M2(D MO.Zlol PYRIMIDINE METABOLISM
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Antiviral Defense Via Nucleotide Depletion in Bacteria
bioRxiv preprint doi: https://doi.org/10.1101/2021.04.26.441389; this version posted April 26, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Antiviral defense via nucleotide depletion in bacteria Nitzan Tal1, Adi Millman1, Avigail Stokar-Avihail1, Taya Fedorenko1, Azita Leavitt1, Sarah Melamed1, Erez Yirmiya1, Carmel Avraham1, Gil Amitai1, Rotem Sorek1,# 1 Department of Molecular Genetics, Weizmann Institute of Science, Rehovot 76100, Israel # Corresponding author: [email protected] Abstract DNA viruses and retroviruses need to consume large quantities of deoxynucleotides (dNTPs) when replicating within infected cells. The human antiviral factor SAMHD1 takes advantage of this vulnerability in the viral life cycle, and inhibits viral replication by degrading dNTPs into their constituent deoxynucleosides and inorganic phosphate. In this study we report that bacteria employ a similar strategy to defend against phage infection. We found a family of defensive dCTP deaminase proteins that, in response to phage infection, convert dCTP into deoxy-uracil nucleotides. A second family of phage resistance genes encode dGTPase enzymes, which degrade dGTP into phosphate-free deoxy- guanosine (dG) and are distant homologs of the human SAMHD1. Our results show that the defensive proteins completely eliminate the specific deoxynucleotide (either dCTP or dGTP) from the nucleotide pool during phage infection, thus starving the phage of an essential DNA building block and halting its replication. Both defensive genes are found in a diverse set of bacterial species and are specifically enriched in Vibrio genomes. Our study demonstrates that manipulation of the deoxynucleotide pool is a potent antiviral strategy shared by both prokaryotes and eukaryotes. -

1/05661 1 Al
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date _ . ... - 12 May 2011 (12.05.2011) W 2 11/05661 1 Al (51) International Patent Classification: (81) Designated States (unless otherwise indicated, for every C12Q 1/00 (2006.0 1) C12Q 1/48 (2006.0 1) kind of national protection available): AE, AG, AL, AM, C12Q 1/42 (2006.01) AO, AT, AU, AZ, BA, BB, BG, BH, BR, BW, BY, BZ, CA, CH, CL, CN, CO, CR, CU, CZ, DE, DK, DM, DO, (21) Number: International Application DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, PCT/US20 10/054171 HN, HR, HU, ID, IL, IN, IS, JP, KE, KG, KM, KN, KP, (22) International Filing Date: KR, KZ, LA, LC, LK, LR, LS, LT, LU, LY, MA, MD, 26 October 2010 (26.10.2010) ME, MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, OM, PE, PG, PH, PL, PT, RO, RS, RU, SC, SD, (25) Filing Language: English SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, TR, (26) Publication Language: English TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. (30) Priority Data: (84) Designated States (unless otherwise indicated, for every 61/255,068 26 October 2009 (26.10.2009) US kind of regional protection available): ARIPO (BW, GH, GM, KE, LR, LS, MW, MZ, NA, SD, SL, SZ, TZ, UG, (71) Applicant (for all designated States except US): ZM, ZW), Eurasian (AM, AZ, BY, KG, KZ, MD, RU, TJ, MYREXIS, INC. -

Supplementary Information
Supplementary information (a) (b) Figure S1. Resistant (a) and sensitive (b) gene scores plotted against subsystems involved in cell regulation. The small circles represent the individual hits and the large circles represent the mean of each subsystem. Each individual score signifies the mean of 12 trials – three biological and four technical. The p-value was calculated as a two-tailed t-test and significance was determined using the Benjamini-Hochberg procedure; false discovery rate was selected to be 0.1. Plots constructed using Pathway Tools, Omics Dashboard. Figure S2. Connectivity map displaying the predicted functional associations between the silver-resistant gene hits; disconnected gene hits not shown. The thicknesses of the lines indicate the degree of confidence prediction for the given interaction, based on fusion, co-occurrence, experimental and co-expression data. Figure produced using STRING (version 10.5) and a medium confidence score (approximate probability) of 0.4. Figure S3. Connectivity map displaying the predicted functional associations between the silver-sensitive gene hits; disconnected gene hits not shown. The thicknesses of the lines indicate the degree of confidence prediction for the given interaction, based on fusion, co-occurrence, experimental and co-expression data. Figure produced using STRING (version 10.5) and a medium confidence score (approximate probability) of 0.4. Figure S4. Metabolic overview of the pathways in Escherichia coli. The pathways involved in silver-resistance are coloured according to respective normalized score. Each individual score represents the mean of 12 trials – three biological and four technical. Amino acid – upward pointing triangle, carbohydrate – square, proteins – diamond, purines – vertical ellipse, cofactor – downward pointing triangle, tRNA – tee, and other – circle. -
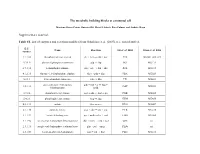
The Metabolic Building Blocks of a Minimal Cell Supplementary
The metabolic building blocks of a minimal cell Mariana Reyes-Prieto, Rosario Gil, Mercè Llabrés, Pere Palmer and Andrés Moya Supplementary material. Table S1. List of enzymes and reactions modified from Gabaldon et. al. (2007). n.i.: non identified. E.C. Name Reaction Gil et. al. 2004 Glass et. al. 2006 number 2.7.1.69 phosphotransferase system glc + pep → g6p + pyr PTS MG041, 069, 429 5.3.1.9 glucose-6-phosphate isomerase g6p ↔ f6p PGI MG111 2.7.1.11 6-phosphofructokinase f6p + atp → fbp + adp PFK MG215 4.1.2.13 fructose-1,6-bisphosphate aldolase fbp ↔ gdp + dhp FBA MG023 5.3.1.1 triose-phosphate isomerase gdp ↔ dhp TPI MG431 glyceraldehyde-3-phosphate gdp + nad + p ↔ bpg + 1.2.1.12 GAP MG301 dehydrogenase nadh 2.7.2.3 phosphoglycerate kinase bpg + adp ↔ 3pg + atp PGK MG300 5.4.2.1 phosphoglycerate mutase 3pg ↔ 2pg GPM MG430 4.2.1.11 enolase 2pg ↔ pep ENO MG407 2.7.1.40 pyruvate kinase pep + adp → pyr + atp PYK MG216 1.1.1.27 lactate dehydrogenase pyr + nadh ↔ lac + nad LDH MG460 1.1.1.94 sn-glycerol-3-phosphate dehydrogenase dhp + nadh → g3p + nad GPS n.i. 2.3.1.15 sn-glycerol-3-phosphate acyltransferase g3p + pal → mag PLSb n.i. 2.3.1.51 1-acyl-sn-glycerol-3-phosphate mag + pal → dag PLSc MG212 acyltransferase 2.7.7.41 phosphatidate cytidyltransferase dag + ctp → cdp-dag + pp CDS MG437 cdp-dag + ser → pser + 2.7.8.8 phosphatidylserine synthase PSS n.i. cmp 4.1.1.65 phosphatidylserine decarboxylase pser → peta PSD n.i. -

Supplementary Informations SI2. Supplementary Table 1
Supplementary Informations SI2. Supplementary Table 1. M9, soil, and rhizosphere media composition. LB in Compound Name Exchange Reaction LB in soil LBin M9 rhizosphere H2O EX_cpd00001_e0 -15 -15 -10 O2 EX_cpd00007_e0 -15 -15 -10 Phosphate EX_cpd00009_e0 -15 -15 -10 CO2 EX_cpd00011_e0 -15 -15 0 Ammonia EX_cpd00013_e0 -7.5 -7.5 -10 L-glutamate EX_cpd00023_e0 0 -0.0283302 0 D-glucose EX_cpd00027_e0 -0.61972444 -0.04098397 0 Mn2 EX_cpd00030_e0 -15 -15 -10 Glycine EX_cpd00033_e0 -0.0068175 -0.00693094 0 Zn2 EX_cpd00034_e0 -15 -15 -10 L-alanine EX_cpd00035_e0 -0.02780553 -0.00823049 0 Succinate EX_cpd00036_e0 -0.0056245 -0.12240603 0 L-lysine EX_cpd00039_e0 0 -10 0 L-aspartate EX_cpd00041_e0 0 -0.03205557 0 Sulfate EX_cpd00048_e0 -15 -15 -10 L-arginine EX_cpd00051_e0 -0.0068175 -0.00948672 0 L-serine EX_cpd00054_e0 0 -0.01004986 0 Cu2+ EX_cpd00058_e0 -15 -15 -10 Ca2+ EX_cpd00063_e0 -15 -100 -10 L-ornithine EX_cpd00064_e0 -0.0068175 -0.00831712 0 H+ EX_cpd00067_e0 -15 -15 -10 L-tyrosine EX_cpd00069_e0 -0.0068175 -0.00233919 0 Sucrose EX_cpd00076_e0 0 -0.02049199 0 L-cysteine EX_cpd00084_e0 -0.0068175 0 0 Cl- EX_cpd00099_e0 -15 -15 -10 Glycerol EX_cpd00100_e0 0 0 -10 Biotin EX_cpd00104_e0 -15 -15 0 D-ribose EX_cpd00105_e0 -0.01862144 0 0 L-leucine EX_cpd00107_e0 -0.03596182 -0.00303228 0 D-galactose EX_cpd00108_e0 -0.25290619 -0.18317325 0 L-histidine EX_cpd00119_e0 -0.0068175 -0.00506825 0 L-proline EX_cpd00129_e0 -0.01102953 0 0 L-malate EX_cpd00130_e0 -0.03649016 -0.79413596 0 D-mannose EX_cpd00138_e0 -0.2540567 -0.05436649 0 Co2 EX_cpd00149_e0 -

Biosynthesis and Function of Modified Bases in Bacteria and Their Viruses
This is an open access article published under a Creative Commons Non-Commercial No Derivative Works (CC-BY-NC-ND) Attribution License, which permits copying and redistribution of the article, and creation of adaptations, all for non-commercial purposes. Review pubs.acs.org/CR Biosynthesis and Function of Modified Bases in Bacteria and Their Viruses Peter Weigele Chemical Biology, New England Biolabs, Ipswich, Massachusetts 01938, United States Elisabeth A. Raleigh* Research, New England Biolabs, Ipswich, Massachusetts 01938, United States *S Supporting Information ABSTRACT: Naturally occurring modification of the canonical A, G, C, and T bases can be found in the DNA of cellular organisms and viruses from all domains of life. Bacterial viruses (bacteriophages) are a particularly rich but still underexploited source of such modified variant nucleotides. The modifications conserve the coding and base-pairing functions of DNA, but add regulatory and protective functions. In prokaryotes, modified bases appear primarily to be part of an arms race between bacteriophages (and other genomic parasites) and their hosts, although, as in eukaryotes, some modifications have been adapted to convey epigenetic information. The first half of this review catalogs the identification and diversity of DNA modifications found in bacteria and bacteriophages. What is known about the biogenesis, context, and function of these modifications are also described. The second part of the review places these DNA modifications in the context of the arms race between bacteria and bacteriophages. It focuses particularly on the defense and counter-defense strategies that turn on direct recognition of the presence of a modified base. Where modification has been shown to affect other DNA transactions, such as expression and chromosome segregation, that is summarized, with reference to recent reviews. -
Generate Metabolic Map Poster
Authors: Zheng Zhao, Delft University of Technology Marcel A. van den Broek, Delft University of Technology S. Aljoscha Wahl, Delft University of Technology Wilbert H. Heijne, DSM Biotechnology Center Roel A. Bovenberg, DSM Biotechnology Center Joseph J. Heijnen, Delft University of Technology An online version of this diagram is available at BioCyc.org. Biosynthetic pathways are positioned in the left of the cytoplasm, degradative pathways on the right, and reactions not assigned to any pathway are in the far right of the cytoplasm. Transporters and membrane proteins are shown on the membrane. Marco A. van den Berg, DSM Biotechnology Center Peter J.T. Verheijen, Delft University of Technology Periplasmic (where appropriate) and extracellular reactions and proteins may also be shown. Pathways are colored according to their cellular function. PchrCyc: Penicillium rubens Wisconsin 54-1255 Cellular Overview Connections between pathways are omitted for legibility. Liang Wu, DSM Biotechnology Center Walter M. van Gulik, Delft University of Technology L-quinate phosphate a sugar a sugar a sugar a sugar multidrug multidrug a dicarboxylate phosphate a proteinogenic 2+ 2+ + met met nicotinate Mg Mg a cation a cation K + L-fucose L-fucose L-quinate L-quinate L-quinate ammonium UDP ammonium ammonium H O pro met amino acid a sugar a sugar a sugar a sugar a sugar a sugar a sugar a sugar a sugar a sugar a sugar K oxaloacetate L-carnitine L-carnitine L-carnitine 2 phosphate quinic acid brain-specific hypothetical hypothetical hypothetical hypothetical -

1 SUPPLEMENTARY INFORMATION 1 2 an Archaeal Symbiont
1 SUPPLEMENTARY INFORMATION 2 3 An archaeal symbiont-host association from the deep terrestrial subsurface 4 5 Katrin Schwank1, Till L. V. Bornemann1, Nina Dombrowski2, Anja Spang2,3, Jillian F. Banfield4 and 6 Alexander J. Probst1* 7 8 9 1 Group for Aquatic Microbial Ecology (GAME), Biofilm Center, Department of Chemistry, 10 University of Duisburg-Essen, Germany 11 2Department of Marine Microbiology and Biogeochemistry (MMB), Royal Netherlands Institute 12 for Sea Research (NIOZ), and Utrecht University, Den Burg, Netherlands 13 3Department of Cell- and Molecular Biology, Science for Life Laboratory, Uppsala University, SE- 14 75123, Uppsala, Sweden 15 4 Department for Earth and Planetary Sciences, University of California, Berkeley, USA 16 17 *to whom the correspondence should be addressed: 18 Alexander J. Probst 19 University of Duisburg-Essen 20 Universitätsstrasse 4 21 45141 Essen 22 [email protected] 23 24 25 Content: 26 1. Supplementary Methods 27 2. Supplementary Tables 28 3. Supplementary Figure 29 4. References 30 31 32 33 1. Supplementary methods 34 Phylogenetic analysis 35 Hmm profiles of the 37 maker genes used by the phylosift v1.01 software [1, 2] were queried 36 against a representative set of archaeal genomes from NCBI (182) and the GOLD database (3) as 37 well as against the huberarchaeal population genome (for details on genomes please see 38 Supplementary Data file 1) using the phylosift search mode [1, 2]. Protein sequences 39 corresponding to each of the maker genes were subsequently aligned using mafft-linsi v7.407 [3] 40 and trimmed with BMGE-1.12 (parameters: -t AA -m BLOSUM30 -h 0.55) [4]. -

Synechocystis Pcc6803 And
STUDIES ON THE PHOTOSYNTHETIC MICROORGANISM SYNECHOCYSTIS PCC6803 AND HOW IT RESPONDS TO THE EFFECTS OF SALT STRESS BY BRADLEY LYNN POSTIER Bachelor of Science Oklahoma State University Stillwater, OK 1996 Submitted to the Faculty of the Graduate College of the Oklahoma State University in partial fulfillment of the requirements for the Degree of DOCTOR OF PHILOSOPHY December, 2003 STUDIES ON THE PHOTOSYNTHETIC MICROORGANISM SYNECHOCYSTIS PCC6803 AND HOW IT RESPONDS TO THE EFFECTS OF SALT STRESS Thesis approved: Dean of the Graduate College 11 ACKNOWLEDGMENTS I would first like to thank my wife Sheridan for all of her support and motivation. Without her, I may have never even attempted graduate school. Since I met her in 1994, she has been everything I could ask for. I look forward to spending the rest of my life with you and our as of yet unborn son. I would also like to acknowledge all the work and effort put forth by my advisor - Dr. Burnap. Rob made every possible effort to support my research financially. He guided my intellectual'maturation throughout my experience here. I don't think anyone else could have shown the patience necessary to help me achieve my goals. I am truly appreciative for what he has helped me achieve and hope that some day I can do the same for other students working under me. To my parents, I would like to say thank you for all the great support guidance and memories. Y cm have been very supportive through all my years in school, allowing me to make all of the important decisions myself, good or bad. -

Supplemental Data Heidel Et Al
Supplemental data Heidel et al. Table of Contents 1. Sequencing strategy and statistics ...................................................................................................... 2 2. Genome structure ............................................................................................................................... 2 2.1 Extrachromosal elements .............................................................................................................. 2 2.2 Chromosome structure ................................................................................................................. 3 2.3 Repetitive elements ...................................................................................................................... 5 3. Coding sequences ................................................................................................................................ 5 3.1 Homopolymer tracts ..................................................................................................................... 5 3.2 Gene families and orthology relationships ................................................................................... 7 3.3 Synteny analysis .......................................................................................................................... 11 4. Protein functional domains ............................................................................................................... 12 5. Protein families ................................................................................................................................. -

(12) United States Patent (10) Patent No.: US 7820,442 B2 Petersen-Mahrt Et Al
USOO7820442B2 (12) United States Patent (10) Patent No.: US 7820,442 B2 Petersen-Mahrt et al. (45) Date of Patent: Oct. 26, 2010 (54) ACTIVATION INDUCED DEAMINASE (AID) Yamanaka et al., Apollipoprotein B mRNA-editing protein induces hepatocellular carcinoma and dysplasia in transgenic animals, Proc. (75) Inventors: Svend K. Petersen-Mahrt, Cambridge Natl. Acad. Sci. USA, Biochemisty, Aug. 1995, vol. 92, pp. 8483 (GB); Reuben Harris, Cambridge (GB); 8487. International Search Report Related To PCT/GB 03/02002. Michael Samuel Neuberger, Cambridge Martin A. et al., Activation-Induced Cytidine Deaminase Turns On (GB); Rupert Christopher Somatic Hypermutation. In Hybridomas, Nature, vol. 415, Feb. 24. Landsdowne Beale, Cambridge (GB) 2002, pp. 802-806. Jarmuz A. et al. An Anthropoid-Specific Locus Of Orphan C To U (73) Assignee: Medical Research Council, London RNA-Editing Enzymes On Chromosome-22, Genomics, vol. 79, No. (GB) 3, Mar. 2002, pp. 285-296. Manis JP, Mechanism And Control OfClass-Switch Recombination, (*) Notice: Subject to any disclaimer, the term of this Trends in Immunology, Elsevier, Cambridge, GB, vol. 23, No. 1, Jan. patent is extended or adjusted under 35 1, 2002, pp. 31-39. U.S.C. 154(b) by 0 days. MacGinnitie A.J. et al., Mutagenesis. Of Apobec-1. The Catalytic Subunit Of The Mammalian Apollipoprotein B mRNA Editing (21) Appl. No.: 10/985,321 Enzyme, Reveals Distinct Domains That Meiate Cytosine Nucleoside Deaminase, RNA Binding, and RNA Editing Activity, The Journal of Biological Chemistry, vol. 270, No. 24. Jun. 16, 1995, (22) Filed: Nov. 10, 2004 pp. 14768-14775. Okazaki IM. et al., The Aid Enzyme Induces Class Switch Recom (65) Prior Publication Data bination. -

12) United States Patent (10
US007635572B2 (12) UnitedO States Patent (10) Patent No.: US 7,635,572 B2 Zhou et al. (45) Date of Patent: Dec. 22, 2009 (54) METHODS FOR CONDUCTING ASSAYS FOR 5,506,121 A 4/1996 Skerra et al. ENZYME ACTIVITY ON PROTEIN 5,510,270 A 4/1996 Fodor et al. MICROARRAYS 5,512,492 A 4/1996 Herron et al. 5,516,635 A 5/1996 Ekins et al. (75) Inventors: Fang X. Zhou, New Haven, CT (US); 5,532,128 A 7/1996 Eggers Barry Schweitzer, Cheshire, CT (US) 5,538,897 A 7/1996 Yates, III et al. s s 5,541,070 A 7/1996 Kauvar (73) Assignee: Life Technologies Corporation, .. S.E. al Carlsbad, CA (US) 5,585,069 A 12/1996 Zanzucchi et al. 5,585,639 A 12/1996 Dorsel et al. (*) Notice: Subject to any disclaimer, the term of this 5,593,838 A 1/1997 Zanzucchi et al. patent is extended or adjusted under 35 5,605,662 A 2f1997 Heller et al. U.S.C. 154(b) by 0 days. 5,620,850 A 4/1997 Bamdad et al. 5,624,711 A 4/1997 Sundberg et al. (21) Appl. No.: 10/865,431 5,627,369 A 5/1997 Vestal et al. 5,629,213 A 5/1997 Kornguth et al. (22) Filed: Jun. 9, 2004 (Continued) (65) Prior Publication Data FOREIGN PATENT DOCUMENTS US 2005/O118665 A1 Jun. 2, 2005 EP 596421 10, 1993 EP 0619321 12/1994 (51) Int. Cl. EP O664452 7, 1995 CI2O 1/50 (2006.01) EP O818467 1, 1998 (52) U.S.