Systematic Detection of Brain Protein-Coding Genes Under Positive Selection During
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Differences Between Human and Chimpanzee Genomes and Their Implications in Gene Expression, Protein Functions and Biochemical Properties of the Two Species Maria V
Suntsova and Buzdin BMC Genomics 2020, 21(Suppl 7):535 https://doi.org/10.1186/s12864-020-06962-8 REVIEW Open Access Differences between human and chimpanzee genomes and their implications in gene expression, protein functions and biochemical properties of the two species Maria V. Suntsova1 and Anton A. Buzdin1,2,3,4* From 11th International Young Scientists School “Systems Biology and Bioinformatics”–SBB-2019 Novosibirsk, Russia. 24-28 June 2019 Abstract Chimpanzees are the closest living relatives of humans. The divergence between human and chimpanzee ancestors dates to approximately 6,5–7,5 million years ago. Genetic features distinguishing us from chimpanzees and making us humans are still of a great interest. After divergence of their ancestor lineages, human and chimpanzee genomes underwent multiple changes including single nucleotide substitutions, deletions and duplications of DNA fragments of different size, insertion of transposable elements and chromosomal rearrangements. Human-specific single nucleotide alterations constituted 1.23% of human DNA, whereas more extended deletions and insertions cover ~ 3% of our genome. Moreover, much higher proportion is made by differential chromosomal inversions and translocations comprising several megabase-long regions or even whole chromosomes. However, despite of extensive knowledge of structural genomic changes accompanying human evolution we still cannot identify with certainty the causative genes of human identity. Most structural gene-influential changes happened at the level of expression regulation, which in turn provoked larger alterations of interactome gene regulation networks. In this review, we summarized the available information about genetic differences between humans and chimpanzees and their potential functional impacts on differential molecular, anatomical, physiological and cognitive peculiarities of these species. -

Protein Identities in Evs Isolated from U87-MG GBM Cells As Determined by NG LC-MS/MS
Protein identities in EVs isolated from U87-MG GBM cells as determined by NG LC-MS/MS. No. Accession Description Σ Coverage Σ# Proteins Σ# Unique Peptides Σ# Peptides Σ# PSMs # AAs MW [kDa] calc. pI 1 A8MS94 Putative golgin subfamily A member 2-like protein 5 OS=Homo sapiens PE=5 SV=2 - [GG2L5_HUMAN] 100 1 1 7 88 110 12,03704523 5,681152344 2 P60660 Myosin light polypeptide 6 OS=Homo sapiens GN=MYL6 PE=1 SV=2 - [MYL6_HUMAN] 100 3 5 17 173 151 16,91913397 4,652832031 3 Q6ZYL4 General transcription factor IIH subunit 5 OS=Homo sapiens GN=GTF2H5 PE=1 SV=1 - [TF2H5_HUMAN] 98,59 1 1 4 13 71 8,048185945 4,652832031 4 P60709 Actin, cytoplasmic 1 OS=Homo sapiens GN=ACTB PE=1 SV=1 - [ACTB_HUMAN] 97,6 5 5 35 917 375 41,70973209 5,478027344 5 P13489 Ribonuclease inhibitor OS=Homo sapiens GN=RNH1 PE=1 SV=2 - [RINI_HUMAN] 96,75 1 12 37 173 461 49,94108966 4,817871094 6 P09382 Galectin-1 OS=Homo sapiens GN=LGALS1 PE=1 SV=2 - [LEG1_HUMAN] 96,3 1 7 14 283 135 14,70620005 5,503417969 7 P60174 Triosephosphate isomerase OS=Homo sapiens GN=TPI1 PE=1 SV=3 - [TPIS_HUMAN] 95,1 3 16 25 375 286 30,77169764 5,922363281 8 P04406 Glyceraldehyde-3-phosphate dehydrogenase OS=Homo sapiens GN=GAPDH PE=1 SV=3 - [G3P_HUMAN] 94,63 2 13 31 509 335 36,03039959 8,455566406 9 Q15185 Prostaglandin E synthase 3 OS=Homo sapiens GN=PTGES3 PE=1 SV=1 - [TEBP_HUMAN] 93,13 1 5 12 74 160 18,68541938 4,538574219 10 P09417 Dihydropteridine reductase OS=Homo sapiens GN=QDPR PE=1 SV=2 - [DHPR_HUMAN] 93,03 1 1 17 69 244 25,77302971 7,371582031 11 P01911 HLA class II histocompatibility antigen, -

Molecular Evolutionary Routes That Lead to Innovations
International Journal of Evolutionary Biology Molecular Evolutionary Routes That Lead to Innovations Guest Editors: Frédéric Brunet, Hideki Innan, Ben-Yang Liao, and Wen Wang Molecular Evolutionary Routes That Lead to Innovations International Journal of Evolutionary Biology Molecular Evolutionary Routes That Lead to Innovations Guest Editors: Fred´ eric´ Brunet, Hideki Innan, Ben-YangLiao, and Wen Wang Copyright © 2012 Hindawi Publishing Corporation. All rights reserved. This is a special issue published in “International Journal of Evolutionary Biology.” All articles are open access articles distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Editorial Board Giacomo Bernardi, USA Kazuho Ikeo, Japan Jeffrey R. Powell, USA Terr y Burke, UK Yoh Iwasa, Japan Hudson Kern Reeve, USA Ignacio Doadrio, Spain Henrik J. Jensen, UK Y. Satta, Japan Simon Easteal, Australia Amitabh Joshi, India Koji Tamura, Japan Santiago F. Elena, Spain Hirohisa Kishino, Japan Yoshio Tateno, Japan Renato Fani, Italy A. Moya, Spain E. N. Trifonov, Israel Dmitry A. Filatov, UK G. Pesole, Italy Eske Willerslev, Denmark F. Gonzalez-Candelas,´ Spain I. Popescu, USA Shozo Yokoyama, Japan D. Graur, USA David Posada, Spain Contents Molecular Evolutionary Routes That Lead to Innovations,Fred´ eric´ Brunet, Hideki Innan, Ben-Yang Liao, and Wen Wang Volume 2012, Article ID 483176, 2 pages Purifying Selection Bias against Microsatellites in Gene -

The Function of NM23-H1/NME1 and Its Homologs in Major Processes Linked to Metastasis
University of Dundee The Function of NM23-H1/NME1 and Its Homologs in Major Processes Linked to Metastasis Mátyási, Barbara; Farkas, Zsolt; Kopper, László; Sebestyén, Anna; Boissan, Mathieu; Mehta, Anil Published in: Pathology and Oncology Research DOI: 10.1007/s12253-020-00797-0 Publication date: 2020 Licence: CC BY Document Version Publisher's PDF, also known as Version of record Link to publication in Discovery Research Portal Citation for published version (APA): Mátyási, B., Farkas, Z., Kopper, L., Sebestyén, A., Boissan, M., Mehta, A., & Takács-Vellai, K. (2020). The Function of NM23-H1/NME1 and Its Homologs in Major Processes Linked to Metastasis. Pathology and Oncology Research, 26(1), 49-61. https://doi.org/10.1007/s12253-020-00797-0 General rights Copyright and moral rights for the publications made accessible in Discovery Research Portal are retained by the authors and/or other copyright owners and it is a condition of accessing publications that users recognise and abide by the legal requirements associated with these rights. • Users may download and print one copy of any publication from Discovery Research Portal for the purpose of private study or research. • You may not further distribute the material or use it for any profit-making activity or commercial gain. • You may freely distribute the URL identifying the publication in the public portal. Take down policy If you believe that this document breaches copyright please contact us providing details, and we will remove access to the work immediately and investigate your -

Differential Gene Expression in Patients with Prostate Cancer and in Patients with Parkinson Disease: an Example of Inverse Comorbidity
Differential Gene Expression in Patients With Prostate Cancer and in Patients With Parkinson Disease: an Example of Inverse Comorbidity Pietro Pepe Ospedale Cannizzaro Simona Vetrano Ospedale Cannizzaro Rossella Cannarella University of Catania Aldo E Calogero University of Catania Giovanna Marchese University of Salerno Maria Ravo University of Salerno Filippo Fraggetta Ospedale Cannizzaro Ludovica Pepe Ospedale Cannizzaro Michele Pennisi Ospedale Cannizzaro Corrado Romano Oasi Maria SS Raffaele Ferri Oasi Maria SS Michele Salemi ( [email protected] ) Oasi Maria SS Research Article Keywords: Prostate cancer, Parkinson disease, Inverse comorbidity, NGS Posted Date: March 11th, 2021 DOI: https://doi.org/10.21203/rs.3.rs-289371/v1 Page 1/11 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 2/11 Abstract Prostate cancer (PCa) is one of the leading causes of death in Western countries. Environmental and genetic factors play a pivotal role in PCa etiology. Timely identication of the genetic causes is useful for an early diagnosis. Parkinson’s disease (PD) is the most frequent neurodegenerative movement disorder; it is associated with the presence of Lewy bodies (LBs) and genetic factors are involved in its pathogenesis. Several studies have indicated that the expression of target genes in patients with PD is inversely related to cancer development; this phenomenon has been named “inverse comorbidity”. The present study was undertaken to evaluate whether a genetic dysregulation occurs in opposite directions in patients with PD or PCa. In the present study, next-generation sequencing (NGS) transcriptome analysis was used to assess whether a genetic dysregulation in opposite directions occurs in patients with PD or PCa. -

Placenta-Derived Exosomes Continuously Increase in Maternal
Sarker et al. Journal of Translational Medicine 2014, 12:204 http://www.translational-medicine.com/content/12/1/204 RESEARCH Open Access Placenta-derived exosomes continuously increase in maternal circulation over the first trimester of pregnancy Suchismita Sarker1, Katherin Scholz-Romero1, Alejandra Perez2, Sebastian E Illanes1,2,3, Murray D Mitchell1, Gregory E Rice1,2 and Carlos Salomon1,2* Abstract Background: Human placenta releases specific nanovesicles (i.e. exosomes) into the maternal circulation during pregnancy, however, the presence of placenta-derived exosomes in maternal blood during early pregnancy remains to be established. The aim of this study was to characterise gestational age related changes in the concentration of placenta-derived exosomes during the first trimester of pregnancy (i.e. from 6 to 12 weeks) in plasma from women with normal pregnancies. Methods: A time-series experimental design was used to establish pregnancy-associated changes in maternal plasma exosome concentrations during the first trimester. A series of plasma were collected from normal healthy women (10 patients) at 6, 7, 8, 9, 10, 11 and 12 weeks of gestation (n = 70). We measured the stability of these vesicles by quantifying and observing their protein and miRNA contents after the freeze/thawing processes. Exosomes were isolated by differential and buoyant density centrifugation using a sucrose continuous gradient and characterised by their size distribution and morphology using the nanoparticles tracking analysis (NTA; Nanosight™) and electron microscopy (EM), respectively. The total number of exosomes and placenta-derived exosomes were determined by quantifying the immunoreactive exosomal marker, CD63 and a placenta-specific marker (Placental Alkaline Phosphatase PLAP). -

Development and Validation of a Protein-Based Risk Score for Cardiovascular Outcomes Among Patients with Stable Coronary Heart Disease
Supplementary Online Content Ganz P, Heidecker B, Hveem K, et al. Development and validation of a protein-based risk score for cardiovascular outcomes among patients with stable coronary heart disease. JAMA. doi: 10.1001/jama.2016.5951 eTable 1. List of 1130 Proteins Measured by Somalogic’s Modified Aptamer-Based Proteomic Assay eTable 2. Coefficients for Weibull Recalibration Model Applied to 9-Protein Model eFigure 1. Median Protein Levels in Derivation and Validation Cohort eTable 3. Coefficients for the Recalibration Model Applied to Refit Framingham eFigure 2. Calibration Plots for the Refit Framingham Model eTable 4. List of 200 Proteins Associated With the Risk of MI, Stroke, Heart Failure, and Death eFigure 3. Hazard Ratios of Lasso Selected Proteins for Primary End Point of MI, Stroke, Heart Failure, and Death eFigure 4. 9-Protein Prognostic Model Hazard Ratios Adjusted for Framingham Variables eFigure 5. 9-Protein Risk Scores by Event Type This supplementary material has been provided by the authors to give readers additional information about their work. Downloaded From: https://jamanetwork.com/ on 10/02/2021 Supplemental Material Table of Contents 1 Study Design and Data Processing ......................................................................................................... 3 2 Table of 1130 Proteins Measured .......................................................................................................... 4 3 Variable Selection and Statistical Modeling ........................................................................................ -

Arnau Soler2019.Pdf
This thesis has been submitted in fulfilment of the requirements for a postgraduate degree (e.g. PhD, MPhil, DClinPsychol) at the University of Edinburgh. Please note the following terms and conditions of use: This work is protected by copyright and other intellectual property rights, which are retained by the thesis author, unless otherwise stated. A copy can be downloaded for personal non-commercial research or study, without prior permission or charge. This thesis cannot be reproduced or quoted extensively from without first obtaining permission in writing from the author. The content must not be changed in any way or sold commercially in any format or medium without the formal permission of the author. When referring to this work, full bibliographic details including the author, title, awarding institution and date of the thesis must be given. Genetic responses to environmental stress underlying major depressive disorder Aleix Arnau Soler Doctor of Philosophy The University of Edinburgh 2019 Declaration I hereby declare that this thesis has been composed by myself and that the work presented within has not been submitted for any other degree or professional qualification. I confirm that the work submitted is my own, except where work which has formed part of jointly-authored publications has been included. My contribution and those of the other authors to this work are indicated below. I confirm that appropriate credit has been given within this thesis where reference has been made to the work of others. I composed this thesis under guidance of Dr. Pippa Thomson. Chapter 2 has been published in PLOS ONE and is attached in the Appendix A, chapter 4 and chapter 5 are published in Translational Psychiatry and are attached in the Appendix C and D, and I expect to submit chapter 6 as a manuscript for publication. -

Assessing the Human Canonical Protein Count[Version 1; Peer Review
F1000Research 2017, 6:448 Last updated: 15 JUL 2020 REVIEW Last rolls of the yoyo: Assessing the human canonical protein count [version 1; peer review: 1 approved, 2 approved with reservations] Christopher Southan IUPHAR/BPS Guide to Pharmacology, Centre for Integrative Physiology, University of Edinburgh, Edinburgh, EH8 9XD, UK First published: 07 Apr 2017, 6:448 Open Peer Review v1 https://doi.org/10.12688/f1000research.11119.1 Latest published: 07 Apr 2017, 6:448 https://doi.org/10.12688/f1000research.11119.1 Reviewer Status Abstract Invited Reviewers In 2004, when the protein estimate from the finished human genome was 1 2 3 only 24,000, the surprise was compounded as reviewed estimates fell to 19,000 by 2014. However, variability in the total canonical protein counts version 1 (i.e. excluding alternative splice forms) of open reading frames (ORFs) in 07 Apr 2017 report report report different annotation portals persists. This work assesses these differences and possible causes. A 16-year analysis of Ensembl and UniProtKB/Swiss-Prot shows convergence to a protein number of ~20,000. The former had shown some yo-yoing, but both have now plateaued. Nine 1 Michael Tress, Spanish National Cancer major annotation portals, reviewed at the beginning of 2017, gave a spread Research Centre (CNIO), Madrid, Spain of counts from 21,819 down to 18,891. The 4-way cross-reference concordance (within UniProt) between Ensembl, Swiss-Prot, Entrez Gene 2 Elspeth A. Bruford , European Molecular and the Human Gene Nomenclature Committee (HGNC) drops to 18,690, Biology Laboratory, Hinxton, UK indicating methodological differences in protein definitions and experimental existence support between sources. -

Shear Stress Modulates Gene Expression in Normal Human Dermal Fibroblasts
University of Calgary PRISM: University of Calgary's Digital Repository Graduate Studies The Vault: Electronic Theses and Dissertations 2017 Shear Stress Modulates Gene Expression in Normal Human Dermal Fibroblasts Zabinyakov, Nikita Zabinyakov, N. (2017). Shear Stress Modulates Gene Expression in Normal Human Dermal Fibroblasts (Unpublished master's thesis). University of Calgary, Calgary, AB. doi:10.11575/PRISM/27775 http://hdl.handle.net/11023/3639 master thesis University of Calgary graduate students retain copyright ownership and moral rights for their thesis. You may use this material in any way that is permitted by the Copyright Act or through licensing that has been assigned to the document. For uses that are not allowable under copyright legislation or licensing, you are required to seek permission. Downloaded from PRISM: https://prism.ucalgary.ca UNIVERSITY OF CALGARY Shear Stress Modulates Gene Expression in Normal Human Dermal Fibroblasts by Nikita Zabinyakov A THESIS SUBMITTED TO THE FACULTY OF GRADUATE STUDIES IN PARTIAL FULFILMENT OF THE REQUIREMENTS FOR THE DEGREE OF MASTER OF SCIENCE GRADUATE PROGRAM IN BIOMEDICAL ENGINEERING CALGARY, ALBERTA JANUARY 2017 © Nikita Zabinyakov 2017 Abstract Applied mechanical forces, such as those resulting from fluid flow, trigger cells to change their functional behavior or phenotype. However, there is little known about how fluid flow affects fibroblasts. The hypothesis of this thesis is that dermal fibroblasts undergo significant changes of expression of differentiation genes after exposure to fluid flow (or shear stress). To test the hypothesis, human dermal fibroblasts were exposed to laminar steady fluid flow for 20 and 40 hours and RNA was collected for microarray analysis. -
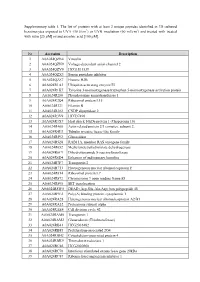
Supplementary Table 1. the List of Proteins with at Least 2 Unique
Supplementary table 1. The list of proteins with at least 2 unique peptides identified in 3D cultured keratinocytes exposed to UVA (30 J/cm2) or UVB irradiation (60 mJ/cm2) and treated with treated with rutin [25 µM] or/and ascorbic acid [100 µM]. Nr Accession Description 1 A0A024QZN4 Vinculin 2 A0A024QZN9 Voltage-dependent anion channel 2 3 A0A024QZV0 HCG1811539 4 A0A024QZX3 Serpin peptidase inhibitor 5 A0A024QZZ7 Histone H2B 6 A0A024R1A3 Ubiquitin-activating enzyme E1 7 A0A024R1K7 Tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein 8 A0A024R280 Phosphoserine aminotransferase 1 9 A0A024R2Q4 Ribosomal protein L15 10 A0A024R321 Filamin B 11 A0A024R382 CNDP dipeptidase 2 12 A0A024R3V9 HCG37498 13 A0A024R3X7 Heat shock 10kDa protein 1 (Chaperonin 10) 14 A0A024R408 Actin related protein 2/3 complex, subunit 2, 15 A0A024R4U3 Tubulin tyrosine ligase-like family 16 A0A024R592 Glucosidase 17 A0A024R5Z8 RAB11A, member RAS oncogene family 18 A0A024R652 Methylenetetrahydrofolate dehydrogenase 19 A0A024R6C9 Dihydrolipoamide S-succinyltransferase 20 A0A024R6D4 Enhancer of rudimentary homolog 21 A0A024R7F7 Transportin 2 22 A0A024R7T3 Heterogeneous nuclear ribonucleoprotein F 23 A0A024R814 Ribosomal protein L7 24 A0A024R872 Chromosome 9 open reading frame 88 25 A0A024R895 SET translocation 26 A0A024R8W0 DEAD (Asp-Glu-Ala-Asp) box polypeptide 48 27 A0A024R9E2 Poly(A) binding protein, cytoplasmic 1 28 A0A024RA28 Heterogeneous nuclear ribonucleoprotein A2/B1 29 A0A024RA52 Proteasome subunit alpha 30 A0A024RAE4 Cell division cycle 42 31 -

Conserved Exchange of Paralog Proteins During Neuronal
bioRxiv preprint doi: https://doi.org/10.1101/2021.07.22.453347; this version posted July 23, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Conserved exchange of paralog proteins during neuronal 2 differentiation 3 4 Domenico Di Fraia1, Mihaela Anitei1, Marie-Therese Mackmull*2, Luca Parca*3, Laura 5 Behrendt1, Amparo Andres-Pons4, Darren Gilmour5, Manuela Helmer Citterich3, Christoph 6 Kaether1, Martin Beck6 and Alessandro Ori1# 7 8 Affiliations 9 1 - Leibniz Institute on Aging - Fritz Lipmann Institute (FLI) Beutenbergstraße 1107745 Jena, Germany 10 2 - ETH Zurich Institute of Molecular Systems Biology Otto-Stern-Weg 3, 8093 Zürich, Switzerland 11 3 - Department of Biology, University of Tor Vergata, Rome, Italy 12 4 - European Molecular Biology Laboratory - EMBL, Meyerhofstraße 1, 69117, Heidelberg, Germany 13 5 - University of Zurich, Department of Molecular Life Sciences, Rämistrasse 71 CH-8006 Zürich, Switzerland 14 6 - Max Planck Institute of Biophysics, department of Molecular Sociology, Max-von-Laue-Straße 3, 60438 Frankfurt am Main 15 16 * contributed equally # 17 correspondence to [email protected] 18 19 20 21 22 Abstract 23 Gene duplication enables the emergence of new functions by lowering the general 24 evolutionary pressure. Previous studies have highlighted the role of specific paralog genes 25 during cell differentiation, e.g., in chromatin remodeling complexes. It remains unexplored 26 whether similar mechanisms extend to other biological functions and whether the regulation 27 of paralog genes is conserved across species.