Mice Deficient in Apobec2 and Apobec3 7271
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Cellular and Molecular Signatures in the Disease Tissue of Early
Cellular and Molecular Signatures in the Disease Tissue of Early Rheumatoid Arthritis Stratify Clinical Response to csDMARD-Therapy and Predict Radiographic Progression Frances Humby1,* Myles Lewis1,* Nandhini Ramamoorthi2, Jason Hackney3, Michael Barnes1, Michele Bombardieri1, Francesca Setiadi2, Stephen Kelly1, Fabiola Bene1, Maria di Cicco1, Sudeh Riahi1, Vidalba Rocher-Ros1, Nora Ng1, Ilias Lazorou1, Rebecca E. Hands1, Desiree van der Heijde4, Robert Landewé5, Annette van der Helm-van Mil4, Alberto Cauli6, Iain B. McInnes7, Christopher D. Buckley8, Ernest Choy9, Peter Taylor10, Michael J. Townsend2 & Costantino Pitzalis1 1Centre for Experimental Medicine and Rheumatology, William Harvey Research Institute, Barts and The London School of Medicine and Dentistry, Queen Mary University of London, Charterhouse Square, London EC1M 6BQ, UK. Departments of 2Biomarker Discovery OMNI, 3Bioinformatics and Computational Biology, Genentech Research and Early Development, South San Francisco, California 94080 USA 4Department of Rheumatology, Leiden University Medical Center, The Netherlands 5Department of Clinical Immunology & Rheumatology, Amsterdam Rheumatology & Immunology Center, Amsterdam, The Netherlands 6Rheumatology Unit, Department of Medical Sciences, Policlinico of the University of Cagliari, Cagliari, Italy 7Institute of Infection, Immunity and Inflammation, University of Glasgow, Glasgow G12 8TA, UK 8Rheumatology Research Group, Institute of Inflammation and Ageing (IIA), University of Birmingham, Birmingham B15 2WB, UK 9Institute of -

Antiviral Defense Via Nucleotide Depletion in Bacteria
bioRxiv preprint doi: https://doi.org/10.1101/2021.04.26.441389; this version posted April 26, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Antiviral defense via nucleotide depletion in bacteria Nitzan Tal1, Adi Millman1, Avigail Stokar-Avihail1, Taya Fedorenko1, Azita Leavitt1, Sarah Melamed1, Erez Yirmiya1, Carmel Avraham1, Gil Amitai1, Rotem Sorek1,# 1 Department of Molecular Genetics, Weizmann Institute of Science, Rehovot 76100, Israel # Corresponding author: [email protected] Abstract DNA viruses and retroviruses need to consume large quantities of deoxynucleotides (dNTPs) when replicating within infected cells. The human antiviral factor SAMHD1 takes advantage of this vulnerability in the viral life cycle, and inhibits viral replication by degrading dNTPs into their constituent deoxynucleosides and inorganic phosphate. In this study we report that bacteria employ a similar strategy to defend against phage infection. We found a family of defensive dCTP deaminase proteins that, in response to phage infection, convert dCTP into deoxy-uracil nucleotides. A second family of phage resistance genes encode dGTPase enzymes, which degrade dGTP into phosphate-free deoxy- guanosine (dG) and are distant homologs of the human SAMHD1. Our results show that the defensive proteins completely eliminate the specific deoxynucleotide (either dCTP or dGTP) from the nucleotide pool during phage infection, thus starving the phage of an essential DNA building block and halting its replication. Both defensive genes are found in a diverse set of bacterial species and are specifically enriched in Vibrio genomes. Our study demonstrates that manipulation of the deoxynucleotide pool is a potent antiviral strategy shared by both prokaryotes and eukaryotes. -

1/05661 1 Al
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date _ . ... - 12 May 2011 (12.05.2011) W 2 11/05661 1 Al (51) International Patent Classification: (81) Designated States (unless otherwise indicated, for every C12Q 1/00 (2006.0 1) C12Q 1/48 (2006.0 1) kind of national protection available): AE, AG, AL, AM, C12Q 1/42 (2006.01) AO, AT, AU, AZ, BA, BB, BG, BH, BR, BW, BY, BZ, CA, CH, CL, CN, CO, CR, CU, CZ, DE, DK, DM, DO, (21) Number: International Application DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, PCT/US20 10/054171 HN, HR, HU, ID, IL, IN, IS, JP, KE, KG, KM, KN, KP, (22) International Filing Date: KR, KZ, LA, LC, LK, LR, LS, LT, LU, LY, MA, MD, 26 October 2010 (26.10.2010) ME, MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, OM, PE, PG, PH, PL, PT, RO, RS, RU, SC, SD, (25) Filing Language: English SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, TR, (26) Publication Language: English TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. (30) Priority Data: (84) Designated States (unless otherwise indicated, for every 61/255,068 26 October 2009 (26.10.2009) US kind of regional protection available): ARIPO (BW, GH, GM, KE, LR, LS, MW, MZ, NA, SD, SL, SZ, TZ, UG, (71) Applicant (for all designated States except US): ZM, ZW), Eurasian (AM, AZ, BY, KG, KZ, MD, RU, TJ, MYREXIS, INC. -

Interactions Between APOBEC3 and Murine Retroviruses: Mechanisms of Restriction and Drug Resistance
University of Pennsylvania ScholarlyCommons Publicly Accessible Penn Dissertations 2013 Interactions Between APOBEC3 and Murine Retroviruses: Mechanisms of Restriction and Drug Resistance Alyssa Lea MacMillan University of Pennsylvania, [email protected] Follow this and additional works at: https://repository.upenn.edu/edissertations Part of the Virology Commons Recommended Citation MacMillan, Alyssa Lea, "Interactions Between APOBEC3 and Murine Retroviruses: Mechanisms of Restriction and Drug Resistance" (2013). Publicly Accessible Penn Dissertations. 894. https://repository.upenn.edu/edissertations/894 This paper is posted at ScholarlyCommons. https://repository.upenn.edu/edissertations/894 For more information, please contact [email protected]. Interactions Between APOBEC3 and Murine Retroviruses: Mechanisms of Restriction and Drug Resistance Abstract APOBEC3 proteins are important for antiretroviral defense in mammals. The activity of these factors has been well characterized in vitro, identifying cytidine deamination as an active source of viral restriction leading to hypermutation of viral DNA synthesized during reverse transcription. These mutations can result in viral lethality via disruption of critical genes, but in some cases is insufficiento t completely obstruct viral replication. This sublethal level of mutagenesis could aid in viral evolution. A cytidine deaminase-independent mechanism of restriction has also been identified, as catalytically inactive proteins are still able to inhibit infection in vitro. Murine retroviruses do not exhibit characteristics of hypermutation by mouse APOBEC3 in vivo. However, human APOBEC3G protein expressed in transgenic mice maintains antiviral restriction and actively deaminates viral genomes. The mechanism by which endogenous APOBEC3 proteins function is unclear. The mouse provides a system amenable to studying the interaction of APOBEC3 and retroviral targets in vivo. -

Global Gene Expression Analysis of the Mouse Colonic Mucosa Treated
BMC Cancer BioMed Central Research article Open Access Global gene expression analysis of the mouse colonic mucosa treated with azoxymethane and dextran sodium sulfate Rikako Suzuki1, Shingo Miyamoto2, Yumiko Yasui1, Shigeyuki Sugie1 and Takuji Tanaka*1 Address: 1Department of Oncologic Pathology, Kanazawa Medical University, 1-1 Daigaku, Uchinada, Ishikawa 920-0293, Japan and 2Division of Food Science and Biotechnology, Graduate School of Agriculture, Kyoto University, Kyoto 606-8502, Japan Email: Rikako Suzuki - [email protected]; Shingo Miyamoto - [email protected]; Yumiko Yasui - y-yasui@kanazawa- med.ac.jp; Shigeyuki Sugie - [email protected]; Takuji Tanaka* - [email protected] * Corresponding author Published: 17 May 2007 Received: 10 July 2006 Accepted: 17 May 2007 BMC Cancer 2007, 7:84 doi:10.1186/1471-2407-7-84 This article is available from: http://www.biomedcentral.com/1471-2407/7/84 © 2007 Suzuki et al; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Abstract Background: Chronic inflammation is well known to be a risk factor for colon cancer. Previously we established a novel mouse model of inflammation-related colon carcinogenesis, which is useful to examine the involvement of inflammation in colon carcinogenesis. To shed light on the alterations in global gene expression in the background of inflammation-related colon cancer and gain further insights into the molecular mechanisms underlying inflammation-related colon carcinogenesis, we conducted a comprehensive DNA microarray analysis using our model. -

Supplementary Information
Supplementary information (a) (b) Figure S1. Resistant (a) and sensitive (b) gene scores plotted against subsystems involved in cell regulation. The small circles represent the individual hits and the large circles represent the mean of each subsystem. Each individual score signifies the mean of 12 trials – three biological and four technical. The p-value was calculated as a two-tailed t-test and significance was determined using the Benjamini-Hochberg procedure; false discovery rate was selected to be 0.1. Plots constructed using Pathway Tools, Omics Dashboard. Figure S2. Connectivity map displaying the predicted functional associations between the silver-resistant gene hits; disconnected gene hits not shown. The thicknesses of the lines indicate the degree of confidence prediction for the given interaction, based on fusion, co-occurrence, experimental and co-expression data. Figure produced using STRING (version 10.5) and a medium confidence score (approximate probability) of 0.4. Figure S3. Connectivity map displaying the predicted functional associations between the silver-sensitive gene hits; disconnected gene hits not shown. The thicknesses of the lines indicate the degree of confidence prediction for the given interaction, based on fusion, co-occurrence, experimental and co-expression data. Figure produced using STRING (version 10.5) and a medium confidence score (approximate probability) of 0.4. Figure S4. Metabolic overview of the pathways in Escherichia coli. The pathways involved in silver-resistance are coloured according to respective normalized score. Each individual score represents the mean of 12 trials – three biological and four technical. Amino acid – upward pointing triangle, carbohydrate – square, proteins – diamond, purines – vertical ellipse, cofactor – downward pointing triangle, tRNA – tee, and other – circle. -

Table SII. Significantly Differentially Expressed Mrnas of GSE23558 Data Series with the Criteria of Adjusted P<0.05 And
Table SII. Significantly differentially expressed mRNAs of GSE23558 data series with the criteria of adjusted P<0.05 and logFC>1.5. Probe ID Adjusted P-value logFC Gene symbol Gene title A_23_P157793 1.52x10-5 6.91 CA9 carbonic anhydrase 9 A_23_P161698 1.14x10-4 5.86 MMP3 matrix metallopeptidase 3 A_23_P25150 1.49x10-9 5.67 HOXC9 homeobox C9 A_23_P13094 3.26x10-4 5.56 MMP10 matrix metallopeptidase 10 A_23_P48570 2.36x10-5 5.48 DHRS2 dehydrogenase A_23_P125278 3.03x10-3 5.40 CXCL11 C-X-C motif chemokine ligand 11 A_23_P321501 1.63x10-5 5.38 DHRS2 dehydrogenase A_23_P431388 2.27x10-6 5.33 SPOCD1 SPOC domain containing 1 A_24_P20607 5.13x10-4 5.32 CXCL11 C-X-C motif chemokine ligand 11 A_24_P11061 3.70x10-3 5.30 CSAG1 chondrosarcoma associated gene 1 A_23_P87700 1.03x10-4 5.25 MFAP5 microfibrillar associated protein 5 A_23_P150979 1.81x10-2 5.25 MUCL1 mucin like 1 A_23_P1691 2.71x10-8 5.12 MMP1 matrix metallopeptidase 1 A_23_P350005 2.53x10-4 5.12 TRIML2 tripartite motif family like 2 A_24_P303091 1.23x10-3 4.99 CXCL10 C-X-C motif chemokine ligand 10 A_24_P923612 1.60x10-5 4.95 PTHLH parathyroid hormone like hormone A_23_P7313 6.03x10-5 4.94 SPP1 secreted phosphoprotein 1 A_23_P122924 2.45x10-8 4.93 INHBA inhibin A subunit A_32_P155460 6.56x10-3 4.91 PICSAR P38 inhibited cutaneous squamous cell carcinoma associated lincRNA A_24_P686965 8.75x10-7 4.82 SH2D5 SH2 domain containing 5 A_23_P105475 7.74x10-3 4.70 SLCO1B3 solute carrier organic anion transporter family member 1B3 A_24_P85099 4.82x10-5 4.67 HMGA2 high mobility group AT-hook 2 A_24_P101651 -
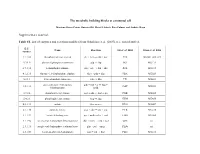
The Metabolic Building Blocks of a Minimal Cell Supplementary
The metabolic building blocks of a minimal cell Mariana Reyes-Prieto, Rosario Gil, Mercè Llabrés, Pere Palmer and Andrés Moya Supplementary material. Table S1. List of enzymes and reactions modified from Gabaldon et. al. (2007). n.i.: non identified. E.C. Name Reaction Gil et. al. 2004 Glass et. al. 2006 number 2.7.1.69 phosphotransferase system glc + pep → g6p + pyr PTS MG041, 069, 429 5.3.1.9 glucose-6-phosphate isomerase g6p ↔ f6p PGI MG111 2.7.1.11 6-phosphofructokinase f6p + atp → fbp + adp PFK MG215 4.1.2.13 fructose-1,6-bisphosphate aldolase fbp ↔ gdp + dhp FBA MG023 5.3.1.1 triose-phosphate isomerase gdp ↔ dhp TPI MG431 glyceraldehyde-3-phosphate gdp + nad + p ↔ bpg + 1.2.1.12 GAP MG301 dehydrogenase nadh 2.7.2.3 phosphoglycerate kinase bpg + adp ↔ 3pg + atp PGK MG300 5.4.2.1 phosphoglycerate mutase 3pg ↔ 2pg GPM MG430 4.2.1.11 enolase 2pg ↔ pep ENO MG407 2.7.1.40 pyruvate kinase pep + adp → pyr + atp PYK MG216 1.1.1.27 lactate dehydrogenase pyr + nadh ↔ lac + nad LDH MG460 1.1.1.94 sn-glycerol-3-phosphate dehydrogenase dhp + nadh → g3p + nad GPS n.i. 2.3.1.15 sn-glycerol-3-phosphate acyltransferase g3p + pal → mag PLSb n.i. 2.3.1.51 1-acyl-sn-glycerol-3-phosphate mag + pal → dag PLSc MG212 acyltransferase 2.7.7.41 phosphatidate cytidyltransferase dag + ctp → cdp-dag + pp CDS MG437 cdp-dag + ser → pser + 2.7.8.8 phosphatidylserine synthase PSS n.i. cmp 4.1.1.65 phosphatidylserine decarboxylase pser → peta PSD n.i. -

Deaminase-Independent Mode of Antiretroviral Action in Human and Mouse APOBEC3 Proteins
microorganisms Review Deaminase-Independent Mode of Antiretroviral Action in Human and Mouse APOBEC3 Proteins Yoshiyuki Hakata 1,* and Masaaki Miyazawa 1,2 1 Department of Immunology, Kindai University Faculty of Medicine, 377-2 Ohno-Higashi, Osaka-Sayama, Osaka 589-8511, Japan; [email protected] 2 Kindai University Anti-Aging Center, 3-4-1 Kowakae, Higashiosaka, Osaka 577-8502, Japan * Correspondence: [email protected]; Tel.: +81-72-367-7660 Received: 8 December 2020; Accepted: 9 December 2020; Published: 12 December 2020 Abstract: Apolipoprotein B mRNA editing enzyme, catalytic polypeptide-like 3 (APOBEC3) proteins (APOBEC3s) are deaminases that convert cytosines to uracils predominantly on a single-stranded DNA, and function as intrinsic restriction factors in the innate immune system to suppress replication of viruses (including retroviruses) and movement of retrotransposons. Enzymatic activity is supposed to be essential for the APOBEC3 antiviral function. However, it is not the only way that APOBEC3s exert their biological function. Since the discovery of human APOBEC3G as a restriction factor for HIV-1, the deaminase-independent mode of action has been observed. At present, it is apparent that both the deaminase-dependent and -independent pathways are tightly involved not only in combating viruses but also in human tumorigenesis. Although the deaminase-dependent pathway has been extensively characterized so far, understanding of the deaminase-independent pathway remains immature. Here, we review existing knowledge regarding the deaminase-independent antiretroviral functions of APOBEC3s and their molecular mechanisms. We also discuss the possible unidentified molecular mechanism for the deaminase-independent antiretroviral function mediated by mouse APOBEC3. Keywords: APOBEC3; deaminase-independent antiretroviral function; innate immunity 1. -

Supplementary Informations SI2. Supplementary Table 1
Supplementary Informations SI2. Supplementary Table 1. M9, soil, and rhizosphere media composition. LB in Compound Name Exchange Reaction LB in soil LBin M9 rhizosphere H2O EX_cpd00001_e0 -15 -15 -10 O2 EX_cpd00007_e0 -15 -15 -10 Phosphate EX_cpd00009_e0 -15 -15 -10 CO2 EX_cpd00011_e0 -15 -15 0 Ammonia EX_cpd00013_e0 -7.5 -7.5 -10 L-glutamate EX_cpd00023_e0 0 -0.0283302 0 D-glucose EX_cpd00027_e0 -0.61972444 -0.04098397 0 Mn2 EX_cpd00030_e0 -15 -15 -10 Glycine EX_cpd00033_e0 -0.0068175 -0.00693094 0 Zn2 EX_cpd00034_e0 -15 -15 -10 L-alanine EX_cpd00035_e0 -0.02780553 -0.00823049 0 Succinate EX_cpd00036_e0 -0.0056245 -0.12240603 0 L-lysine EX_cpd00039_e0 0 -10 0 L-aspartate EX_cpd00041_e0 0 -0.03205557 0 Sulfate EX_cpd00048_e0 -15 -15 -10 L-arginine EX_cpd00051_e0 -0.0068175 -0.00948672 0 L-serine EX_cpd00054_e0 0 -0.01004986 0 Cu2+ EX_cpd00058_e0 -15 -15 -10 Ca2+ EX_cpd00063_e0 -15 -100 -10 L-ornithine EX_cpd00064_e0 -0.0068175 -0.00831712 0 H+ EX_cpd00067_e0 -15 -15 -10 L-tyrosine EX_cpd00069_e0 -0.0068175 -0.00233919 0 Sucrose EX_cpd00076_e0 0 -0.02049199 0 L-cysteine EX_cpd00084_e0 -0.0068175 0 0 Cl- EX_cpd00099_e0 -15 -15 -10 Glycerol EX_cpd00100_e0 0 0 -10 Biotin EX_cpd00104_e0 -15 -15 0 D-ribose EX_cpd00105_e0 -0.01862144 0 0 L-leucine EX_cpd00107_e0 -0.03596182 -0.00303228 0 D-galactose EX_cpd00108_e0 -0.25290619 -0.18317325 0 L-histidine EX_cpd00119_e0 -0.0068175 -0.00506825 0 L-proline EX_cpd00129_e0 -0.01102953 0 0 L-malate EX_cpd00130_e0 -0.03649016 -0.79413596 0 D-mannose EX_cpd00138_e0 -0.2540567 -0.05436649 0 Co2 EX_cpd00149_e0 -

1 APOBEC-Mediated Mutagenesis in Urothelial Carcinoma Is Associated
bioRxiv preprint doi: https://doi.org/10.1101/123802; this version posted April 4, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. APOBEC-mediated mutagenesis in urothelial carcinoma is associated with improved survival, mutations in DNA damage response genes, and immune response Alexander P. Glaser MD, Damiano Fantini PhD, Kalen J. Rimar MD, Joshua J. Meeks MD PhD APG, DF, KJR, JJM: Northwestern University, Department of Urology, Chicago, IL, 60607 Running title: APOBEC mutagenesis in bladder cancer *Corresponding author: Joshua J. Meeks, MD PhD 303 E. Chicago Ave. Tarry 16-703 Chicago, IL 60611 Email: [email protected] Keywords (4-6): • Urinary bladder neoplasms • APOBEC Deaminases • Mutagenesis • DNA damage • Interferon Abbreviations and Acronyms: TCGA – The Cancer Genome Atlas ssDNA – single stranded DNA APOBEC –apolipoprotein B mRNA editing catalytic polypeptide-like GCAC – Genome Data Analysis Center MAF – mutation annotation format “APOBEC-high” – tumors enriched for APOBEC mutagenesis “APOBEC-low” – tumors not enriched for APOBEC mutagenesis 1 bioRxiv preprint doi: https://doi.org/10.1101/123802; this version posted April 4, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract: Background: The APOBEC family of enzymes is responsible for a mutation signature characterized by a TCW>T/G mutation. APOBEC-mediated mutagenesis is implicated in a wide variety of tumors, including bladder cancer. In this study, we explore the APOBEC mutational signature in bladder cancer and the relationship with specific mutations, molecular subtype, gene expression, and survival. -

AID Downregulation Is a Novel Function of the DNMT Inhibitor 5-Aza-Deoxycytidine
www.impactjournals.com/oncotarget/ Oncotarget, January, Vol. 5, No. 1 AID downregulation is a novel function of the DNMT inhibitor 5-aza-deoxycytidine Chiou-Tsun Tsai1, Pei-Ming Yang1, Ting-Rong Chern3, Shu-Hui Chuang1, Jung-Hsin Lin2,3,4, Lars Klemm5, Markus Müschen5 and Ching-Chow Chen1 1 Department of Pharmacology, College of Medicine, National Taiwan University, Taipei, Taiwan 2 School of Pharmacy, College of Medicine, National Taiwan University, Taipei, Taiwan 3 Research Center for Applied Sciences, Academia Sinica, Taipei, Taiwan 4 Institute of Biomedical Science, Academia Sinica, Taipei, Taiwan 5 Department of Laboratory Medicine, University of California San Francisco, San Francisco, California Correspondence to: Ching-Chow Chen, email: [email protected] Keywords: AID, 5-aza-CdR, Zebularine, DNMT1 Received: August 23, 2013 Accepted: November 23, 2013 Published: November 25, 2013 This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. ABSTRACT: Activation-induced cytidine deaminase (AID) was originally identified as an inducer of somatic hypermutation (SHM) and class switch recombination (CSR) in immunoglobulin genes. However, AID can also cause mutations in host genes and contribute to cancer progression and drug resistance. In this study, molecular docking showed the interaction of free 5-aza-CdR and Zebularine (Zeb) with AID. However, only 5-aza-CdR-incorporated ssDNA bound to the active site of AID and inhibited AID expression through proteasomal degradation. 5-aza-CdR demonstrated cytotoxicity against AID-positive and -negative hematopoietic cancer cells.