Transcriptional and Epigenetic Regulation of Oligodendrocyte Development and Myelination in the Central Nervous System
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Hes5 Regulates the Transition Timing of Neurogenesis and Gliogenesis In
© 2017. Published by The Company of Biologists Ltd | Development (2017) 144, 3156-3167 doi:10.1242/dev.147256 RESEARCH ARTICLE Hes5 regulates the transition timing of neurogenesis and gliogenesis in mammalian neocortical development Shama Bansod1,2, Ryoichiro Kageyama1,2,3,4 and Toshiyuki Ohtsuka1,2,3,* ABSTRACT has been reported that the high mobility group AT-hook (Hgma) During mammalian neocortical development, neural stem/progenitor genes regulate gene expression by modulating chromatin structure cells (NSCs) sequentially give rise to deep layer neurons and (Ozturk et al., 2014), maintain neurogenic NSCs, and inhibit superficial layer neurons through mid- to late-embryonic stages, gliogenesis during early- to mid-embryonic stages through global shifting to gliogenic phase at perinatal stages. Previously, we found opening of the chromatin state (Kishi et al., 2012). However, the that the Hes genes inhibit neuronal differentiation and maintain mechanism by which the expression of these epigenetic factors is NSCs. Here, we generated transgenic mice that overexpress Hes5 in controlled remains to be analyzed. NSCs of the central nervous system, and found that the transition Here, we found that Hes5, a transcriptional repressor acting timing from deep to superficial layer neurogenesis was shifted earlier, as an effector of Notch signaling, regulates the timing of while gliogenesis precociously occurred in the developing neocortex neurogenesis and gliogenesis via alteration in the expression of Hes5-overexpressing mice. By contrast, the transition from deep to levels of epigenetic factors. Notch signaling contributes to the superficial layer neurogenesis and the onset of gliogenesis were elaboration of cellular diversity during the development of various delayed in Hes5 knockout (KO) mice. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Transcriptome Dynamics and Potential Roles of Sox6 in the Postnatal Heart
RESEARCH ARTICLE Transcriptome Dynamics and Potential Roles of Sox6 in the Postnatal Heart Chung-Il An1☯*, Yasunori Ichihashi2☯¤a¤b*, Jie Peng3, Neelima R. Sinha2, Nobuko Hagiwara1* 1 Division of Cardiovascular Medicine, Department of Internal Medicine, University of California Davis, Davis, California, United States of America, 2 Department of Plant Biology, University of California Davis, Davis, California, United States of America, 3 Department of Statistics, University of California Davis, Davis, California, United States of America ☯ These authors contributed equally to this work. a11111 ¤a Current address: RIKEN Center for Sustainable Resource Science, Yokohama, Kanagawa, Japan ¤b Current address: JST, PRESTO, Kawaguchi, Saitama, Japan * [email protected] (CA); [email protected] (YI); [email protected] (NH) Abstract OPEN ACCESS The postnatal heart undergoes highly coordinated developmental processes culminating in Citation: An C-I, Ichihashi Y, Peng J, Sinha NR, the complex physiologic properties of the adult heart. The molecular mechanisms of postna- Hagiwara N (2016) Transcriptome Dynamics and tal heart development remain largely unexplored despite their important clinical implications. Potential Roles of Sox6 in the Postnatal Heart. To gain an integrated view of the dynamic changes in gene expression during postnatal PLoS ONE 11(11): e0166574. doi:10.1371/journal. heart development at the organ level, time-series transcriptome analyses of the postnatal pone.0166574 hearts of neonatal through adult mice (P1, P7, P14, P30, and P60) were performed using a Editor: Katherine Yutzey, Cincinnati Children's newly developed bioinformatics pipeline. We identified functional gene clusters by principal Hospital Medical Center, UNITED STATES component analysis with self-organizing map clustering which revealed organized, discrete Received: July 16, 2016 gene expression patterns corresponding to biological functions associated with the neona- Accepted: October 31, 2016 tal, juvenile and adult stages of postnatal heart development. -

Prox1regulates the Subtype-Specific Development of Caudal Ganglionic
The Journal of Neuroscience, September 16, 2015 • 35(37):12869–12889 • 12869 Development/Plasticity/Repair Prox1 Regulates the Subtype-Specific Development of Caudal Ganglionic Eminence-Derived GABAergic Cortical Interneurons X Goichi Miyoshi,1 Allison Young,1 Timothy Petros,1 Theofanis Karayannis,1 Melissa McKenzie Chang,1 Alfonso Lavado,2 Tomohiko Iwano,3 Miho Nakajima,4 Hiroki Taniguchi,5 Z. Josh Huang,5 XNathaniel Heintz,4 Guillermo Oliver,2 Fumio Matsuzaki,3 Robert P. Machold,1 and Gord Fishell1 1Department of Neuroscience and Physiology, NYU Neuroscience Institute, Smilow Research Center, New York University School of Medicine, New York, New York 10016, 2Department of Genetics & Tumor Cell Biology, St. Jude Children’s Research Hospital, Memphis, Tennessee 38105, 3Laboratory for Cell Asymmetry, RIKEN Center for Developmental Biology, Kobe 650-0047, Japan, 4Laboratory of Molecular Biology, Howard Hughes Medical Institute, GENSAT Project, The Rockefeller University, New York, New York 10065, and 5Cold Spring Harbor Laboratory, Cold Spring Harbor, New York 11724 Neurogliaform (RELNϩ) and bipolar (VIPϩ) GABAergic interneurons of the mammalian cerebral cortex provide critical inhibition locally within the superficial layers. While these subtypes are known to originate from the embryonic caudal ganglionic eminence (CGE), the specific genetic programs that direct their positioning, maturation, and integration into the cortical network have not been eluci- dated. Here, we report that in mice expression of the transcription factor Prox1 is selectively maintained in postmitotic CGE-derived cortical interneuron precursors and that loss of Prox1 impairs the integration of these cells into superficial layers. Moreover, Prox1 differentially regulates the postnatal maturation of each specific subtype originating from the CGE (RELN, Calb2/VIP, and VIP). -
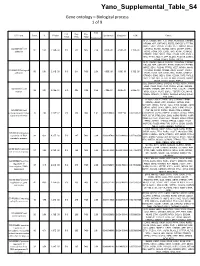
Supplemental Tables4.Pdf
Yano_Supplemental_Table_S4 Gene ontology – Biological process 1 of 9 Fold List Pop Pop GO Term Count % PValue Bonferroni Benjamini FDR Genes Total Hits Total Enrichment DLC1, CADM1, NELL2, CLSTN1, PCDHGA8, CTNNB1, NRCAM, APP, CNTNAP2, FERT2, RAPGEF1, PTPRM, MPDZ, SDK1, PCDH9, PTPRS, VEZT, NRXN1, MYH9, GO:0007155~cell CTNNA2, NCAM1, NCAM2, DDR1, LSAMP, CNTN1, 50 5.61 2.14E-08 510 311 7436 2.34 4.50E-05 4.50E-05 3.70E-05 adhesion ROR2, VCAN, DST, LIMS1, TNC, ASTN1, CTNND2, CTNND1, CDH2, NEO1, CDH4, CD24A, FAT3, PVRL3, TRO, TTYH1, MLLT4, LPP, NLGN1, PCDH19, LAMA1, ITGA9, CDH13, CDON, PSPC1 DLC1, CADM1, NELL2, CLSTN1, PCDHGA8, CTNNB1, NRCAM, APP, CNTNAP2, FERT2, RAPGEF1, PTPRM, MPDZ, SDK1, PCDH9, PTPRS, VEZT, NRXN1, MYH9, GO:0022610~biological CTNNA2, NCAM1, NCAM2, DDR1, LSAMP, CNTN1, 50 5.61 2.14E-08 510 311 7436 2.34 4.50E-05 4.50E-05 3.70E-05 adhesion ROR2, VCAN, DST, LIMS1, TNC, ASTN1, CTNND2, CTNND1, CDH2, NEO1, CDH4, CD24A, FAT3, PVRL3, TRO, TTYH1, MLLT4, LPP, NLGN1, PCDH19, LAMA1, ITGA9, CDH13, CDON, PSPC1 DCC, ENAH, PLXNA2, CAPZA2, ATP5B, ASTN1, PAX6, ZEB2, CDH2, CDH4, GLI3, CD24A, EPHB1, NRCAM, GO:0006928~cell CTTNBP2, EDNRB, APP, PTK2, ETV1, CLASP2, STRBP, 36 4.04 3.46E-07 510 205 7436 2.56 7.28E-04 3.64E-04 5.98E-04 motion NRG1, DCLK1, PLAT, SGPL1, TGFBR1, EVL, MYH9, YWHAE, NCKAP1, CTNNA2, SEMA6A, EPHA4, NDEL1, FYN, LRP6 PLXNA2, ADCY5, PAX6, GLI3, CTNNB1, LPHN2, EDNRB, LPHN3, APP, CSNK2A1, GPR45, NRG1, RAPGEF1, WWOX, SGPL1, TLE4, SPEN, NCAM1, DDR1, GRB10, GRM3, GNAQ, HIPK1, GNB1, HIPK2, PYGO1, GO:0007166~cell RNF138, ROR2, CNTN1, -

TF Activation Profiling Plate Array II Signosis, Inc
Signosis, Inc. Innovative Plate Assay Solutions TF Activation Profiling Plate Array II Catalog Number: FA-1002 (For Research Use Only) Introduction Materials Provided with the Kit Transcription factors (TFs) are a group of cellular proteins that play essential roles in regulating gene Component Qty Store at expression. They act as sensors to monitor cellular 96-Well Plates (with 2 RT changes and convert signals into gene expression. aluminum adhesive seal) Often, a specific cellular signal pathway can activate Isolation Columns 2 RT multiple TFs. The expression of a specific gene can Elution Buffer 400µL RT also be under the control of multiple TFs. Thus, TF Plate Hybridization Buffer 20mL RT monitoring the activation of multiple TFs 5X Plate Hybridization Wash 60mL RT simultaneously is critical to understanding the Buffer molecular mechanism of cellular regulation underlying 5X Detection Wash Buffer 60mL RT cell signaling and gene expression. Signosis, Inc.’s TF Blocking Buffer 60mL RT Activation Profiling Plate Array II is used for Filter Wash Buffer 5mL 4°C monitoring 96 different TFs simultaneously from one Filter Binding Buffer 1mL 4°C sample. Substrate A 2mL 4°C Substrate B 2mL 4°C Principle of the assay Streptavidin-HRP Conjugate 40µL 4°C Substrate Dilution Buffer 16mL 4°C Signosis, Inc.’s TF Activation Profiling Plate Array II TF Binding Buffer Mix 60µL -20°C is used for monitoring the activation of multiple TFs TF Probe Mix II 20µL -20°C simultaneously. With this technology a series of biotin-labeled probes are made based on the consensus sequences of TF DNA-binding sites. -

The GATA2 Transcription Factor Negatively Regulates the Proliferation of Neuronal Progenitors
RESEARCH ARTICLE 2155 Development 133, 2155-2165 (2006) doi:10.1242/dev.02377 The GATA2 transcription factor negatively regulates the proliferation of neuronal progenitors Abeer El Wakil*, Cédric Francius*,†, Annie Wolff, Jocelyne Pleau-Varet† and Jeannette Nardelli†,§ Postmitotic neurons are produced from a pool of cycling progenitors in an orderly fashion that requires proper spatial and temporal coordination of proliferation, fate determination, differentiation and morphogenesis. This probably relies on complex interplay between mechanisms that control cell cycle, specification and differentiation. In this respect, we have studied the possible implication of GATA2, a transcription factor that is involved in several neuronal specification pathways, in the control of the proliferation of neural progenitors in the embryonic spinal cord. Using gain- and loss-of-function manipulations, we have shown that Gata2 can drive neural progenitors out of the cycle and, to some extent, into differentiation. This correlates with the control of cyclin D1 transcription and of the expression of the p27/Kip1 protein. Interestingly, this functional aspect is not only associated with silencing of the Notch pathway but also appears to be independent of proneural function. Consistently, GATA2 also controls the proliferation capacity of mouse embryonic neuroepithelial cells in culture. Indeed, Gata2 inactivation enhances the proliferation rate in these cells. By contrast, GATA2 overexpression is sufficient to force such cells and neuroblastoma cells to stop dividing but not to drive either type of cell into differentiation. Furthermore, a non-cell autonomous effect of Gata2 expression was observed in vivo as well as in vitro. Hence, our data have provided evidence for the ability of Gata2 to inhibit the proliferation of neural progenitors, and they further suggest that, in this regard, Gata2 can operate independently of neuronal differentiation. -

Olig1 and Sox10 Interact Synergistically to Drivemyelin Basic
The Journal of Neuroscience, December 26, 2007 • 27(52):14375–14382 • 14375 Cellular/Molecular Olig1 and Sox10 Interact Synergistically to Drive Myelin Basic Protein Transcription in Oligodendrocytes Huiliang Li,1 Yan Lu,2 Hazel K. Smith,1 and William D. Richardson1 1Wolfson Institute for Biomedical Research and Department of Biology, University College London, London WC1E 6BT, United Kingdom, and 2Medical Research Council, Clinical Sciences Centre, Imperial College London, London W12 0NN, United Kingdom The oligodendrocyte lineage genes (Olig1/2), encoding basic helix-loop-helix transcription factors, were first identified in screens for master regulators of oligodendrocyte development. OLIG1 is important for differentiation of oligodendrocyte precursors into myelin- forming oligodendrocytes during development and is thought to play a crucial role in remyelination during multiple sclerosis. However, itisstillunclearhowOLIG1interactswithitstranscriptionalcofactorsandDNAtargets.OLIG1wasreportedlyrestrictedtomammals,but we demonstrate here that zebrafish and other teleosts also possess an OLIG1 homolog. In zebrafish, as in mammals, Olig1 is expressed in the oligodendrocyte lineage. Olig1 associates physically with another myelin-associated transcription factor, Sox10, and the Olig1/Sox10 complex activates mbp (myelin basic protein) transcription via conserved DNA sequence motifs in the mbp promoter region. In contrast, Olig2 does not bind to Sox10 in zebrafish, although both OLIG1 and OLIG2 bind SOX10 in mouse. Key words: Olig1; Olig2; Sox10; Mbp; oligodendrocyte; myelin; zebrafish; mouse; evolution; development Introduction directly regulates Mbp transcription (Stolt et al., 2002), and over- Myelin, the multilayered glial sheath around axons, is one of the expression of SOX10 alone is sufficient to induce myelin gene defining features of jawed vertebrates (gnathostomes). It is expression in embryonic chick spinal cord (Liu et al., 2007). -

Genome-Wide DNA Methylation Analysis of KRAS Mutant Cell Lines Ben Yi Tew1,5, Joel K
www.nature.com/scientificreports OPEN Genome-wide DNA methylation analysis of KRAS mutant cell lines Ben Yi Tew1,5, Joel K. Durand2,5, Kirsten L. Bryant2, Tikvah K. Hayes2, Sen Peng3, Nhan L. Tran4, Gerald C. Gooden1, David N. Buckley1, Channing J. Der2, Albert S. Baldwin2 ✉ & Bodour Salhia1 ✉ Oncogenic RAS mutations are associated with DNA methylation changes that alter gene expression to drive cancer. Recent studies suggest that DNA methylation changes may be stochastic in nature, while other groups propose distinct signaling pathways responsible for aberrant methylation. Better understanding of DNA methylation events associated with oncogenic KRAS expression could enhance therapeutic approaches. Here we analyzed the basal CpG methylation of 11 KRAS-mutant and dependent pancreatic cancer cell lines and observed strikingly similar methylation patterns. KRAS knockdown resulted in unique methylation changes with limited overlap between each cell line. In KRAS-mutant Pa16C pancreatic cancer cells, while KRAS knockdown resulted in over 8,000 diferentially methylated (DM) CpGs, treatment with the ERK1/2-selective inhibitor SCH772984 showed less than 40 DM CpGs, suggesting that ERK is not a broadly active driver of KRAS-associated DNA methylation. KRAS G12V overexpression in an isogenic lung model reveals >50,600 DM CpGs compared to non-transformed controls. In lung and pancreatic cells, gene ontology analyses of DM promoters show an enrichment for genes involved in diferentiation and development. Taken all together, KRAS-mediated DNA methylation are stochastic and independent of canonical downstream efector signaling. These epigenetically altered genes associated with KRAS expression could represent potential therapeutic targets in KRAS-driven cancer. Activating KRAS mutations can be found in nearly 25 percent of all cancers1. -

Interindividual Regulation of the BCRP/ABCG2 Transporter in Term Human Placentas
DMD Fast Forward. Published on January 31, 2018 as DOI: 10.1124/dmd.117.079228 This article has not been copyedited and formatted. The final version may differ from this version. DMD #79228 Title Page Interindividual Regulation of the BCRP/ABCG2 Transporter in Term Human Placentas Kristin M Bircsak, Jamie E Moscovitz, Xia Wen, Faith Archer, Poi Yu Sofia Yuen, Moiz Mohammed, Naureen Memon, Barry I Weinberger, Laura M Saba, Anna M Vetrano, Lauren M Aleksunes Downloaded from Department of Pharmacology and Toxicology, Rutgers, The State University of New Jersey, Ernest Mario School of Pharmacy, Piscataway, NJ, USA (K.M.B., J.E.M., X.W., L.M.A.), dmd.aspetjournals.org Department of Pediatrics, Rutgers University Robert Wood Johnson Medical School, New Brunswick, NJ, USA (F.A., P.Y.S.Y, M.M., N.M., A.M.V.), Hofstra Northwell School of Medicine, Cohen Children’s Medical Center of New York, New Hyde Park, NY, USA (B.I.W.), at ASPET Journals on October 2, 2021 Department of Pharmaceutical Sciences, Skaggs School of Pharmacy and Pharmaceutical Sciences, University of Colorado, Aurora, CO, USA (L.S.), Environmental and Occupational Health Sciences Institute, Rutgers, The State University of New Jersey, Piscataway, NJ, USA (L.M.A.), Lipid Center, Rutgers, The State University of New Jersey, Piscataway, NJ, USA (L.M.A.) 1 DMD Fast Forward. Published on January 31, 2018 as DOI: 10.1124/dmd.117.079228 This article has not been copyedited and formatted. The final version may differ from this version. DMD #79228 Running Title Page Running title: Interindividual -

Accompanies CD8 T Cell Effector Function Global DNA Methylation
Global DNA Methylation Remodeling Accompanies CD8 T Cell Effector Function Christopher D. Scharer, Benjamin G. Barwick, Benjamin A. Youngblood, Rafi Ahmed and Jeremy M. Boss This information is current as of October 1, 2021. J Immunol 2013; 191:3419-3429; Prepublished online 16 August 2013; doi: 10.4049/jimmunol.1301395 http://www.jimmunol.org/content/191/6/3419 Downloaded from Supplementary http://www.jimmunol.org/content/suppl/2013/08/20/jimmunol.130139 Material 5.DC1 References This article cites 81 articles, 25 of which you can access for free at: http://www.jimmunol.org/content/191/6/3419.full#ref-list-1 http://www.jimmunol.org/ Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists by guest on October 1, 2021 • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Global DNA Methylation Remodeling Accompanies CD8 T Cell Effector Function Christopher D. Scharer,* Benjamin G. Barwick,* Benjamin A. Youngblood,*,† Rafi Ahmed,*,† and Jeremy M. -

SUPPLEMENTARY MATERIAL Bone Morphogenetic Protein 4 Promotes
www.intjdevbiol.com doi: 10.1387/ijdb.160040mk SUPPLEMENTARY MATERIAL corresponding to: Bone morphogenetic protein 4 promotes craniofacial neural crest induction from human pluripotent stem cells SUMIYO MIMURA, MIKA SUGA, KAORI OKADA, MASAKI KINEHARA, HIROKI NIKAWA and MIHO K. FURUE* *Address correspondence to: Miho Kusuda Furue. Laboratory of Stem Cell Cultures, National Institutes of Biomedical Innovation, Health and Nutrition, 7-6-8, Saito-Asagi, Ibaraki, Osaka 567-0085, Japan. Tel: 81-72-641-9819. Fax: 81-72-641-9812. E-mail: [email protected] Full text for this paper is available at: http://dx.doi.org/10.1387/ijdb.160040mk TABLE S1 PRIMER LIST FOR QRT-PCR Gene forward reverse AP2α AATTTCTCAACCGACAACATT ATCTGTTTTGTAGCCAGGAGC CDX2 CTGGAGCTGGAGAAGGAGTTTC ATTTTAACCTGCCTCTCAGAGAGC DLX1 AGTTTGCAGTTGCAGGCTTT CCCTGCTTCATCAGCTTCTT FOXD3 CAGCGGTTCGGCGGGAGG TGAGTGAGAGGTTGTGGCGGATG GAPDH CAAAGTTGTCATGGATGACC CCATGGAGAAGGCTGGGG MSX1 GGATCAGACTTCGGAGAGTGAACT GCCTTCCCTTTAACCCTCACA NANOG TGAACCTCAGCTACAAACAG TGGTGGTAGGAAGAGTAAAG OCT4 GACAGGGGGAGGGGAGGAGCTAGG CTTCCCTCCAACCAGTTGCCCCAAA PAX3 TTGCAATGGCCTCTCAC AGGGGAGAGCGCGTAATC PAX6 GTCCATCTTTGCTTGGGAAA TAGCCAGGTTGCGAAGAACT p75 TCATCCCTGTCTATTGCTCCA TGTTCTGCTTGCAGCTGTTC SOX9 AATGGAGCAGCGAAATCAAC CAGAGAGATTTAGCACACTGATC SOX10 GACCAGTACCCGCACCTG CGCTTGTCACTTTCGTTCAG Suppl. Fig. S1. Comparison of the gene expression profiles of the ES cells and the cells induced by NC and NC-B condition. Scatter plots compares the normalized expression of every gene on the array (refer to Table S3). The central line