Characterization of Transcription Factor Networks Involved in Umbilical Cord Blood CD34+ Stem Cells-Derived Erythropoiesis
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Discovery of Progenitor Cell Signatures by Time-Series Synexpression Analysis During Drosophila Embryonic Cell Immortalization
Correction DEVELOPMENTAL BIOLOGY Correction for “Discovery of progenitor cell signatures by time- series synexpression analysis during Drosophila embryonic cell immortalization,” by Mary-Lee Dequéant, Delphine Fagegaltier, Yanhui Hu, Kerstin Spirohn, Amanda Simcox, Gregory J. Hannon, and Norbert Perrimon, which appeared in issue 42, October 20, 2015, of Proc Natl Acad Sci USA (112:12974–12979; first published October 5, 2015; 10.1073/pnas.1517729112). The authors note that Delphine Fagegaltier should be credited for designing research and performing research. The authors also note that Delphine Fagegaltier, Amanda Simcox, and Gregory J. Hannon should be credited for contributing to the writing of the paper. The corrected author contributions footnote appears below. Author contributions: M.-L.D., D.F., A.S., G.J.H., and N.P. designed research; M.-L.D., D.F., K.S., and A.S. performed research; M.-L.D., D.F., and Y.H. analyzed data; and M.-L.D. and N.P. wrote the paper with contributions from D.F., A.S., and G.J.H. www.pnas.org/cgi/doi/10.1073/pnas.1520482112 E6408 | PNAS | November 17, 2015 | vol. 112 | no. 46 www.pnas.org Downloaded by guest on September 25, 2021 Discovery of progenitor cell signatures by time-series synexpression analysis during Drosophila embryonic cell immortalization Mary-Lee Dequéanta,1, Delphine Fagegaltierb, Yanhui Hua, Kerstin Spirohna, Amanda Simcoxc, Gregory J. Hannond, and Norbert Perrimona,e,1 aDepartment of Genetics, Harvard Medical School, Boston, MA 02115, bCold Spring Harbor Laboratories, Cold Spring Harbor, NY 11724; cDepartment of Molecular Genetics, The Ohio State University, Columbus, OH 43210; dHoward Hughes Medical Institute, Cold Spring Harbor Laboratories, Cold Spring Harbor, NY 11724; and eHoward Hughes Medical Institute, Harvard Medical School, Boston, MA 02115 Contributed by Norbert Perrimon, September 10, 2015 (sent for review May 18, 2015; reviewed by Peter Cherbas, Gary Karpen, and Renato Paro) The use of time series profiling to identify groups of functionally population contributing to adult muscles (4–7). -

The GATA2 Transcription Factor Negatively Regulates the Proliferation of Neuronal Progenitors
RESEARCH ARTICLE 2155 Development 133, 2155-2165 (2006) doi:10.1242/dev.02377 The GATA2 transcription factor negatively regulates the proliferation of neuronal progenitors Abeer El Wakil*, Cédric Francius*,†, Annie Wolff, Jocelyne Pleau-Varet† and Jeannette Nardelli†,§ Postmitotic neurons are produced from a pool of cycling progenitors in an orderly fashion that requires proper spatial and temporal coordination of proliferation, fate determination, differentiation and morphogenesis. This probably relies on complex interplay between mechanisms that control cell cycle, specification and differentiation. In this respect, we have studied the possible implication of GATA2, a transcription factor that is involved in several neuronal specification pathways, in the control of the proliferation of neural progenitors in the embryonic spinal cord. Using gain- and loss-of-function manipulations, we have shown that Gata2 can drive neural progenitors out of the cycle and, to some extent, into differentiation. This correlates with the control of cyclin D1 transcription and of the expression of the p27/Kip1 protein. Interestingly, this functional aspect is not only associated with silencing of the Notch pathway but also appears to be independent of proneural function. Consistently, GATA2 also controls the proliferation capacity of mouse embryonic neuroepithelial cells in culture. Indeed, Gata2 inactivation enhances the proliferation rate in these cells. By contrast, GATA2 overexpression is sufficient to force such cells and neuroblastoma cells to stop dividing but not to drive either type of cell into differentiation. Furthermore, a non-cell autonomous effect of Gata2 expression was observed in vivo as well as in vitro. Hence, our data have provided evidence for the ability of Gata2 to inhibit the proliferation of neural progenitors, and they further suggest that, in this regard, Gata2 can operate independently of neuronal differentiation. -

Olig1 and Sox10 Interact Synergistically to Drivemyelin Basic
The Journal of Neuroscience, December 26, 2007 • 27(52):14375–14382 • 14375 Cellular/Molecular Olig1 and Sox10 Interact Synergistically to Drive Myelin Basic Protein Transcription in Oligodendrocytes Huiliang Li,1 Yan Lu,2 Hazel K. Smith,1 and William D. Richardson1 1Wolfson Institute for Biomedical Research and Department of Biology, University College London, London WC1E 6BT, United Kingdom, and 2Medical Research Council, Clinical Sciences Centre, Imperial College London, London W12 0NN, United Kingdom The oligodendrocyte lineage genes (Olig1/2), encoding basic helix-loop-helix transcription factors, were first identified in screens for master regulators of oligodendrocyte development. OLIG1 is important for differentiation of oligodendrocyte precursors into myelin- forming oligodendrocytes during development and is thought to play a crucial role in remyelination during multiple sclerosis. However, itisstillunclearhowOLIG1interactswithitstranscriptionalcofactorsandDNAtargets.OLIG1wasreportedlyrestrictedtomammals,but we demonstrate here that zebrafish and other teleosts also possess an OLIG1 homolog. In zebrafish, as in mammals, Olig1 is expressed in the oligodendrocyte lineage. Olig1 associates physically with another myelin-associated transcription factor, Sox10, and the Olig1/Sox10 complex activates mbp (myelin basic protein) transcription via conserved DNA sequence motifs in the mbp promoter region. In contrast, Olig2 does not bind to Sox10 in zebrafish, although both OLIG1 and OLIG2 bind SOX10 in mouse. Key words: Olig1; Olig2; Sox10; Mbp; oligodendrocyte; myelin; zebrafish; mouse; evolution; development Introduction directly regulates Mbp transcription (Stolt et al., 2002), and over- Myelin, the multilayered glial sheath around axons, is one of the expression of SOX10 alone is sufficient to induce myelin gene defining features of jawed vertebrates (gnathostomes). It is expression in embryonic chick spinal cord (Liu et al., 2007). -

SUPPLEMENTARY MATERIAL Bone Morphogenetic Protein 4 Promotes
www.intjdevbiol.com doi: 10.1387/ijdb.160040mk SUPPLEMENTARY MATERIAL corresponding to: Bone morphogenetic protein 4 promotes craniofacial neural crest induction from human pluripotent stem cells SUMIYO MIMURA, MIKA SUGA, KAORI OKADA, MASAKI KINEHARA, HIROKI NIKAWA and MIHO K. FURUE* *Address correspondence to: Miho Kusuda Furue. Laboratory of Stem Cell Cultures, National Institutes of Biomedical Innovation, Health and Nutrition, 7-6-8, Saito-Asagi, Ibaraki, Osaka 567-0085, Japan. Tel: 81-72-641-9819. Fax: 81-72-641-9812. E-mail: [email protected] Full text for this paper is available at: http://dx.doi.org/10.1387/ijdb.160040mk TABLE S1 PRIMER LIST FOR QRT-PCR Gene forward reverse AP2α AATTTCTCAACCGACAACATT ATCTGTTTTGTAGCCAGGAGC CDX2 CTGGAGCTGGAGAAGGAGTTTC ATTTTAACCTGCCTCTCAGAGAGC DLX1 AGTTTGCAGTTGCAGGCTTT CCCTGCTTCATCAGCTTCTT FOXD3 CAGCGGTTCGGCGGGAGG TGAGTGAGAGGTTGTGGCGGATG GAPDH CAAAGTTGTCATGGATGACC CCATGGAGAAGGCTGGGG MSX1 GGATCAGACTTCGGAGAGTGAACT GCCTTCCCTTTAACCCTCACA NANOG TGAACCTCAGCTACAAACAG TGGTGGTAGGAAGAGTAAAG OCT4 GACAGGGGGAGGGGAGGAGCTAGG CTTCCCTCCAACCAGTTGCCCCAAA PAX3 TTGCAATGGCCTCTCAC AGGGGAGAGCGCGTAATC PAX6 GTCCATCTTTGCTTGGGAAA TAGCCAGGTTGCGAAGAACT p75 TCATCCCTGTCTATTGCTCCA TGTTCTGCTTGCAGCTGTTC SOX9 AATGGAGCAGCGAAATCAAC CAGAGAGATTTAGCACACTGATC SOX10 GACCAGTACCCGCACCTG CGCTTGTCACTTTCGTTCAG Suppl. Fig. S1. Comparison of the gene expression profiles of the ES cells and the cells induced by NC and NC-B condition. Scatter plots compares the normalized expression of every gene on the array (refer to Table S3). The central line -

Hormonal Regulation of Oligodendrogenesis I: Effects Across the Lifespan
biomolecules Review Hormonal Regulation of Oligodendrogenesis I: Effects across the Lifespan Kimberly L. P. Long 1,*,†,‡ , Jocelyn M. Breton 1,‡,§ , Matthew K. Barraza 2 , Olga S. Perloff 3 and Daniela Kaufer 1,4,5 1 Helen Wills Neuroscience Institute, University of California, Berkeley, CA 94720, USA; [email protected] (J.M.B.); [email protected] (D.K.) 2 Department of Molecular and Cellular Biology, University of California, Berkeley, CA 94720, USA; [email protected] 3 Memory and Aging Center, Department of Neurology, University of California, San Francisco, CA 94143, USA; [email protected] 4 Department of Integrative Biology, University of California, Berkeley, CA 94720, USA 5 Canadian Institute for Advanced Research, Toronto, ON M5G 1M1, Canada * Correspondence: [email protected] † Current address: Department of Psychiatry and Behavioral Sciences, University of California, San Francisco, CA 94143, USA. ‡ These authors contributed equally to this work. § Current address: Department of Psychiatry, Columbia University, New York, NY 10027, USA. Abstract: The brain’s capacity to respond to changing environments via hormonal signaling is critical to fine-tuned function. An emerging body of literature highlights a role for myelin plasticity as a prominent type of experience-dependent plasticity in the adult brain. Myelin plasticity is driven by oligodendrocytes (OLs) and their precursor cells (OPCs). OPC differentiation regulates the trajectory of myelin production throughout development, and importantly, OPCs maintain the ability to proliferate and generate new OLs throughout adulthood. The process of oligodendrogenesis, Citation: Long, K.L.P.; Breton, J.M.; the‘creation of new OLs, can be dramatically influenced during early development and in adulthood Barraza, M.K.; Perloff, O.S.; Kaufer, D. -

Inhibition of Mitochondrial Complex II in Neuronal Cells Triggers Unique
www.nature.com/scientificreports OPEN Inhibition of mitochondrial complex II in neuronal cells triggers unique pathways culminating in autophagy with implications for neurodegeneration Sathyanarayanan Ranganayaki1, Neema Jamshidi2, Mohamad Aiyaz3, Santhosh‑Kumar Rashmi4, Narayanappa Gayathri4, Pulleri Kandi Harsha5, Balasundaram Padmanabhan6 & Muchukunte Mukunda Srinivas Bharath7* Mitochondrial dysfunction and neurodegeneration underlie movement disorders such as Parkinson’s disease, Huntington’s disease and Manganism among others. As a corollary, inhibition of mitochondrial complex I (CI) and complex II (CII) by toxins 1‑methyl‑4‑phenylpyridinium (MPP+) and 3‑nitropropionic acid (3‑NPA) respectively, induced degenerative changes noted in such neurodegenerative diseases. We aimed to unravel the down‑stream pathways associated with CII inhibition and compared with CI inhibition and the Manganese (Mn) neurotoxicity. Genome‑wide transcriptomics of N27 neuronal cells exposed to 3‑NPA, compared with MPP+ and Mn revealed varied transcriptomic profle. Along with mitochondrial and synaptic pathways, Autophagy was the predominant pathway diferentially regulated in the 3‑NPA model with implications for neuronal survival. This pathway was unique to 3‑NPA, as substantiated by in silico modelling of the three toxins. Morphological and biochemical validation of autophagy markers in the cell model of 3‑NPA revealed incomplete autophagy mediated by mechanistic Target of Rapamycin Complex 2 (mTORC2) pathway. Interestingly, Brain Derived Neurotrophic Factor -

Appendix 2. Significantly Differentially Regulated Genes in Term Compared with Second Trimester Amniotic Fluid Supernatant
Appendix 2. Significantly Differentially Regulated Genes in Term Compared With Second Trimester Amniotic Fluid Supernatant Fold Change in term vs second trimester Amniotic Affymetrix Duplicate Fluid Probe ID probes Symbol Entrez Gene Name 1019.9 217059_at D MUC7 mucin 7, secreted 424.5 211735_x_at D SFTPC surfactant protein C 416.2 206835_at STATH statherin 363.4 214387_x_at D SFTPC surfactant protein C 295.5 205982_x_at D SFTPC surfactant protein C 288.7 1553454_at RPTN repetin solute carrier family 34 (sodium 251.3 204124_at SLC34A2 phosphate), member 2 238.9 206786_at HTN3 histatin 3 161.5 220191_at GKN1 gastrokine 1 152.7 223678_s_at D SFTPA2 surfactant protein A2 130.9 207430_s_at D MSMB microseminoprotein, beta- 99.0 214199_at SFTPD surfactant protein D major histocompatibility complex, class II, 96.5 210982_s_at D HLA-DRA DR alpha 96.5 221133_s_at D CLDN18 claudin 18 94.4 238222_at GKN2 gastrokine 2 93.7 1557961_s_at D LOC100127983 uncharacterized LOC100127983 93.1 229584_at LRRK2 leucine-rich repeat kinase 2 HOXD cluster antisense RNA 1 (non- 88.6 242042_s_at D HOXD-AS1 protein coding) 86.0 205569_at LAMP3 lysosomal-associated membrane protein 3 85.4 232698_at BPIFB2 BPI fold containing family B, member 2 84.4 205979_at SCGB2A1 secretoglobin, family 2A, member 1 84.3 230469_at RTKN2 rhotekin 2 82.2 204130_at HSD11B2 hydroxysteroid (11-beta) dehydrogenase 2 81.9 222242_s_at KLK5 kallikrein-related peptidase 5 77.0 237281_at AKAP14 A kinase (PRKA) anchor protein 14 76.7 1553602_at MUCL1 mucin-like 1 76.3 216359_at D MUC7 mucin 7, -

Functional Characterization of TBR1 Variants in Neurodevelopmental Disorder Received: 14 May 2018 Joery Den Hoed1, Elliot Sollis1, Hanka Venselaar2, Sara B
www.nature.com/scientificreports OPEN Functional characterization of TBR1 variants in neurodevelopmental disorder Received: 14 May 2018 Joery den Hoed1, Elliot Sollis1, Hanka Venselaar2, Sara B. Estruch1, Pelagia Deriziotis1 & Accepted: 31 August 2018 Simon E. Fisher 1,3 Published: xx xx xxxx Recurrent de novo variants in the TBR1 transcription factor are implicated in the etiology of sporadic autism spectrum disorders (ASD). Disruptions include missense variants located in the T-box DNA- binding domain and previous work has demonstrated that they disrupt TBR1 protein function. Recent screens of thousands of simplex families with sporadic ASD cases uncovered additional T-box variants in TBR1 but their etiological relevance is unclear. We performed detailed functional analyses of de novo missense TBR1 variants found in the T-box of ASD cases, assessing many aspects of protein function, including subcellular localization, transcriptional activity and protein-interactions. Only two of the three tested variants severely disrupted TBR1 protein function, despite in silico predictions that all would be deleterious. Furthermore, we characterized a putative interaction with BCL11A, a transcription factor that was recently implicated in a neurodevelopmental syndrome involving developmental delay and language defcits. Our fndings enhance understanding of molecular functions of TBR1, as well as highlighting the importance of functional testing of variants that emerge from next-generation sequencing, to decipher their contributions to neurodevelopmental disorders like ASD. TBR1 (T-box brain, 1; OMIM *604616) encodes a neuron-specifc transcription factor of the T-box family1. Te TBR1 protein is highly expressed in the deep layers of the cortex, where it is involved in diferentiation of subsets of projection neurons2–4. -
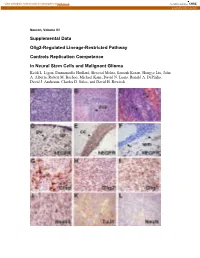
Supplemental Data Olig2-Regulated Lineage-Restricted Pathway Controls Replication Competence in Neural Stem Cells and Malignant
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Caltech Authors Neuron, Volume 53 Supplemental Data Olig2-Regulated Lineage-Restricted Pathway Controls Replication Competence in Neural Stem Cells and Malignant Glioma Keith L. Ligon, Emmanuelle Huillard, Shwetal Mehta, Santosh Kesari, Hongye Liu, John A. Alberta, Robert M. Bachoo, Michael Kane, David N. Louis, Ronald A. DePinho, David J. Anderson, Charles D. Stiles, and David H. Rowitch Figure S1. Olig1/2+/- Ink4a/Arf-/-EGFRvIII Gliomas Express Characteristic Morphologic and Immunophenotypic Features of Human Malignant Gliomas (A) Tumors exhibit dense cellularity and atypia. (B) Characteristic infiltration of host SVZ stem cell niche. (C) Although occasional tumors exhibited pseudopalisading necrosis (n, arrowhead) and hemorrhage (h, arrowhead) similar to human GBM (Astrocytoma WHO Grade IV), most tumors lacked these features. (D) IHC for hEGFR highlights hallmark feature of human gliomas, including perivascular (pv) and subpial (sp) accumulation of tumor cells, (E) striking white matter tropism of tumor cells in corpus callosum (cc), and (F) distant single cell infiltration of cerebellar white matter (wm, arrows). (G-J) Immunohistochemical markers characteristic of human tumors (Gfap, Olig2, Olig1, Nestin) are present. (K and L) Weak staining for the early neuronal marker TuJ1 and absence of the differentiated neuronal marker, NeuN similar to human glioma and consistent with heterogeneous lines of partial differentiation. Figure S2. Olig1/2+/- Ink4a/Arf-/-EGFRvIII Neurospheres Are Multipotent and Olig1/2-/- Ink4a/Arf-/-EGFRvIII Neurospheres Are Bipotent Olig1/2+/- or Olig1/2-/- Ink4a/Arf-/-EGFRvIII neurospheres were allowed to differentiate for 6 days in medium without EGF. -

Reduction of BCL11A in Hematopoietic Stem Cells Through
Science Bulletin 64 (2019) 1562–1564 Contents lists available at ScienceDirect Science Bulletin journal homepage: www.elsevier.com/locate/scib Research Highlight Reduction of BCL11A in hematopoietic stem cells through gene editing: new strategy to ameliorate the severe b-globin disorders sickle cell disease ⇑ Weiqi Hong, Mengyuan Huang, Yuquan Wei, Xiawei Wei Laboratory of Aging Research and Cancer Drug Target, State Key Laboratory of Biotherapy, National Clinical Research Center for Geriatrics, West China Hospital, Sichuan University, Chengdu 610041, China Site-specific gene editing is of great importance in precise Sickle-cell anemia is a prototypical monogenic disorder caused medicine. Two conventional genome editing methods, Zine finger by mutation of b-globin subunit. It is a promising therapy strategy nucleases (ZFNs) and transcription activator-like effector nucleases to induct fetal hemoglobin (HbF, a2c2) by re-expressing the paral- (TALENs), are based on protein-DNA recognition, with tedious ogous c-globin genes (HBG1/2) for severe b-globin disorders sickle work in constructing target protein [1,2]. Developed from immune cell disease (SCD) and b-thalassemia [9]. Researches in the past response of bacteria, CRISPR/Cas9 has been widely investigated as have shown that the core of the +58 erythroid enhancer of BCL11A a promising tool for therapeutic genome editing in clinical settings was crucial for repression of HBF in adult stage erythroid. Wu et al nowadays [3,4]. This system succeeds in gene deletion, insertion found that chemically modified synthetic sgRNAs (MS-sgRNAs) and frameshift mutations with higher efficiency, less cost, was more efficient than in vitro transcribed sgRNAs. Targeting improved flexibility and simplified designing process [5]. -

Transcriptome Alterations of Vascular Smooth Muscle Cells in Aortic Wall of Myocardial Infarction Patients
This document is downloaded from DR‑NTU (https://dr.ntu.edu.sg) Nanyang Technological University, Singapore. Transcriptome alterations of vascular smooth muscle cells in aortic wall of myocardial infarction patients Wongsurawat, Thidathip; Woo, Chin Cheng; Giannakakis, Antonis; Lin, Xiao Yun; Cheow, Esther Sok Hwee; Lee, Chuen Neng; Richards, Mark; Sze, Siu Kwan; Nookaew, Intawat; Sorokin, Vitaly; Kuznetsov, Vladimir Andreevich 2018 Wongsurawat, T., Woo, C. C., Giannakakis, A., Lin, X. Y., Cheow, E. S. H., Lee, C. N., et al. (2018). Transcriptome alterations of vascular smooth muscle cells in aortic wall of myocardial infarction patients. Data in Brief, 17, 1112‑1135. https://hdl.handle.net/10356/85590 https://doi.org/10.1016/j.dib.2018.01.108 © 2018 The Authors. Published by Elsevier Inc. This is an open access article under the CC BY license (http://creativecommons.org/licenses/by/4.0/). Downloaded on 09 Oct 2021 06:21:01 SGT Data in Brief 17 (2018) 1112–1135 Contents lists available at ScienceDirect Data in Brief journal homepage: www.elsevier.com/locate/dib Data Article Transcriptome alterations of vascular smooth muscle cells in aortic wall of myocardial infarction patients Thidathip Wongsurawat a,b, Chin Cheng Woo c, Antonis Giannakakis a, Xiao Yun Lin d, Esther Sok Hwee Cheow e, Chuen Neng Lee c,d, Mark Richards f,g, Siu Kwan Sze e, Intawat Nookaew b, Vladimir A. Kuznetsov a,h, Vitaly Sorokin c,d,⁎ a Department of Genome and Gene Expression Data Analysis, Bioinformatics Institute, Agency for Science, Technology and Research (A*STAR), -

Transcriptional and Epigenetic Characterization of Early Striosomes
bioRxiv preprint doi: https://doi.org/10.1101/2020.05.19.105171; this version posted May 22, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Transcriptional and epigenetic characterization of early striosomes identifies Foxf2 and Olig2 as factors required for development of striatal compartmentation and neuronal phenotypic differentiation Maria-Daniela Cirnaru1, Sicheng Song2, Kizito-Tshitoko Tshilenge3, Chuhyon Corwin1, Justyna Mleczko1, Carlos Galicia Aguirre3, Houda Benlhabib2, Jaroslav Bendl4,5,6, Jordi Creus Muncunill1, Pasha Apontes4,5,6, John F. Fullard4,5,6, Peter Carlsson7, Panos Roussos4,5,6,8, Sean D. Mooney2, Lisa M. Ellerby3, Michelle E. Ehrlich1* 1. Department of Neurology, Icahn School of Medicine at Mount Sinai, New York, New York 10029, USA. 2. Department of Epidemiology, University of Washington, Seattle, WA, USA. 3. Buck Institute for Research on Aging, Novato, CA, 94945, USA. 4. Pamela Sklar Division of Psychiatric Genomics, Icahn School of Medicine at Mount Sinai, One Gustave L. Levy Place, New York, NY, 10029, USA. 5. Institute for Genomics and Multiscale Biology, Department of Genetics and Genomic Sciences, Icahn School of Medicine at Mount Sinai, One Gustave L. Levy Place, New York, NY, 10029, USA. 6. Department of Psychiatry, Icahn School of Medicine at Mount Sinai, One Gustave L. Levy Place, New York, NY, 10029, USA. 7. Department of Chemistry and Molecular Biology, University of Gothenburg, Gothenburg, Sweden. 8. Mental Illness Research, Education, and Clinical Center (VISN 2 South), James J. Peters VA Medical Center, Bronx, NY 10468, USA.