniques to PDB to identify evolutionary conserved struc- tural patterns, e.g. active sites. This analysis retrieved 3- and 4-bodies with assumed two- and three-way residue interaction that have been selected from a distribution analysis of residue triplets. A subset of the mined patterns is assumed to represent an active site, which should be confirmed by annotations gathered by automatic literature analysis. -

Nucleoside Diphosphate Kinases (Ndpks) in Animal Development
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Repository of the Academy's Library Nucleoside diphosphate kinases (NDPKs) in animal development Krisztina Takács-Vellai, Tibor Vellai, Zsolt Farkas & Anil Mehta Cellular and Molecular Life Sciences ISSN 1420-682X Volume 72 Number 8 Cell. Mol. Life Sci. (2015) 72:1447-1462 DOI 10.1007/s00018-014-1803-0 1 23 Your article is protected by copyright and all rights are held exclusively by Springer Basel. This e-offprint is for personal use only and shall not be self-archived in electronic repositories. If you wish to self-archive your article, please use the accepted manuscript version for posting on your own website. You may further deposit the accepted manuscript version in any repository, provided it is only made publicly available 12 months after official publication or later and provided acknowledgement is given to the original source of publication and a link is inserted to the published article on Springer's website. The link must be accompanied by the following text: "The final publication is available at link.springer.com”. 1 23 Author's personal copy Cell. Mol. Life Sci. (2015) 72:1447–1462 DOI 10.1007/s00018-014-1803-0 Cellular and Molecular Life Sciences REVIEW Nucleoside diphosphate kinases (NDPKs) in animal development Krisztina Taka´cs-Vellai • Tibor Vellai • Zsolt Farkas • Anil Mehta Received: 21 July 2014 / Revised: 4 December 2014 / Accepted: 8 December 2014 / Published online: 24 December 2014 Ó Springer Basel 2014 Abstract In textbooks of biochemistry, nucleoside Introduction diphosphate conversion to a triphosphate by nucleoside diphosphate ‘kinases’ (NDPKs, also named NME or NM23 Nucleoside diphosphate kinases (NDPKs) were identified proteins) merits a few lines of text.